Denoising Diffusion Probabilistic Models in Six Simple Steps
Feb 10, 2024Richard E. Turner, Cristiana-Diana Diaconu, Stratis Markou, Aliaksandra Shysheya, Andrew Y. K. Foong, Bruno Mlodozeniec
Denoising Diffusion Probabilistic Models (DDPMs) are a very popular class of deep generative model that have been successfully applied to a diverse range of problems including image and video generation, protein and material synthesis, weather forecasting, and neural surrogates of partial differential equations. Despite their ubiquity it is hard to find an introduction to DDPMs which is simple, comprehensive, clean and clear. The compact explanations necessary in research papers are not able to elucidate all of the different design steps taken to formulate the DDPM and the rationale of the steps that are presented is often omitted to save space. Moreover, the expositions are typically presented from the variational lower bound perspective which is unnecessary and arguably harmful as it obfuscates why the method is working and suggests generalisations that do not perform well in practice. On the other hand, perspectives that take the continuous time-limit are beautiful and general, but they have a high barrier-to-entry as they require background knowledge of stochastic differential equations and probability flow. In this note, we distill down the formulation of the DDPM into six simple steps each of which comes with a clear rationale. We assume that the reader is familiar with fundamental topics in machine learning including basic probabilistic modelling, Gaussian distributions, maximum likelihood estimation, and deep learning.
Meta- (out-of-context) learning in neural networks
Oct 24, 2023Dmitrii Krasheninnikov, Egor Krasheninnikov, Bruno Mlodozeniec, David Krueger
Brown et al. (2020) famously introduced the phenomenon of in-context learning in large language models (LLMs). We establish the existence of a phenomenon we call meta-out-of-context learning (meta-OCL) via carefully designed synthetic experiments with LLMs. Our results suggest that meta-OCL leads LLMs to more readily "internalize" the semantic content of text that is, or appears to be, broadly useful (such as true statements, or text from authoritative sources) and use it in appropriate circumstances. We further demonstrate meta-OCL in a synthetic computer vision setting, and propose two hypotheses for the emergence of meta-OCL: one relying on the way models store knowledge in their parameters, and another suggesting that the implicit gradient alignment bias of gradient-descent-based optimizers may be responsible. Finally, we reflect on what our results might imply about capabilities of future AI systems, and discuss potential risks. Our code can be found at https://github.com/krasheninnikov/internalization.
Hyperparameter Optimization through Neural Network Partitioning
Apr 28, 2023Bruno Mlodozeniec, Matthias Reisser, Christos Louizos
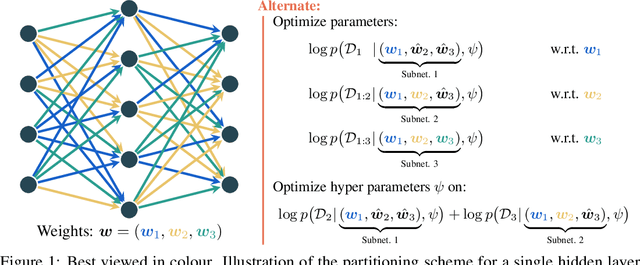
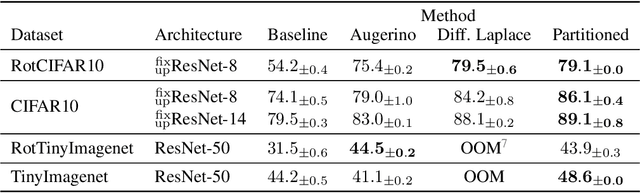
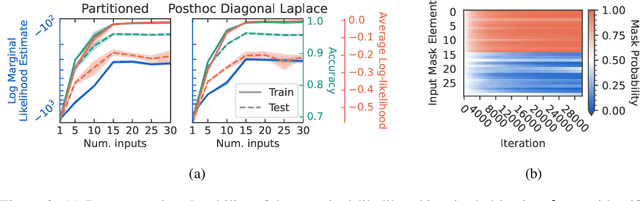
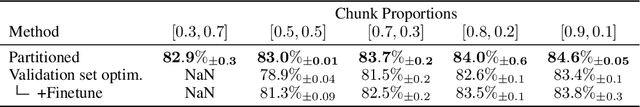
Well-tuned hyperparameters are crucial for obtaining good generalization behavior in neural networks. They can enforce appropriate inductive biases, regularize the model and improve performance -- especially in the presence of limited data. In this work, we propose a simple and efficient way for optimizing hyperparameters inspired by the marginal likelihood, an optimization objective that requires no validation data. Our method partitions the training data and a neural network model into $K$ data shards and parameter partitions, respectively. Each partition is associated with and optimized only on specific data shards. Combining these partitions into subnetworks allows us to define the ``out-of-training-sample" loss of a subnetwork, i.e., the loss on data shards unseen by the subnetwork, as the objective for hyperparameter optimization. We demonstrate that we can apply this objective to optimize a variety of different hyperparameters in a single training run while being significantly computationally cheaper than alternative methods aiming to optimize the marginal likelihood for neural networks. Lastly, we also focus on optimizing hyperparameters in federated learning, where retraining and cross-validation are particularly challenging.
Timewarp: Transferable Acceleration of Molecular Dynamics by Learning Time-Coarsened Dynamics
Feb 02, 2023Leon Klein, Andrew Y. K. Foong, Tor Erlend Fjelde, Bruno Mlodozeniec, Marc Brockschmidt, Sebastian Nowozin, Frank Noé, Ryota Tomioka
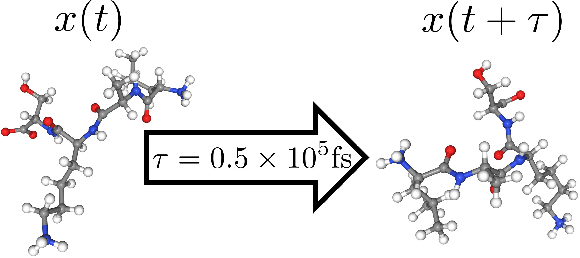
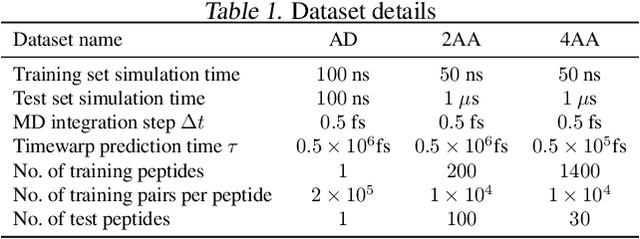
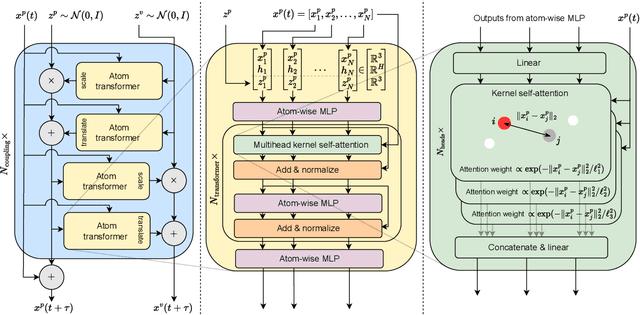

Molecular dynamics (MD) simulation is a widely used technique to simulate molecular systems, most commonly at the all-atom resolution where the equations of motion are integrated with timesteps on the order of femtoseconds ($1\textrm{fs}=10^{-15}\textrm{s}$). MD is often used to compute equilibrium properties, which requires sampling from an equilibrium distribution such as the Boltzmann distribution. However, many important processes, such as binding and folding, occur over timescales of milliseconds or beyond, and cannot be efficiently sampled with conventional MD. Furthermore, new MD simulations need to be performed from scratch for each molecular system studied. We present Timewarp, an enhanced sampling method which uses a normalising flow as a proposal distribution in a Markov chain Monte Carlo method targeting the Boltzmann distribution. The flow is trained offline on MD trajectories and learns to make large steps in time, simulating the molecular dynamics of $10^{5} - 10^{6}\:\textrm{fs}$. Crucially, Timewarp is transferable between molecular systems: once trained, we show that it generalises to unseen small peptides (2-4 amino acids), exploring their metastable states and providing wall-clock acceleration when sampling compared to standard MD. Our method constitutes an important step towards developing general, transferable algorithms for accelerating MD.
Ensemble Distribution Distillation
May 30, 2019Andrey Malinin, Bruno Mlodozeniec, Mark Gales




Ensembles of models often yield improvements in system performance. They have also been empirically shown to yield robust measures of uncertainty, and are capable of distinguishing between different forms of uncertainty. However, ensembles come at a computational and memory cost which may be prohibitive for many applications. There has been significant work done on the distillation of an ensemble into a single model. Such approaches decrease computational cost and allow a single model to achieve an accuracy comparable to that of an ensemble. However, information about the diversity of the ensemble, which can yield estimates of different forms of uncertainty, is lost. Recently, a new class of models called Prior Networks has been proposed, which allows a single neural network to explicitly model a distribution over output distributions. In this work, Ensemble Distillation and Prior Networks are combined to yield a novel approach called Ensemble Distribution Distillation (EnD$^2$), in which the distribution of an ensemble is distilled into a single Prior Network. This enables a single model to retain both the improved classification performance as well as measures of diversity of the ensemble. The properties of EnD$^2$ have been investigated on both an artificial dataset, and on the CIFAR-10 dataset, where it is shown that EnD$^2$ can approach the performance of an ensemble, and outperforms both standard DNNs and standard Ensemble Distillation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge