PAM-UNet: Shifting Attention on Region of Interest in Medical Images
May 02, 2024Abhijit Das, Debesh Jha, Vandan Gorade, Koushik Biswas, Hongyi Pan, Zheyuan Zhang, Daniela P. Ladner, Yury Velichko, Amir Borhani, Ulas Bagci
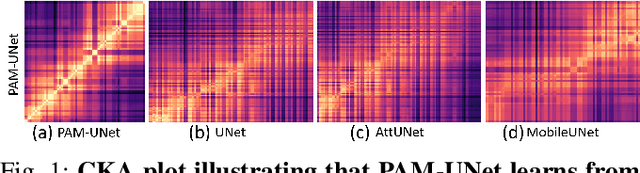
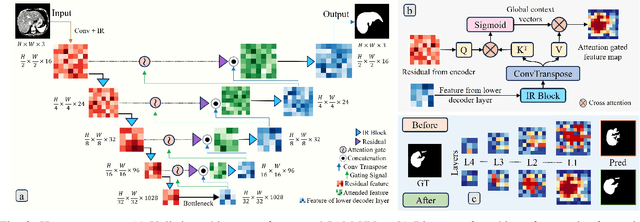
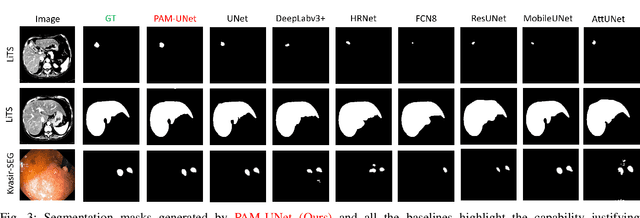

Computer-aided segmentation methods can assist medical personnel in improving diagnostic outcomes. While recent advancements like UNet and its variants have shown promise, they face a critical challenge: balancing accuracy with computational efficiency. Shallow encoder architectures in UNets often struggle to capture crucial spatial features, leading in inaccurate and sparse segmentation. To address this limitation, we propose a novel \underline{P}rogressive \underline{A}ttention based \underline{M}obile \underline{UNet} (\underline{PAM-UNet}) architecture. The inverted residual (IR) blocks in PAM-UNet help maintain a lightweight framework, while layerwise \textit{Progressive Luong Attention} ($\mathcal{PLA}$) promotes precise segmentation by directing attention toward regions of interest during synthesis. Our approach prioritizes both accuracy and speed, achieving a commendable balance with a mean IoU of 74.65 and a dice score of 82.87, while requiring only 1.32 floating-point operations per second (FLOPS) on the Liver Tumor Segmentation Benchmark (LiTS) 2017 dataset. These results highlight the importance of developing efficient segmentation models to accelerate the adoption of AI in clinical practice.
Detection of Peri-Pancreatic Edema using Deep Learning and Radiomics Techniques
Apr 25, 2024Ziliang Hong, Debesh Jha, Koushik Biswas, Zheyuan Zhang, Yury Velichko, Cemal Yazici, Temel Tirkes, Amir Borhani, Baris Turkbey, Alpay Medetalibeyoglu, Gorkem Durak, Ulas Bagci
Identifying peri-pancreatic edema is a pivotal indicator for identifying disease progression and prognosis, emphasizing the critical need for accurate detection and assessment in pancreatitis diagnosis and management. This study \textit{introduces a novel CT dataset sourced from 255 patients with pancreatic diseases, featuring annotated pancreas segmentation masks and corresponding diagnostic labels for peri-pancreatic edema condition}. With the novel dataset, we first evaluate the efficacy of the \textit{LinTransUNet} model, a linear Transformer based segmentation algorithm, to segment the pancreas accurately from CT imaging data. Then, we use segmented pancreas regions with two distinctive machine learning classifiers to identify existence of peri-pancreatic edema: deep learning-based models and a radiomics-based eXtreme Gradient Boosting (XGBoost). The LinTransUNet achieved promising results, with a dice coefficient of 80.85\%, and mIoU of 68.73\%. Among the nine benchmarked classification models for peri-pancreatic edema detection, \textit{Swin-Tiny} transformer model demonstrated the highest recall of $98.85 \pm 0.42$ and precision of $98.38\pm 0.17$. Comparatively, the radiomics-based XGBoost model achieved an accuracy of $79.61\pm4.04$ and recall of $91.05\pm3.28$, showcasing its potential as a supplementary diagnostic tool given its rapid processing speed and reduced training time. Our code is available \url{https://github.com/NUBagciLab/Peri-Pancreatic-Edema-Detection}.
Explainable Transformer Prototypes for Medical Diagnoses
Mar 11, 2024Ugur Demir, Debesh Jha, Zheyuan Zhang, Elif Keles, Bradley Allen, Aggelos K. Katsaggelos, Ulas Bagci
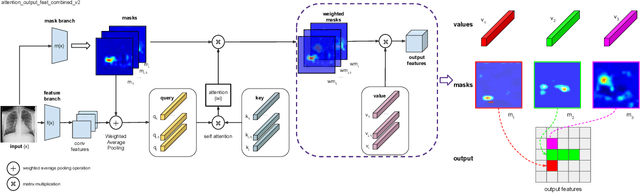
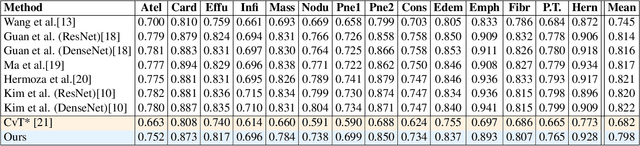
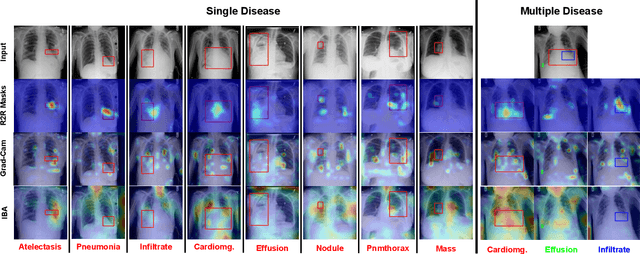
Deployments of artificial intelligence in medical diagnostics mandate not just accuracy and efficacy but also trust, emphasizing the need for explainability in machine decisions. The recent trend in automated medical image diagnostics leans towards the deployment of Transformer-based architectures, credited to their impressive capabilities. Since the self-attention feature of transformers contributes towards identifying crucial regions during the classification process, they enhance the trustability of the methods. However, the complex intricacies of these attention mechanisms may fall short of effectively pinpointing the regions of interest directly influencing AI decisions. Our research endeavors to innovate a unique attention block that underscores the correlation between 'regions' rather than 'pixels'. To address this challenge, we introduce an innovative system grounded in prototype learning, featuring an advanced self-attention mechanism that goes beyond conventional ad-hoc visual explanation techniques by offering comprehensible visual insights. A combined quantitative and qualitative methodological approach was used to demonstrate the effectiveness of the proposed method on the large-scale NIH chest X-ray dataset. Experimental results showed that our proposed method offers a promising direction for explainability, which can lead to the development of more trustable systems, which can facilitate easier and rapid adoption of such technology into routine clinics. The code is available at www.github.com/NUBagcilab/r2r_proto.
Harmonized Spatial and Spectral Learning for Robust and Generalized Medical Image Segmentation
Jan 18, 2024Vandan Gorade, Sparsh Mittal, Debesh Jha, Rekha Singhal, Ulas Bagci
Deep learning has demonstrated remarkable achievements in medical image segmentation. However, prevailing deep learning models struggle with poor generalization due to (i) intra-class variations, where the same class appears differently in different samples, and (ii) inter-class independence, resulting in difficulties capturing intricate relationships between distinct objects, leading to higher false negative cases. This paper presents a novel approach that synergies spatial and spectral representations to enhance domain-generalized medical image segmentation. We introduce the innovative Spectral Correlation Coefficient objective to improve the model's capacity to capture middle-order features and contextual long-range dependencies. This objective complements traditional spatial objectives by incorporating valuable spectral information. Extensive experiments reveal that optimizing this objective with existing architectures like UNet and TransUNet significantly enhances generalization, interpretability, and noise robustness, producing more confident predictions. For instance, in cardiac segmentation, we observe a 0.81 pp and 1.63 pp (pp = percentage point) improvement in DSC over UNet and TransUNet, respectively. Our interpretability study demonstrates that, in most tasks, objectives optimized with UNet outperform even TransUNet by introducing global contextual information alongside local details. These findings underscore the versatility and effectiveness of our proposed method across diverse imaging modalities and medical domains.
CT Liver Segmentation via PVT-based Encoding and Refined Decoding
Jan 17, 2024Debesh Jha, Nikhil Kumar Tomar, Koushik Biswas, Gorkem Durak, Alpay Medetalibeyoglu, Matthew Antalek, Yury Velichko, Daniela Ladner, Amir Borhani, Ulas Bagci
Accurate liver segmentation from CT scans is essential for computer-aided diagnosis and treatment planning. Recently, Vision Transformers achieved a competitive performance in computer vision tasks compared to convolutional neural networks due to their exceptional ability to learn global representations. However, they often struggle with scalability, memory constraints, and computational inefficiency, particularly in handling high-resolution medical images. To overcome scalability and efficiency issues, we propose a novel deep learning approach, \textit{\textbf{PVTFormer}}, that is built upon a pretrained pyramid vision transformer (PVT v2) combined with advanced residual upsampling and decoder block. By integrating a refined feature channel approach with hierarchical decoding strategy, PVTFormer generates high quality segmentation masks by enhancing semantic features. Rigorous evaluation of the proposed method on Liver Tumor Segmentation Benchmark (LiTS) 2017 demonstrates that our proposed architecture not only achieves a high dice coefficient of 86.78\%, mIoU of 78.46\%, but also obtains a low HD of 3.50. The results underscore PVTFormer's efficacy in setting a new benchmark for state-of-the-art liver segmentation methods. The source code of the proposed PVTFormer is available at \url{https://github.com/DebeshJha/PVTFormer}.
Pose Guidance by Supervision: A Framework for Clothes-Changing Person Re-Identification
Dec 09, 2023Quoc-Huy Trinh, Nhat-Tan Bui, Phuoc-Thao Vo Thi, Hai-Dang Nguyen, Debesh Jha, Ulas Bagci, Minh-Triet Tran
Person Re-Identification (ReID) task seeks to enhance the tracking of multiple individuals by surveillance cameras. It provides additional support for multimodal tasks, including text-based person retrieval and human matching. One of the primary challenges in ReID is clothes-changing, which means the same person wears different clothes. While previous methods have achieved competitive results in maintaining clothing data consistency and handling clothing change data, they still tend to rely excessively on clothing information, thus limiting performance due to the dynamic nature of human appearances. To mitigate this challenge, we propose the Pose Guidance by Supervision (PGS) framework, an effective framework for learning pose guidance within the ReID task. This approach leverages pose knowledge and human part information from the pre-trained features to guide the network focus on clothes-irrelevant information, thus alleviating the clothes' influence on the deep learning model. Extensive experiments on five benchmark datasets demonstrate that our framework achieves competitive results compared with other state-of-the-art methods, which holds promise for developing robust models in the ReID task. Our code is available at https://github.com/huyquoctrinh/PGS.
Rethinking Intermediate Layers design in Knowledge Distillation for Kidney and Liver Tumor Segmentation
Nov 28, 2023Vandan Gorade, Sparsh Mittal, Debesh Jha, Ulas Bagci
Knowledge distillation(KD) has demonstrated remarkable success across various domains, but its application to medical imaging tasks, such as kidney and liver tumor segmentation, has encountered challenges. Many existing KD methods are not specifically tailored for these tasks. Moreover, prevalent KD methods often lack a careful consideration of what and from where to distill knowledge from the teacher to the student. This oversight may lead to issues like the accumulation of training bias within shallower student layers, potentially compromising the effectiveness of KD. To address these challenges, we propose Hierarchical Layer-selective Feedback Distillation (HLFD). HLFD strategically distills knowledge from a combination of middle layers to earlier layers and transfers final layer knowledge to intermediate layers at both the feature and pixel levels. This design allows the model to learn higher-quality representations from earlier layers, resulting in a robust and compact student model. Extensive quantitative evaluations reveal that HLFD outperforms existing methods by a significant margin. For example, in the kidney segmentation task, HLFD surpasses the student model (without KD) by over 10pp, significantly improving its focus on tumor-specific features. From a qualitative standpoint, the student model trained using HLFD excels at suppressing irrelevant information and can focus sharply on tumor-specific details, which opens a new pathway for more efficient and accurate diagnostic tools.
SynergyNet: Bridging the Gap between Discrete and Continuous Representations for Precise Medical Image Segmentation
Oct 26, 2023Vandan Gorade, Sparsh Mittal, Debesh Jha, Ulas Bagci
In recent years, continuous latent space (CLS) and discrete latent space (DLS) deep learning models have been proposed for medical image analysis for improved performance. However, these models encounter distinct challenges. CLS models capture intricate details but often lack interpretability in terms of structural representation and robustness due to their emphasis on low-level features. Conversely, DLS models offer interpretability, robustness, and the ability to capture coarse-grained information thanks to their structured latent space. However, DLS models have limited efficacy in capturing fine-grained details. To address the limitations of both DLS and CLS models, we propose SynergyNet, a novel bottleneck architecture designed to enhance existing encoder-decoder segmentation frameworks. SynergyNet seamlessly integrates discrete and continuous representations to harness complementary information and successfully preserves both fine and coarse-grained details in the learned representations. Our extensive experiment on multi-organ segmentation and cardiac datasets demonstrates that SynergyNet outperforms other state of the art methods, including TransUNet: dice scores improving by 2.16%, and Hausdorff scores improving by 11.13%, respectively. When evaluating skin lesion and brain tumor segmentation datasets, we observe a remarkable improvement of 1.71% in Intersection-over Union scores for skin lesion segmentation and of 8.58% for brain tumor segmentation. Our innovative approach paves the way for enhancing the overall performance and capabilities of deep learning models in the critical domain of medical image analysis.
EMIT-Diff: Enhancing Medical Image Segmentation via Text-Guided Diffusion Model
Oct 19, 2023Zheyuan Zhang, Lanhong Yao, Bin Wang, Debesh Jha, Elif Keles, Alpay Medetalibeyoglu, Ulas Bagci
Large-scale, big-variant, and high-quality data are crucial for developing robust and successful deep-learning models for medical applications since they potentially enable better generalization performance and avoid overfitting. However, the scarcity of high-quality labeled data always presents significant challenges. This paper proposes a novel approach to address this challenge by developing controllable diffusion models for medical image synthesis, called EMIT-Diff. We leverage recent diffusion probabilistic models to generate realistic and diverse synthetic medical image data that preserve the essential characteristics of the original medical images by incorporating edge information of objects to guide the synthesis process. In our approach, we ensure that the synthesized samples adhere to medically relevant constraints and preserve the underlying structure of imaging data. Due to the random sampling process by the diffusion model, we can generate an arbitrary number of synthetic images with diverse appearances. To validate the effectiveness of our proposed method, we conduct an extensive set of medical image segmentation experiments on multiple datasets, including Ultrasound breast (+13.87%), CT spleen (+0.38%), and MRI prostate (+7.78%), achieving significant improvements over the baseline segmentation methods. For the first time, to our best knowledge, the promising results demonstrate the effectiveness of our EMIT-Diff for medical image segmentation tasks and show the feasibility of introducing a first-ever text-guided diffusion model for general medical image segmentation tasks. With carefully designed ablation experiments, we investigate the influence of various data augmentation ratios, hyper-parameter settings, patch size for generating random merging mask settings, and combined influence with different network architectures.
Domain Generalization with Fourier Transform and Soft Thresholding
Sep 25, 2023Hongyi Pan, Bin Wang, Zheyuan Zhang, Xin Zhu, Debesh Jha, Ahmet Enis Cetin, Concetto Spampinato, Ulas Bagci
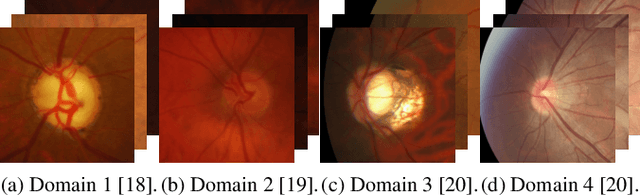
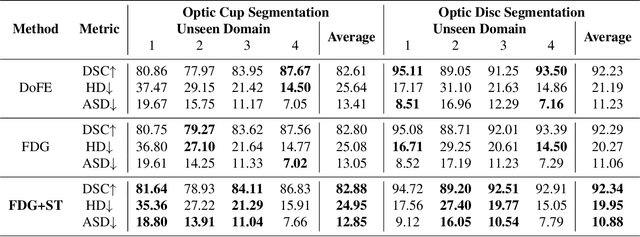
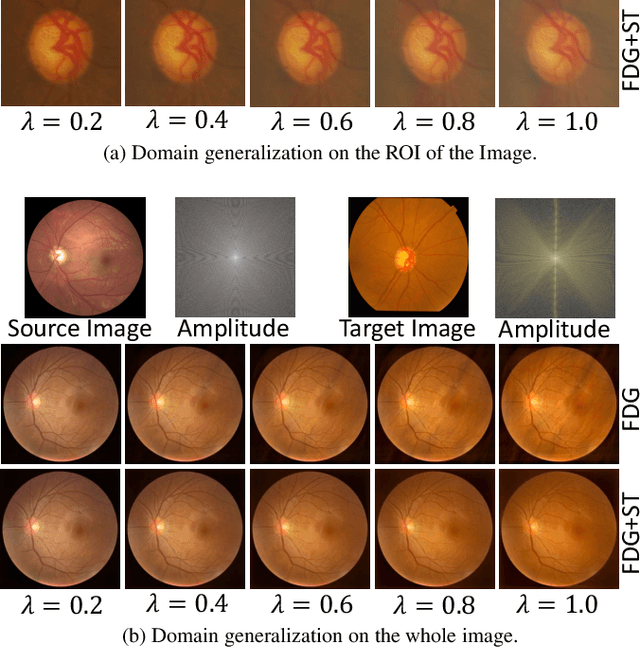
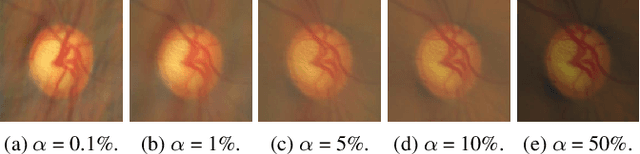
Domain generalization aims to train models on multiple source domains so that they can generalize well to unseen target domains. Among many domain generalization methods, Fourier-transform-based domain generalization methods have gained popularity primarily because they exploit the power of Fourier transformation to capture essential patterns and regularities in the data, making the model more robust to domain shifts. The mainstream Fourier-transform-based domain generalization swaps the Fourier amplitude spectrum while preserving the phase spectrum between the source and the target images. However, it neglects background interference in the amplitude spectrum. To overcome this limitation, we introduce a soft-thresholding function in the Fourier domain. We apply this newly designed algorithm to retinal fundus image segmentation, which is important for diagnosing ocular diseases but the neural network's performance can degrade across different sources due to domain shifts. The proposed technique basically enhances fundus image augmentation by eliminating small values in the Fourier domain and providing better generalization. The innovative nature of the soft thresholding fused with Fourier-transform-based domain generalization improves neural network models' performance by reducing the target images' background interference significantly. Experiments on public data validate our approach's effectiveness over conventional and state-of-the-art methods with superior segmentation metrics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge