Inductive-Deductive Strategy Reuse for Multi-Turn Instructional Dialogues
Apr 17, 2024Jiao Ou, Jiayu Wu, Che Liu, Fuzheng Zhang, Di Zhang, Kun Gai
Aligning large language models (LLMs) with human expectations requires high-quality instructional dialogues, which can be achieved by raising diverse, in-depth, and insightful instructions that deepen interactions. Existing methods target instructions from real instruction dialogues as a learning goal and fine-tune a user simulator for posing instructions. However, the user simulator struggles to implicitly model complex dialogue flows and pose high-quality instructions. In this paper, we take inspiration from the cognitive abilities inherent in human learning and propose the explicit modeling of complex dialogue flows through instructional strategy reuse. Specifically, we first induce high-level strategies from various real instruction dialogues. These strategies are applied to new dialogue scenarios deductively, where the instructional strategies facilitate high-quality instructions. Experimental results show that our method can generate diverse, in-depth, and insightful instructions for a given dialogue history. The constructed multi-turn instructional dialogues can outperform competitive baselines on the downstream chat model.
BIMCV-R: A Landmark Dataset for 3D CT Text-Image Retrieval
Mar 24, 2024Yinda Chen, Che Liu, Xiaoyu Liu, Rossella Arcucci, Zhiwei Xiong
The burgeoning integration of 3D medical imaging into healthcare has led to a substantial increase in the workload of medical professionals. To assist clinicians in their diagnostic processes and alleviate their workload, the development of a robust system for retrieving similar case studies presents a viable solution. While the concept holds great promise, the field of 3D medical text-image retrieval is currently limited by the absence of robust evaluation benchmarks and curated datasets. To remedy this, our study presents a groundbreaking dataset, BIMCV-R (This dataset will be released upon acceptance.), which includes an extensive collection of 8,069 3D CT volumes, encompassing over 2 million slices, paired with their respective radiological reports. Expanding upon the foundational work of our dataset, we craft a retrieval strategy, MedFinder. This approach employs a dual-stream network architecture, harnessing the potential of large language models to advance the field of medical image retrieval beyond existing text-image retrieval solutions. It marks our preliminary step towards developing a system capable of facilitating text-to-image, image-to-text, and keyword-based retrieval tasks.
Electrocardiogram Instruction Tuning for Report Generation
Mar 13, 2024Zhongwei Wan, Che Liu, Xin Wang, Chaofan Tao, Hui Shen, Zhenwu Peng, Jie Fu, Rossella Arcucci, Huaxiu Yao, Mi Zhang
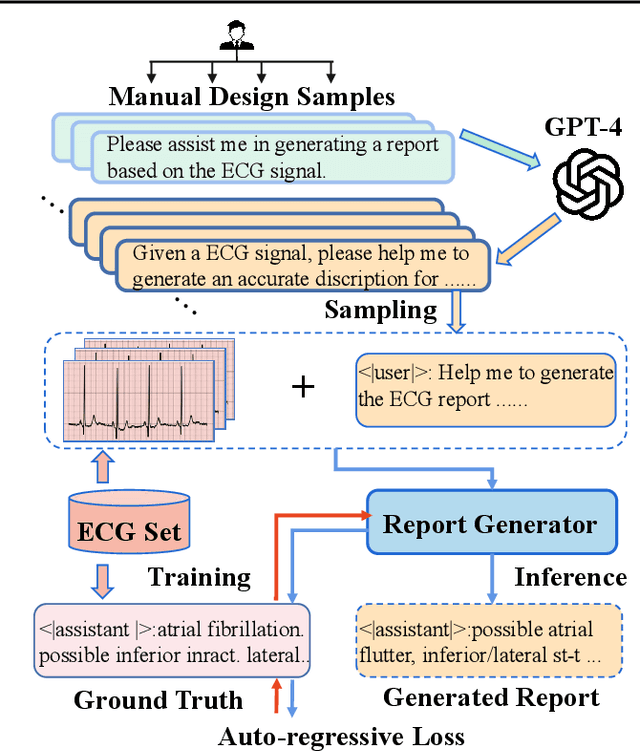
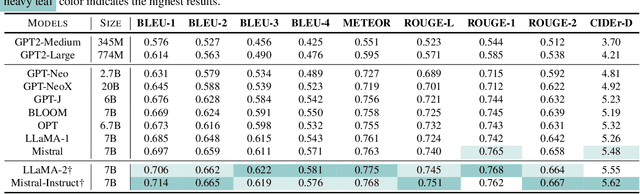
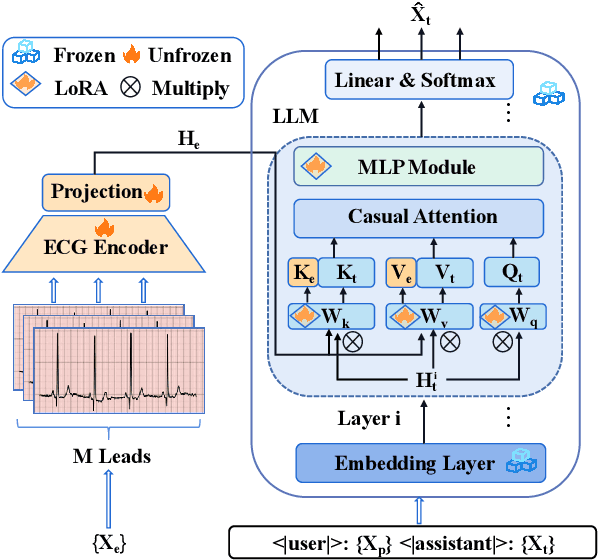
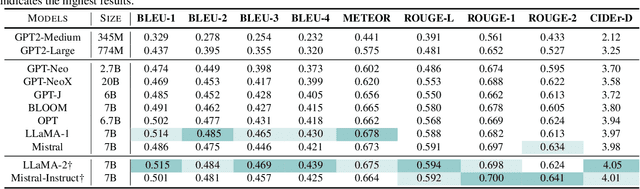
Electrocardiogram (ECG) serves as the primary non-invasive diagnostic tool for cardiac conditions monitoring, are crucial in assisting clinicians. Recent studies have concentrated on classifying cardiac conditions using ECG data but have overlooked ECG report generation, which is not only time-consuming but also requires clinical expertise. To automate ECG report generation and ensure its versatility, we propose the Multimodal ECG Instruction Tuning (MEIT) framework, the \textit{first} attempt to tackle ECG report generation with LLMs and multimodal instructions. To facilitate future research, we establish a benchmark to evaluate MEIT with various LLMs backbones across two large-scale ECG datasets. Our approach uniquely aligns the representations of the ECG signal and the report, and we conduct extensive experiments to benchmark MEIT with nine open source LLMs, using more than 800,000 ECG reports. MEIT's results underscore the superior performance of instruction-tuned LLMs, showcasing their proficiency in quality report generation, zero-shot capabilities, and resilience to signal perturbation. These findings emphasize the efficacy of our MEIT framework and its potential for real-world clinical application.
Zero-Shot ECG Classification with Multimodal Learning and Test-time Clinical Knowledge Enhancement
Mar 11, 2024Che Liu, Zhongwei Wan, Cheng Ouyang, Anand Shah, Wenjia Bai, Rossella Arcucci
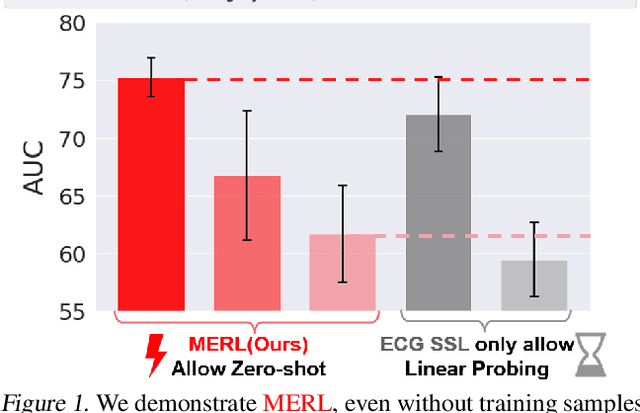

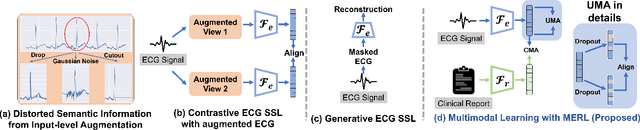
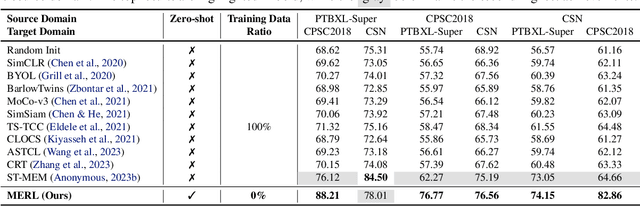
Electrocardiograms (ECGs) are non-invasive diagnostic tools crucial for detecting cardiac arrhythmic diseases in clinical practice. While ECG Self-supervised Learning (eSSL) methods show promise in representation learning from unannotated ECG data, they often overlook the clinical knowledge that can be found in reports. This oversight and the requirement for annotated samples for downstream tasks limit eSSL's versatility. In this work, we address these issues with the Multimodal ECG Representation Learning (MERL}) framework. Through multimodal learning on ECG records and associated reports, MERL is capable of performing zero-shot ECG classification with text prompts, eliminating the need for training data in downstream tasks. At test time, we propose the Clinical Knowledge Enhanced Prompt Engineering (CKEPE) approach, which uses Large Language Models (LLMs) to exploit external expert-verified clinical knowledge databases, generating more descriptive prompts and reducing hallucinations in LLM-generated content to boost zero-shot classification. Based on MERL, we perform the first benchmark across six public ECG datasets, showing the superior performance of MERL compared against eSSL methods. Notably, MERL achieves an average AUC score of 75.2% in zero-shot classification (without training data), 3.2% higher than linear probed eSSL methods with 10\% annotated training data, averaged across all six datasets.
ARKS: Active Retrieval in Knowledge Soup for Code Generation
Feb 19, 2024Hongjin Su, Shuyang Jiang, Yuhang Lai, Haoyuan Wu, Boao Shi, Che Liu, Qian Liu, Tao Yu
Recently the retrieval-augmented generation (RAG) paradigm has raised much attention for its potential in incorporating external knowledge into large language models (LLMs) without further training. While widely explored in natural language applications, its utilization in code generation remains under-explored. In this paper, we introduce Active Retrieval in Knowledge Soup (ARKS), an advanced strategy for generalizing large language models for code. In contrast to relying on a single source, we construct a knowledge soup integrating web search, documentation, execution feedback, and evolved code snippets. We employ an active retrieval strategy that iteratively refines the query and updates the knowledge soup. To assess the performance of ARKS, we compile a new benchmark comprising realistic coding problems associated with frequently updated libraries and long-tail programming languages. Experimental results on ChatGPT and CodeLlama demonstrate a substantial improvement in the average execution accuracy of ARKS on LLMs. The analysis confirms the effectiveness of our proposed knowledge soup and active retrieval strategies, offering rich insights into the construction of effective retrieval-augmented code generation (RACG) pipelines. Our model, code, and data are available at https://arks-codegen.github.io.
Enhancing Role-playing Systems through Aggressive Queries: Evaluation and Improvement
Feb 16, 2024Yihong Tang, Jiao Ou, Che Liu, Fuzheng Zhang, Di Zhang, Kun Gai
The advent of Large Language Models (LLMs) has propelled dialogue generation into new realms, particularly in the field of role-playing systems (RPSs). While enhanced with ordinary role-relevant training dialogues, existing LLM-based RPSs still struggle to align with roles when handling intricate and trapped queries in boundary scenarios. In this paper, we design the Modular ORchestrated Trap-setting Interaction SystEm (MORTISE) to benchmark and improve the role-playing LLMs' performance. MORTISE can produce highly role-relevant aggressive queries through the collaborative effort of multiple LLM-based modules, and formulate corresponding responses to create an adversarial training dataset via a consistent response generator. We select 190 Chinese and English roles to construct aggressive queries to benchmark existing role-playing LLMs. Through comprehensive evaluation, we find that existing models exhibit a general deficiency in role alignment capabilities. We further select 180 of the roles to collect an adversarial training dataset (named RoleAD) and retain the other 10 roles for testing. Experiments on models improved by RoleAD indicate that our adversarial dataset ameliorates this deficiency, with the improvements demonstrating a degree of generalizability in ordinary scenarios.
Freeze the backbones: A Parameter-Efficient Contrastive Approach to Robust Medical Vision-Language Pre-training
Jan 02, 2024Jiuming Qin, Che Liu, Sibo Cheng, Yike Guo, Rossella Arcucci
Modern healthcare often utilises radiographic images alongside textual reports for diagnostics, encouraging the use of Vision-Language Self-Supervised Learning (VL-SSL) with large pre-trained models to learn versatile medical vision representations. However, most existing VL-SSL frameworks are trained end-to-end, which is computation-heavy and can lose vital prior information embedded in pre-trained encoders. To address both issues, we introduce the backbone-agnostic Adaptor framework, which preserves medical knowledge in pre-trained image and text encoders by keeping them frozen, and employs a lightweight Adaptor module for cross-modal learning. Experiments on medical image classification and segmentation tasks across three datasets reveal that our framework delivers competitive performance while cutting trainable parameters by over 90% compared to current pre-training approaches. Notably, when fine-tuned with just 1% of data, Adaptor outperforms several Transformer-based methods trained on full datasets in medical image segmentation.
Efficient Large Language Models: A Survey
Dec 23, 2023Zhongwei Wan, Xin Wang, Che Liu, Samiul Alam, Yu Zheng, Jiachen Liu, Zhongnan Qu, Shen Yan, Yi Zhu, Quanlu Zhang, Mosharaf Chowdhury, Mi Zhang
Large Language Models (LLMs) have demonstrated remarkable capabilities in important tasks such as natural language understanding, language generation, and complex reasoning and have the potential to make a substantial impact on our society. Such capabilities, however, come with the considerable resources they demand, highlighting the strong need to develop effective techniques for addressing their efficiency challenges. In this survey, we provide a systematic and comprehensive review of efficient LLMs research. We organize the literature in a taxonomy consisting of three main categories, covering distinct yet interconnected efficient LLMs topics from model-centric, data-centric, and framework-centric perspective, respectively. We have also created a GitHub repository where we compile the papers featured in this survey at https://github.com/AIoT-MLSys-Lab/EfficientLLMs, and will actively maintain this repository and incorporate new research as it emerges. We hope our survey can serve as a valuable resource to help researchers and practitioners gain a systematic understanding of the research developments in efficient LLMs and inspire them to contribute to this important and exciting field.
T3D: Towards 3D Medical Image Understanding through Vision-Language Pre-training
Dec 05, 2023Che Liu, Cheng Ouyang, Yinda Chen, Cesar César Quilodrán-Casas, Lei Ma, Jie Fu, Yike Guo, Anand Shah, Wenjia Bai, Rossella Arcucci
Expert annotation of 3D medical image for downstream analysis is resource-intensive, posing challenges in clinical applications. Visual self-supervised learning (vSSL), though effective for learning visual invariance, neglects the incorporation of domain knowledge from medicine. To incorporate medical knowledge into visual representation learning, vision-language pre-training (VLP) has shown promising results in 2D image. However, existing VLP approaches become generally impractical when applied to high-resolution 3D medical images due to GPU hardware constraints and the potential loss of critical details caused by downsampling, which is the intuitive solution to hardware constraints. To address the above limitations, we introduce T3D, the first VLP framework designed for high-resolution 3D medical images. T3D incorporates two text-informed pretext tasks: (\lowerromannumeral{1}) text-informed contrastive learning; (\lowerromannumeral{2}) text-informed image restoration. These tasks focus on learning 3D visual representations from high-resolution 3D medical images and integrating clinical knowledge from radiology reports, without distorting information through forced alignment of downsampled volumes with detailed anatomical text. Trained on a newly curated large-scale dataset of 3D medical images and radiology reports, T3D significantly outperforms current vSSL methods in tasks like organ and tumor segmentation, as well as disease classification. This underlines T3D's potential in representation learning for 3D medical image analysis. All data and code will be available upon acceptance.
G2D: From Global to Dense Radiography Representation Learning via Vision-Language Pre-training
Dec 03, 2023Che Liu, Cheng Ouyang, Sibo Cheng, Anand Shah, Wenjia Bai, Rossella Arcucci
Recently, medical vision-language pre-training (VLP) has reached substantial progress to learn global visual representation from medical images and their paired radiology reports. However, medical imaging tasks in real world usually require finer granularity in visual features. These tasks include visual localization tasks (e.g., semantic segmentation, object detection) and visual grounding task. Yet, current medical VLP methods face challenges in learning these fine-grained features, as they primarily focus on brute-force alignment between image patches and individual text tokens for local visual feature learning, which is suboptimal for downstream dense prediction tasks. In this work, we propose a new VLP framework, named \textbf{G}lobal to \textbf{D}ense level representation learning (G2D) that achieves significantly improved granularity and more accurate grounding for the learned features, compared to existing medical VLP approaches. In particular, G2D learns dense and semantically-grounded image representations via a pseudo segmentation task parallel with the global vision-language alignment. Notably, generating pseudo segmentation targets does not incur extra trainable parameters: they are obtained on the fly during VLP with a parameter-free processor. G2D achieves superior performance across 6 medical imaging tasks and 25 diseases, particularly in semantic segmentation, which necessitates fine-grained, semantically-grounded image features. In this task, G2D surpasses peer models even when fine-tuned with just 1\% of the training data, compared to the 100\% used by these models. The code will be released upon acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge