HeAR -- Health Acoustic Representations
Mar 04, 2024Sebastien Baur, Zaid Nabulsi, Wei-Hung Weng, Jake Garrison, Louis Blankemeier, Sam Fishman, Christina Chen, Sujay Kakarmath, Minyoi Maimbolwa, Nsala Sanjase, Brian Shuma, Yossi Matias, Greg S. Corrado, Shwetak Patel, Shravya Shetty, Shruthi Prabhakara, Monde Muyoyeta, Diego Ardila
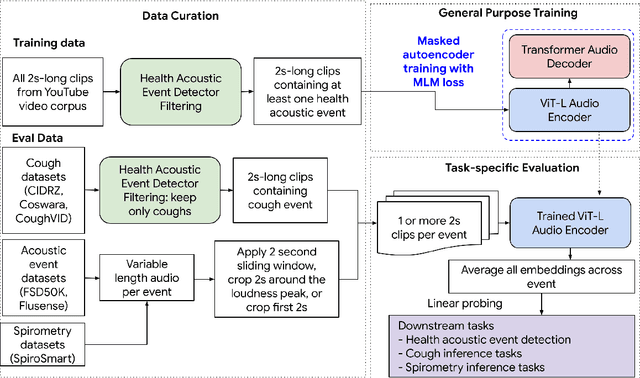
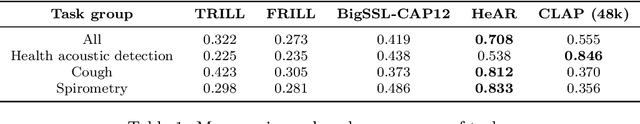
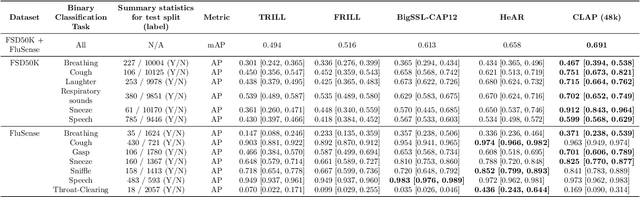
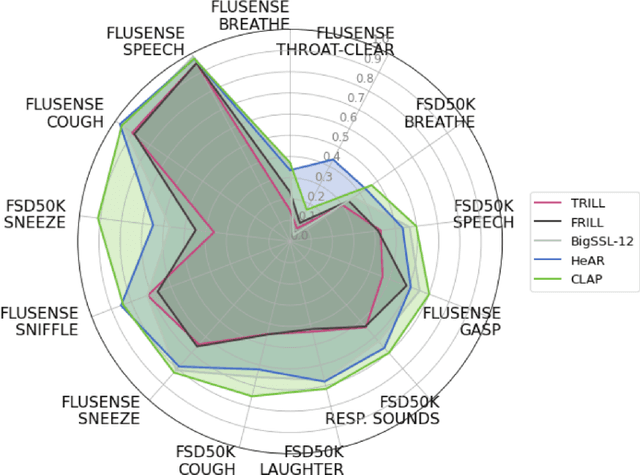
Health acoustic sounds such as coughs and breaths are known to contain useful health signals with significant potential for monitoring health and disease, yet are underexplored in the medical machine learning community. The existing deep learning systems for health acoustics are often narrowly trained and evaluated on a single task, which is limited by data and may hinder generalization to other tasks. To mitigate these gaps, we develop HeAR, a scalable self-supervised learning-based deep learning system using masked autoencoders trained on a large dataset of 313 million two-second long audio clips. Through linear probes, we establish HeAR as a state-of-the-art health audio embedding model on a benchmark of 33 health acoustic tasks across 6 datasets. By introducing this work, we hope to enable and accelerate further health acoustics research.
Closing the AI generalization gap by adjusting for dermatology condition distribution differences across clinical settings
Feb 23, 2024Rajeev V. Rikhye, Aaron Loh, Grace Eunhae Hong, Preeti Singh, Margaret Ann Smith, Vijaytha Muralidharan, Doris Wong, Rory Sayres, Michelle Phung, Nicolas Betancourt, Bradley Fong, Rachna Sahasrabudhe, Khoban Nasim, Alec Eschholz, Basil Mustafa, Jan Freyberg, Terry Spitz, Yossi Matias, Greg S. Corrado, Katherine Chou, Dale R. Webster, Peggy Bui, Yuan Liu, Yun Liu, Justin Ko, Steven Lin
Recently, there has been great progress in the ability of artificial intelligence (AI) algorithms to classify dermatological conditions from clinical photographs. However, little is known about the robustness of these algorithms in real-world settings where several factors can lead to a loss of generalizability. Understanding and overcoming these limitations will permit the development of generalizable AI that can aid in the diagnosis of skin conditions across a variety of clinical settings. In this retrospective study, we demonstrate that differences in skin condition distribution, rather than in demographics or image capture mode are the main source of errors when an AI algorithm is evaluated on data from a previously unseen source. We demonstrate a series of steps to close this generalization gap, requiring progressively more information about the new source, ranging from the condition distribution to training data enriched for data less frequently seen during training. Our results also suggest comparable performance from end-to-end fine tuning versus fine tuning solely the classification layer on top of a frozen embedding model. Our approach can inform the adaptation of AI algorithms to new settings, based on the information and resources available.
Domain-specific optimization and diverse evaluation of self-supervised models for histopathology
Oct 20, 2023Jeremy Lai, Faruk Ahmed, Supriya Vijay, Tiam Jaroensri, Jessica Loo, Saurabh Vyawahare, Saloni Agarwal, Fayaz Jamil, Yossi Matias, Greg S. Corrado, Dale R. Webster, Jonathan Krause, Yun Liu, Po-Hsuan Cameron Chen, Ellery Wulczyn, David F. Steiner
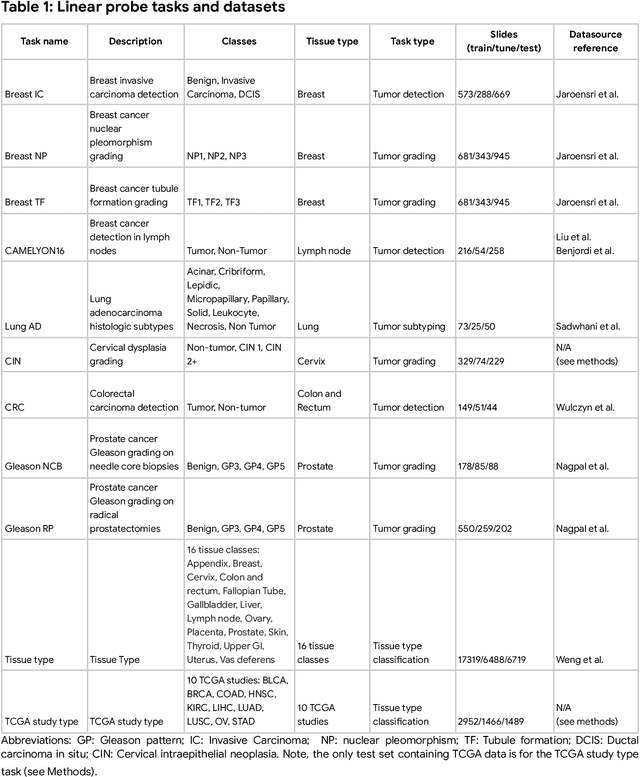
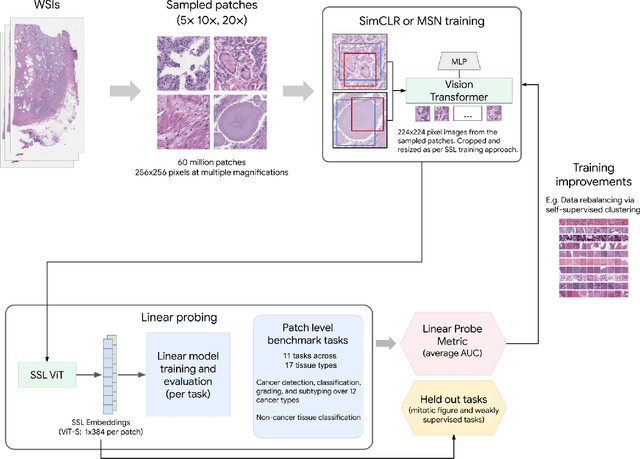
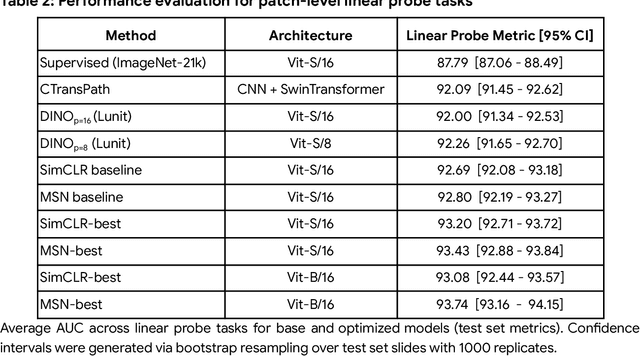
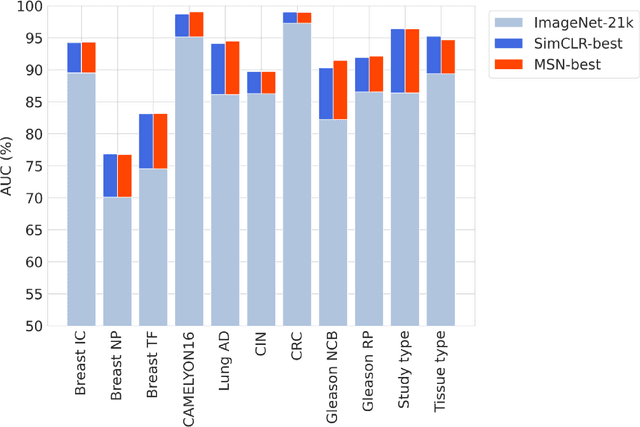
Task-specific deep learning models in histopathology offer promising opportunities for improving diagnosis, clinical research, and precision medicine. However, development of such models is often limited by availability of high-quality data. Foundation models in histopathology that learn general representations across a wide range of tissue types, diagnoses, and magnifications offer the potential to reduce the data, compute, and technical expertise necessary to develop task-specific deep learning models with the required level of model performance. In this work, we describe the development and evaluation of foundation models for histopathology via self-supervised learning (SSL). We first establish a diverse set of benchmark tasks involving 17 unique tissue types and 12 unique cancer types and spanning different optimal magnifications and task types. Next, we use this benchmark to explore and evaluate histopathology-specific SSL methods followed by further evaluation on held out patch-level and weakly supervised tasks. We found that standard SSL methods thoughtfully applied to histopathology images are performant across our benchmark tasks and that domain-specific methodological improvements can further increase performance. Our findings reinforce the value of using domain-specific SSL methods in pathology, and establish a set of high quality foundation models to enable further research across diverse applications.
ELIXR: Towards a general purpose X-ray artificial intelligence system through alignment of large language models and radiology vision encoders
Aug 02, 2023Shawn Xu, Lin Yang, Christopher Kelly, Marcin Sieniek, Timo Kohlberger, Martin Ma, Wei-Hung Weng, Attila Kiraly, Sahar Kazemzadeh, Zakkai Melamed, Jungyeon Park, Patricia Strachan, Yun Liu, Chuck Lau, Preeti Singh, Christina Chen, Mozziyar Etemadi, Sreenivasa Raju Kalidindi, Yossi Matias, Katherine Chou, Greg S. Corrado, Shravya Shetty, Daniel Tse, Shruthi Prabhakara, Daniel Golden, Rory Pilgrim, Krish Eswaran, Andrew Sellergren
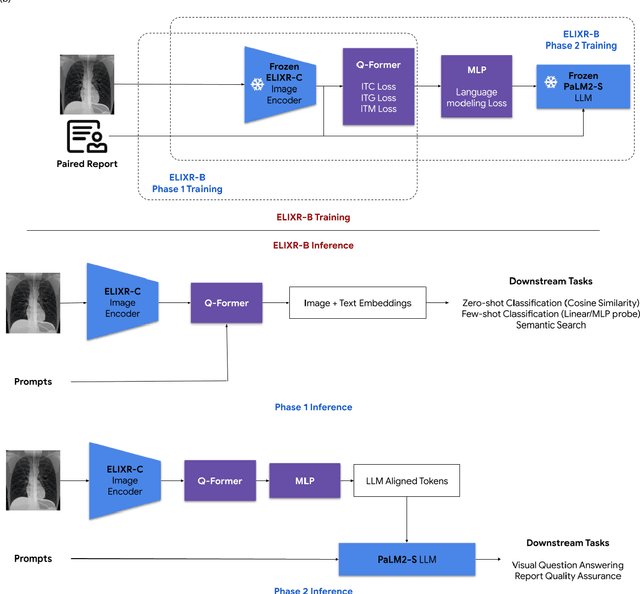
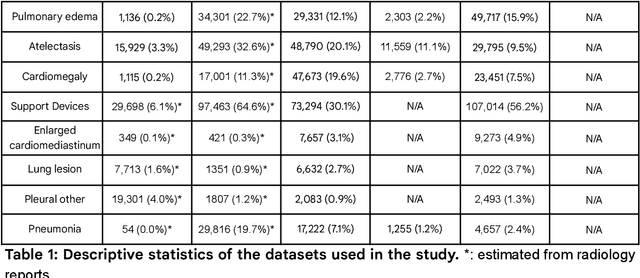
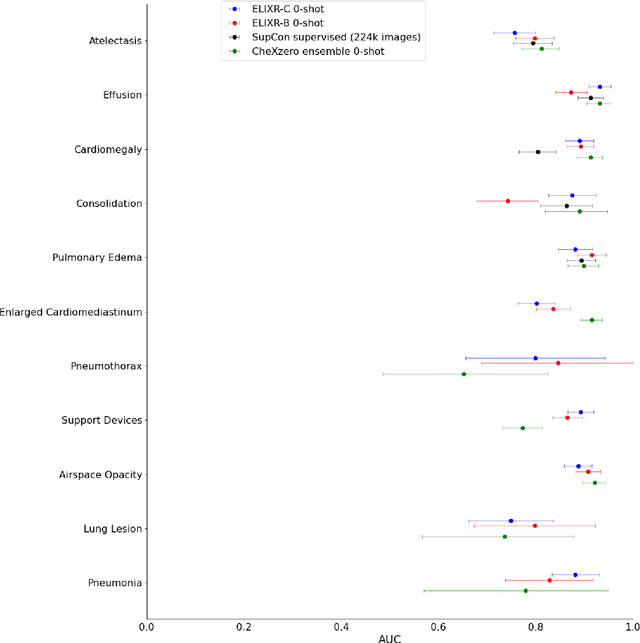
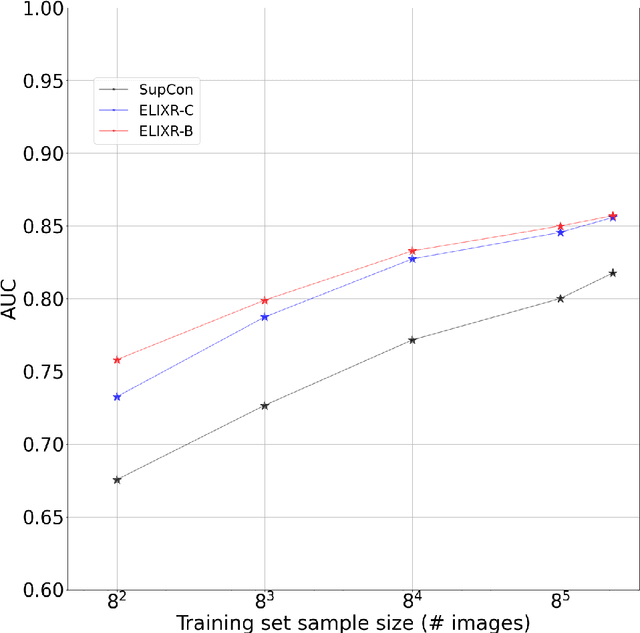
Our approach, which we call Embeddings for Language/Image-aligned X-Rays, or ELIXR, leverages a language-aligned image encoder combined or grafted onto a fixed LLM, PaLM 2, to perform a broad range of tasks. We train this lightweight adapter architecture using images paired with corresponding free-text radiology reports from the MIMIC-CXR dataset. ELIXR achieved state-of-the-art performance on zero-shot chest X-ray (CXR) classification (mean AUC of 0.850 across 13 findings), data-efficient CXR classification (mean AUCs of 0.893 and 0.898 across five findings (atelectasis, cardiomegaly, consolidation, pleural effusion, and pulmonary edema) for 1% (~2,200 images) and 10% (~22,000 images) training data), and semantic search (0.76 normalized discounted cumulative gain (NDCG) across nineteen queries, including perfect retrieval on twelve of them). Compared to existing data-efficient methods including supervised contrastive learning (SupCon), ELIXR required two orders of magnitude less data to reach similar performance. ELIXR also showed promise on CXR vision-language tasks, demonstrating overall accuracies of 58.7% and 62.5% on visual question answering and report quality assurance tasks, respectively. These results suggest that ELIXR is a robust and versatile approach to CXR AI.
Towards Generalist Biomedical AI
Jul 26, 2023Tao Tu, Shekoofeh Azizi, Danny Driess, Mike Schaekermann, Mohamed Amin, Pi-Chuan Chang, Andrew Carroll, Chuck Lau, Ryutaro Tanno, Ira Ktena, Basil Mustafa, Aakanksha Chowdhery, Yun Liu, Simon Kornblith, David Fleet, Philip Mansfield, Sushant Prakash, Renee Wong, Sunny Virmani, Christopher Semturs, S Sara Mahdavi, Bradley Green, Ewa Dominowska, Blaise Aguera y Arcas, Joelle Barral, Dale Webster, Greg S. Corrado, Yossi Matias, Karan Singhal, Pete Florence, Alan Karthikesalingam, Vivek Natarajan
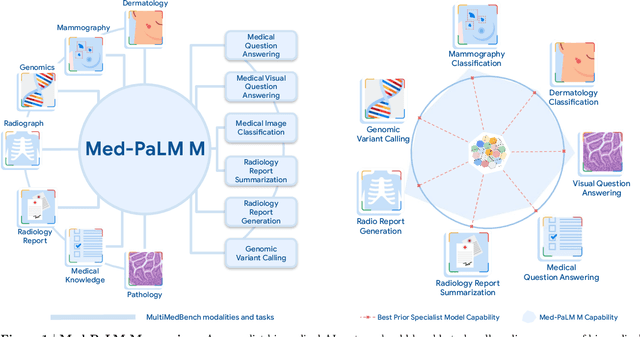
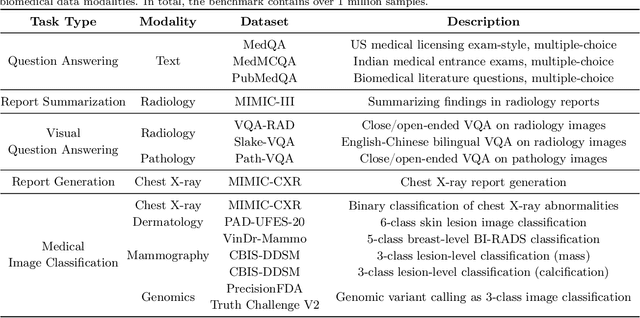

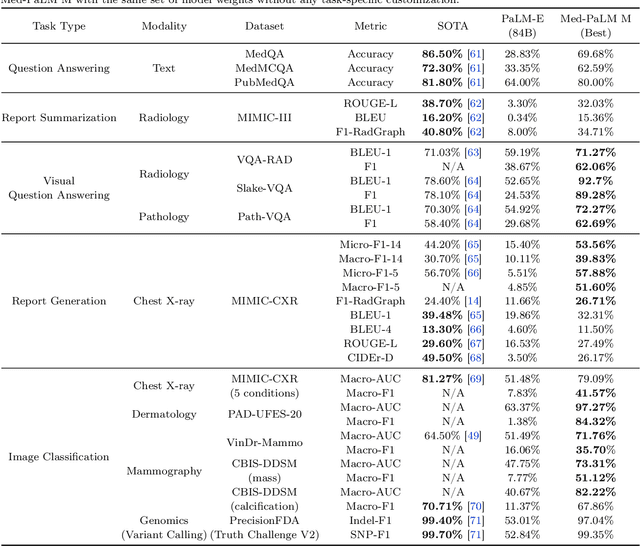
Medicine is inherently multimodal, with rich data modalities spanning text, imaging, genomics, and more. Generalist biomedical artificial intelligence (AI) systems that flexibly encode, integrate, and interpret this data at scale can potentially enable impactful applications ranging from scientific discovery to care delivery. To enable the development of these models, we first curate MultiMedBench, a new multimodal biomedical benchmark. MultiMedBench encompasses 14 diverse tasks such as medical question answering, mammography and dermatology image interpretation, radiology report generation and summarization, and genomic variant calling. We then introduce Med-PaLM Multimodal (Med-PaLM M), our proof of concept for a generalist biomedical AI system. Med-PaLM M is a large multimodal generative model that flexibly encodes and interprets biomedical data including clinical language, imaging, and genomics with the same set of model weights. Med-PaLM M reaches performance competitive with or exceeding the state of the art on all MultiMedBench tasks, often surpassing specialist models by a wide margin. We also report examples of zero-shot generalization to novel medical concepts and tasks, positive transfer learning across tasks, and emergent zero-shot medical reasoning. To further probe the capabilities and limitations of Med-PaLM M, we conduct a radiologist evaluation of model-generated (and human) chest X-ray reports and observe encouraging performance across model scales. In a side-by-side ranking on 246 retrospective chest X-rays, clinicians express a pairwise preference for Med-PaLM M reports over those produced by radiologists in up to 40.50% of cases, suggesting potential clinical utility. While considerable work is needed to validate these models in real-world use cases, our results represent a milestone towards the development of generalist biomedical AI systems.
Evaluating AI systems under uncertain ground truth: a case study in dermatology
Jul 05, 2023David Stutz, Ali Taylan Cemgil, Abhijit Guha Roy, Tatiana Matejovicova, Melih Barsbey, Patricia Strachan, Mike Schaekermann, Jan Freyberg, Rajeev Rikhye, Beverly Freeman, Javier Perez Matos, Umesh Telang, Dale R. Webster, Yuan Liu, Greg S. Corrado, Yossi Matias, Pushmeet Kohli, Yun Liu, Arnaud Doucet, Alan Karthikesalingam
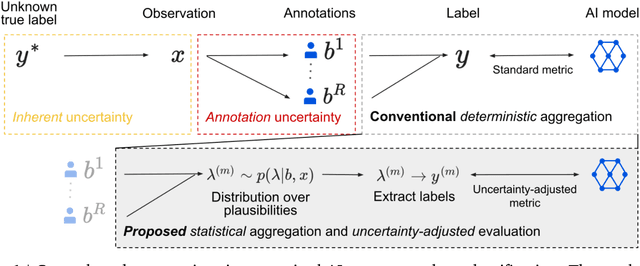
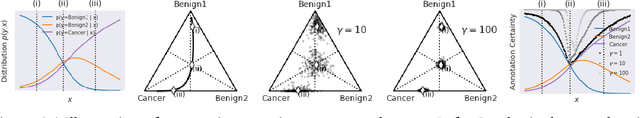
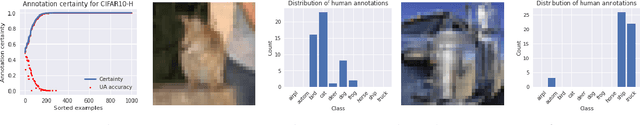
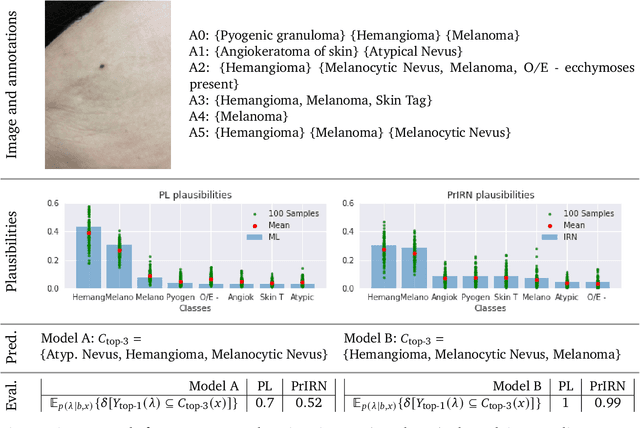
For safety, AI systems in health undergo thorough evaluations before deployment, validating their predictions against a ground truth that is assumed certain. However, this is actually not the case and the ground truth may be uncertain. Unfortunately, this is largely ignored in standard evaluation of AI models but can have severe consequences such as overestimating the future performance. To avoid this, we measure the effects of ground truth uncertainty, which we assume decomposes into two main components: annotation uncertainty which stems from the lack of reliable annotations, and inherent uncertainty due to limited observational information. This ground truth uncertainty is ignored when estimating the ground truth by deterministically aggregating annotations, e.g., by majority voting or averaging. In contrast, we propose a framework where aggregation is done using a statistical model. Specifically, we frame aggregation of annotations as posterior inference of so-called plausibilities, representing distributions over classes in a classification setting, subject to a hyper-parameter encoding annotator reliability. Based on this model, we propose a metric for measuring annotation uncertainty and provide uncertainty-adjusted metrics for performance evaluation. We present a case study applying our framework to skin condition classification from images where annotations are provided in the form of differential diagnoses. The deterministic adjudication process called inverse rank normalization (IRN) from previous work ignores ground truth uncertainty in evaluation. Instead, we present two alternative statistical models: a probabilistic version of IRN and a Plackett-Luce-based model. We find that a large portion of the dataset exhibits significant ground truth uncertainty and standard IRN-based evaluation severely over-estimates performance without providing uncertainty estimates.
Using generative AI to investigate medical imagery models and datasets
Jun 01, 2023Oran Lang, Doron Yaya-Stupp, Ilana Traynis, Heather Cole-Lewis, Chloe R. Bennett, Courtney Lyles, Charles Lau, Christopher Semturs, Dale R. Webster, Greg S. Corrado, Avinatan Hassidim, Yossi Matias, Yun Liu, Naama Hammel, Boris Babenko
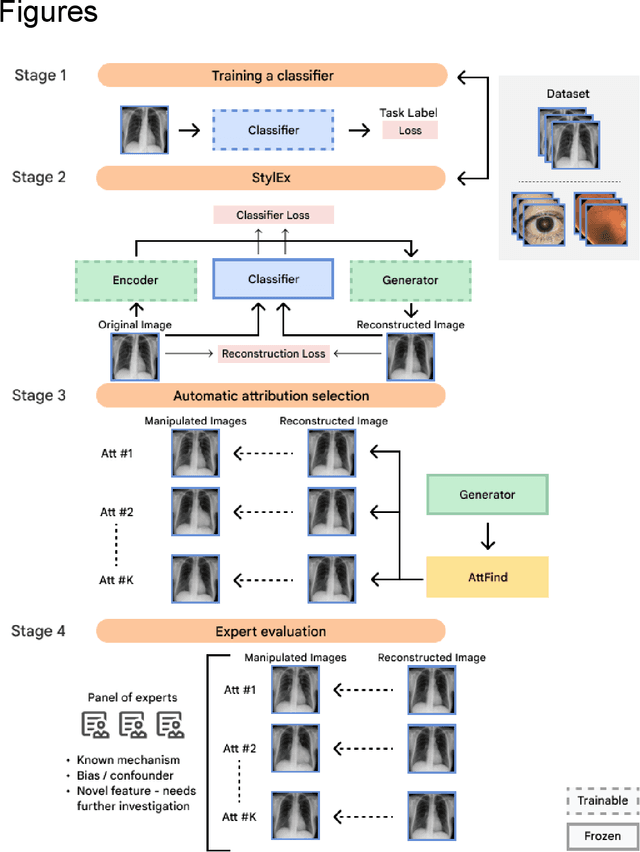
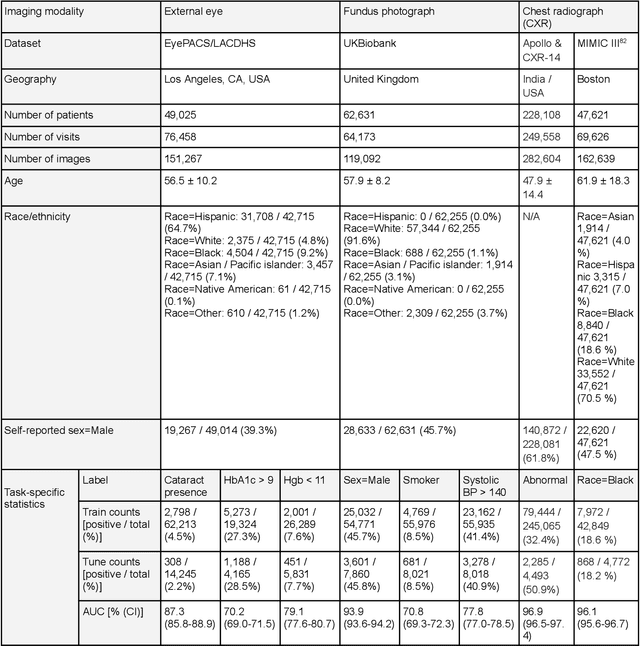
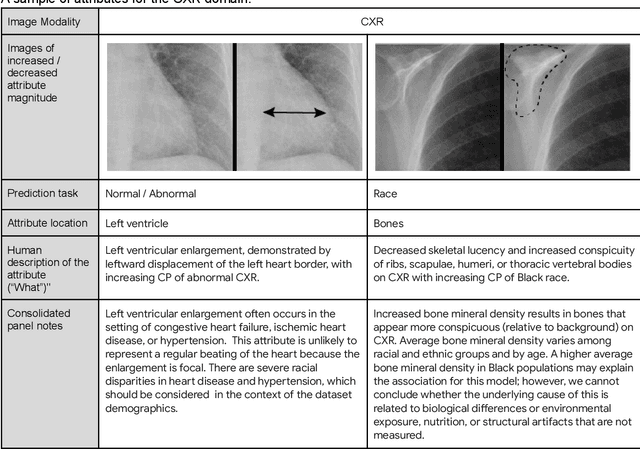
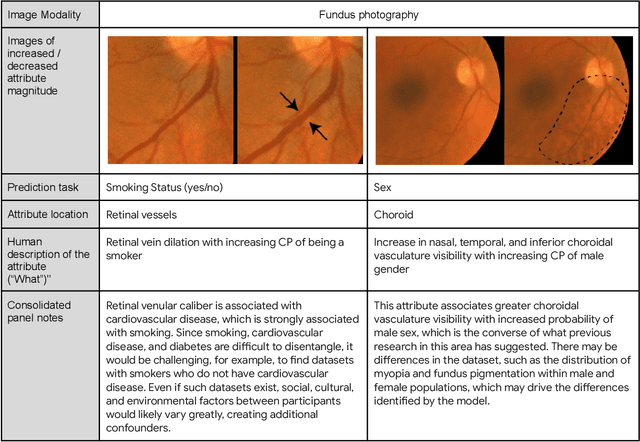
AI models have shown promise in many medical imaging tasks. However, our ability to explain what signals these models have learned is severely lacking. Explanations are needed in order to increase the trust in AI-based models, and could enable novel scientific discovery by uncovering signals in the data that are not yet known to experts. In this paper, we present a method for automatic visual explanations leveraging team-based expertise by generating hypotheses of what visual signals in the images are correlated with the task. We propose the following 4 steps: (i) Train a classifier to perform a given task (ii) Train a classifier guided StyleGAN-based image generator (StylEx) (iii) Automatically detect and visualize the top visual attributes that the classifier is sensitive towards (iv) Formulate hypotheses for the underlying mechanisms, to stimulate future research. Specifically, we present the discovered attributes to an interdisciplinary panel of experts so that hypotheses can account for social and structural determinants of health. We demonstrate results on eight prediction tasks across three medical imaging modalities: retinal fundus photographs, external eye photographs, and chest radiographs. We showcase examples of attributes that capture clinically known features, confounders that arise from factors beyond physiological mechanisms, and reveal a number of physiologically plausible novel attributes. Our approach has the potential to enable researchers to better understand, improve their assessment, and extract new knowledge from AI-based models. Importantly, we highlight that attributes generated by our framework can capture phenomena beyond physiology or pathophysiology, reflecting the real world nature of healthcare delivery and socio-cultural factors. Finally, we intend to release code to enable researchers to train their own StylEx models and analyze their predictive tasks.
Towards Expert-Level Medical Question Answering with Large Language Models
May 16, 2023Karan Singhal, Tao Tu, Juraj Gottweis, Rory Sayres, Ellery Wulczyn, Le Hou, Kevin Clark, Stephen Pfohl, Heather Cole-Lewis, Darlene Neal, Mike Schaekermann, Amy Wang, Mohamed Amin, Sami Lachgar, Philip Mansfield, Sushant Prakash, Bradley Green, Ewa Dominowska, Blaise Aguera y Arcas, Nenad Tomasev, Yun Liu, Renee Wong, Christopher Semturs, S. Sara Mahdavi, Joelle Barral, Dale Webster, Greg S. Corrado, Yossi Matias, Shekoofeh Azizi, Alan Karthikesalingam, Vivek Natarajan
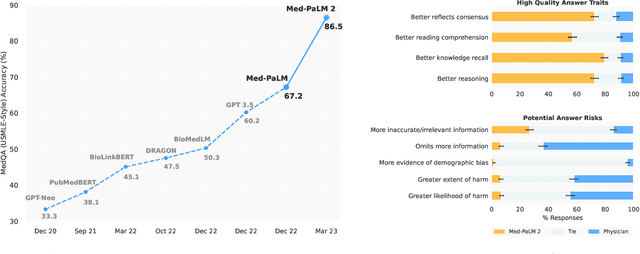
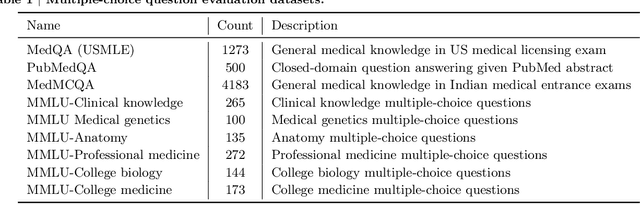

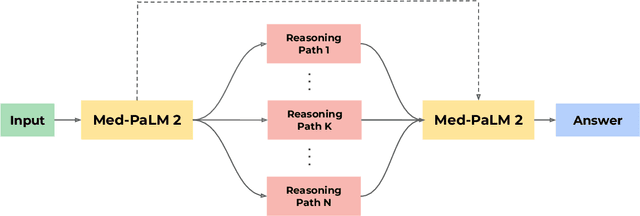
Recent artificial intelligence (AI) systems have reached milestones in "grand challenges" ranging from Go to protein-folding. The capability to retrieve medical knowledge, reason over it, and answer medical questions comparably to physicians has long been viewed as one such grand challenge. Large language models (LLMs) have catalyzed significant progress in medical question answering; Med-PaLM was the first model to exceed a "passing" score in US Medical Licensing Examination (USMLE) style questions with a score of 67.2% on the MedQA dataset. However, this and other prior work suggested significant room for improvement, especially when models' answers were compared to clinicians' answers. Here we present Med-PaLM 2, which bridges these gaps by leveraging a combination of base LLM improvements (PaLM 2), medical domain finetuning, and prompting strategies including a novel ensemble refinement approach. Med-PaLM 2 scored up to 86.5% on the MedQA dataset, improving upon Med-PaLM by over 19% and setting a new state-of-the-art. We also observed performance approaching or exceeding state-of-the-art across MedMCQA, PubMedQA, and MMLU clinical topics datasets. We performed detailed human evaluations on long-form questions along multiple axes relevant to clinical applications. In pairwise comparative ranking of 1066 consumer medical questions, physicians preferred Med-PaLM 2 answers to those produced by physicians on eight of nine axes pertaining to clinical utility (p < 0.001). We also observed significant improvements compared to Med-PaLM on every evaluation axis (p < 0.001) on newly introduced datasets of 240 long-form "adversarial" questions to probe LLM limitations. While further studies are necessary to validate the efficacy of these models in real-world settings, these results highlight rapid progress towards physician-level performance in medical question answering.
Predicting Cardiovascular Disease Risk using Photoplethysmography and Deep Learning
May 09, 2023Wei-Hung Weng, Sebastien Baur, Mayank Daswani, Christina Chen, Lauren Harrell, Sujay Kakarmath, Mariam Jabara, Babak Behsaz, Cory Y. McLean, Yossi Matias, Greg S. Corrado, Shravya Shetty, Shruthi Prabhakara, Yun Liu, Goodarz Danaei, Diego Ardila
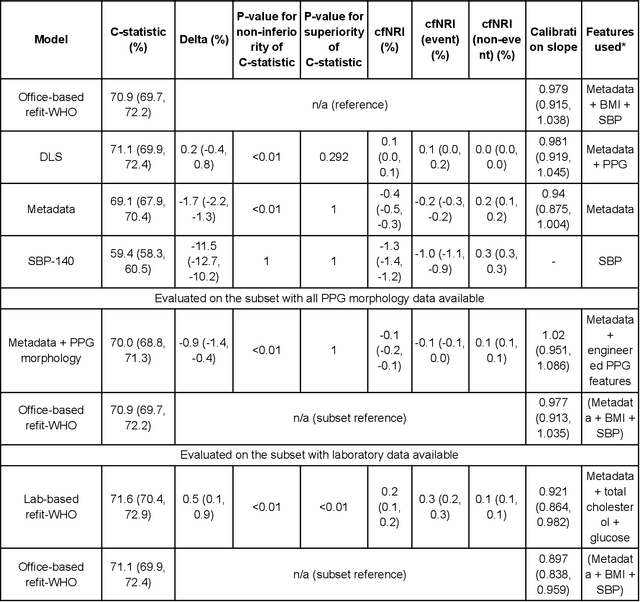
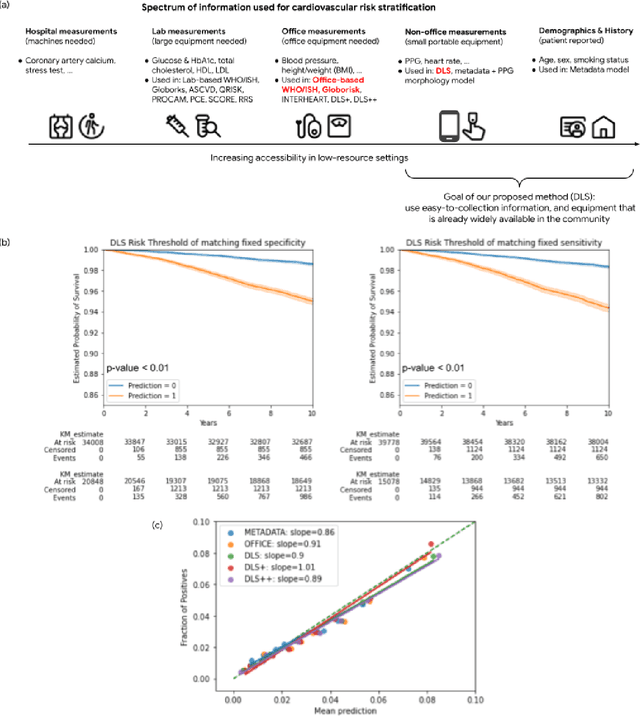
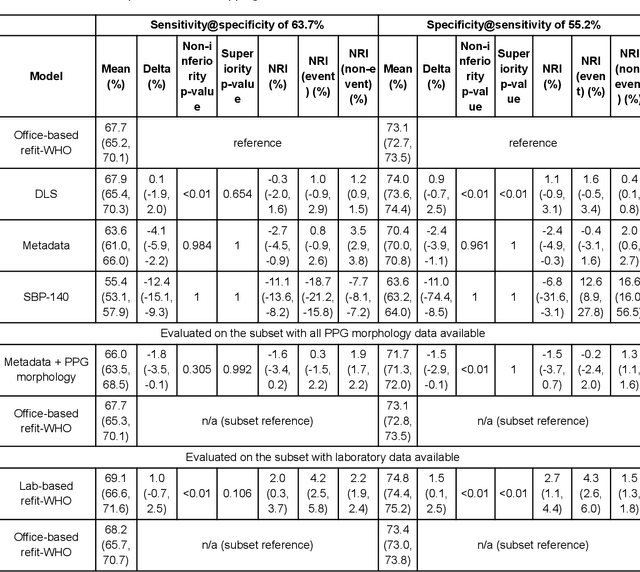
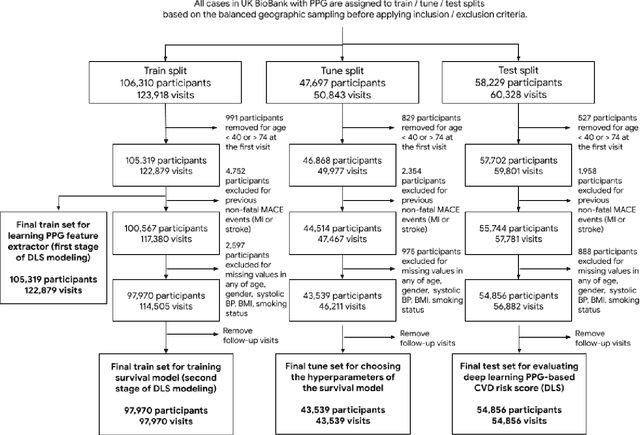
Cardiovascular diseases (CVDs) are responsible for a large proportion of premature deaths in low- and middle-income countries. Early CVD detection and intervention is critical in these populations, yet many existing CVD risk scores require a physical examination or lab measurements, which can be challenging in such health systems due to limited accessibility. Here we investigated the potential to use photoplethysmography (PPG), a sensing technology available on most smartphones that can potentially enable large-scale screening at low cost, for CVD risk prediction. We developed a deep learning PPG-based CVD risk score (DLS) to predict the probability of having major adverse cardiovascular events (MACE: non-fatal myocardial infarction, stroke, and cardiovascular death) within ten years, given only age, sex, smoking status and PPG as predictors. We compared the DLS with the office-based refit-WHO score, which adopts the shared predictors from WHO and Globorisk scores (age, sex, smoking status, height, weight and systolic blood pressure) but refitted on the UK Biobank (UKB) cohort. In UKB cohort, DLS's C-statistic (71.1%, 95% CI 69.9-72.4) was non-inferior to office-based refit-WHO score (70.9%, 95% CI 69.7-72.2; non-inferiority margin of 2.5%, p<0.01). The calibration of the DLS was satisfactory, with a 1.8% mean absolute calibration error. Adding DLS features to the office-based score increased the C-statistic by 1.0% (95% CI 0.6-1.4). DLS predicts ten-year MACE risk comparable with the office-based refit-WHO score. It provides a proof-of-concept and suggests the potential of a PPG-based approach strategies for community-based primary prevention in resource-limited regions.
Large Language Models Encode Clinical Knowledge
Dec 26, 2022Karan Singhal, Shekoofeh Azizi, Tao Tu, S. Sara Mahdavi, Jason Wei, Hyung Won Chung, Nathan Scales, Ajay Tanwani, Heather Cole-Lewis, Stephen Pfohl, Perry Payne, Martin Seneviratne, Paul Gamble, Chris Kelly, Nathaneal Scharli, Aakanksha Chowdhery, Philip Mansfield, Blaise Aguera y Arcas, Dale Webster, Greg S. Corrado, Yossi Matias, Katherine Chou, Juraj Gottweis, Nenad Tomasev, Yun Liu, Alvin Rajkomar, Joelle Barral, Christopher Semturs, Alan Karthikesalingam, Vivek Natarajan
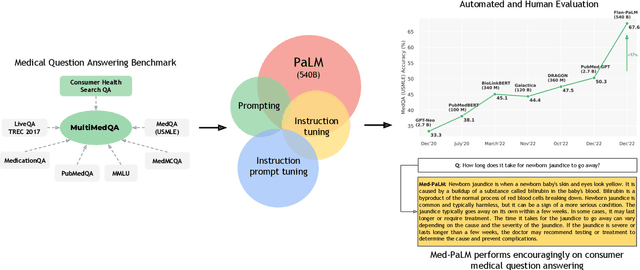
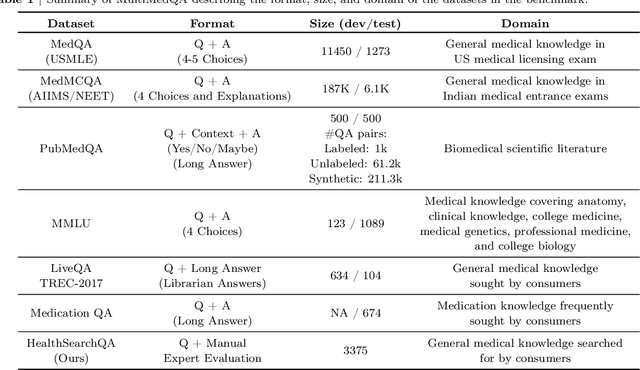
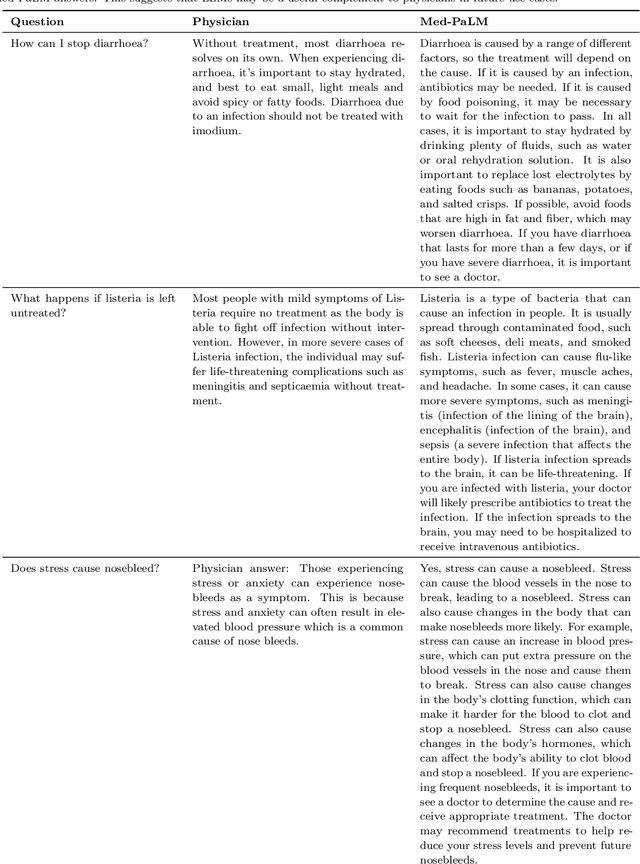
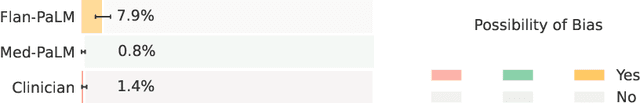
Large language models (LLMs) have demonstrated impressive capabilities in natural language understanding and generation, but the quality bar for medical and clinical applications is high. Today, attempts to assess models' clinical knowledge typically rely on automated evaluations on limited benchmarks. There is no standard to evaluate model predictions and reasoning across a breadth of tasks. To address this, we present MultiMedQA, a benchmark combining six existing open question answering datasets spanning professional medical exams, research, and consumer queries; and HealthSearchQA, a new free-response dataset of medical questions searched online. We propose a framework for human evaluation of model answers along multiple axes including factuality, precision, possible harm, and bias. In addition, we evaluate PaLM (a 540-billion parameter LLM) and its instruction-tuned variant, Flan-PaLM, on MultiMedQA. Using a combination of prompting strategies, Flan-PaLM achieves state-of-the-art accuracy on every MultiMedQA multiple-choice dataset (MedQA, MedMCQA, PubMedQA, MMLU clinical topics), including 67.6% accuracy on MedQA (US Medical License Exam questions), surpassing prior state-of-the-art by over 17%. However, human evaluation reveals key gaps in Flan-PaLM responses. To resolve this we introduce instruction prompt tuning, a parameter-efficient approach for aligning LLMs to new domains using a few exemplars. The resulting model, Med-PaLM, performs encouragingly, but remains inferior to clinicians. We show that comprehension, recall of knowledge, and medical reasoning improve with model scale and instruction prompt tuning, suggesting the potential utility of LLMs in medicine. Our human evaluations reveal important limitations of today's models, reinforcing the importance of both evaluation frameworks and method development in creating safe, helpful LLM models for clinical applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge