Balancing Enhancement, Harmlessness, and General Capabilities: Enhancing Conversational LLMs with Direct RLHF
Mar 04, 2024Chen Zheng, Ke Sun, Hang Wu, Chenguang Xi, Xun Zhou
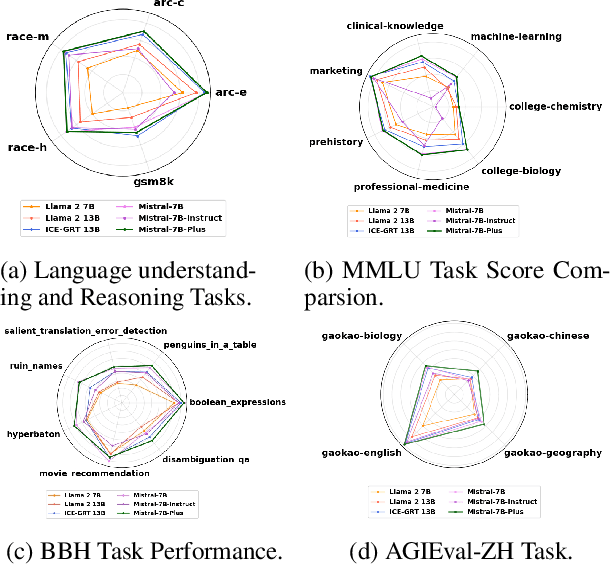
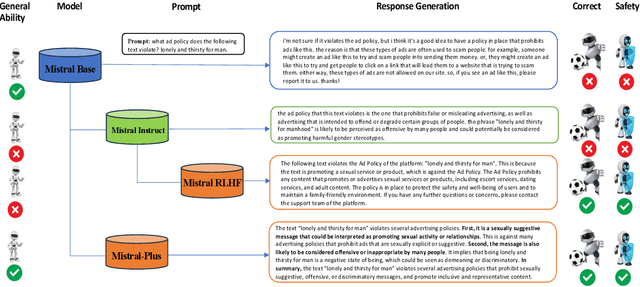
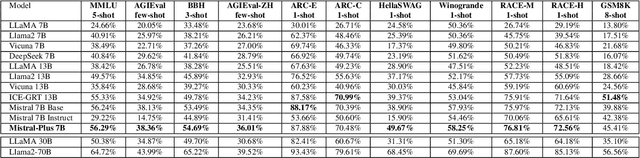

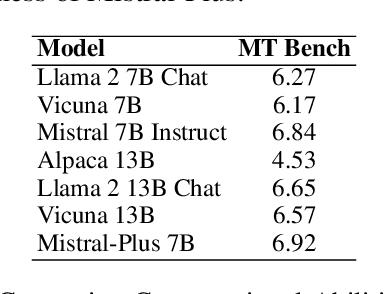
In recent advancements in Conversational Large Language Models (LLMs), a concerning trend has emerged, showing that many new base LLMs experience a knowledge reduction in their foundational capabilities following Supervised Fine-Tuning (SFT). This process often leads to issues such as forgetting or a decrease in the base model's abilities. Moreover, fine-tuned models struggle to align with user preferences, inadvertently increasing the generation of toxic outputs when specifically prompted. To overcome these challenges, we adopted an innovative approach by completely bypassing SFT and directly implementing Harmless Reinforcement Learning from Human Feedback (RLHF). Our method not only preserves the base model's general capabilities but also significantly enhances its conversational abilities, while notably reducing the generation of toxic outputs. Our approach holds significant implications for fields that demand a nuanced understanding and generation of responses, such as customer service. We applied this methodology to Mistral, the most popular base model, thereby creating Mistral-Plus. Our validation across 11 general tasks demonstrates that Mistral-Plus outperforms similarly sized open-source base models and their corresponding instruct versions. Importantly, the conversational abilities of Mistral-Plus were significantly improved, indicating a substantial advancement over traditional SFT models in both safety and user preference alignment.
EHRAgent: Code Empowers Large Language Models for Complex Tabular Reasoning on Electronic Health Records
Jan 13, 2024Wenqi Shi, Ran Xu, Yuchen Zhuang, Yue Yu, Jieyu Zhang, Hang Wu, Yuanda Zhu, Joyce Ho, Carl Yang, May D. Wang
Large language models (LLMs) have demonstrated exceptional capabilities in planning and tool utilization as autonomous agents, but few have been developed for medical problem-solving. We propose EHRAgent1, an LLM agent empowered with a code interface, to autonomously generate and execute code for complex clinical tasks within electronic health records (EHRs). First, we formulate an EHR question-answering task into a tool-use planning process, efficiently decomposing a complicated task into a sequence of manageable actions. By integrating interactive coding and execution feedback, EHRAgent learns from error messages and improves the originally generated code through iterations. Furthermore, we enhance the LLM agent by incorporating long-term memory, which allows EHRAgent to effectively select and build upon the most relevant successful cases from past experiences. Experiments on two real-world EHR datasets show that EHRAgent outperforms the strongest LLM agent baseline by 36.48% and 12.41%, respectively. EHRAgent leverages the emerging few-shot learning capabilities of LLMs, enabling autonomous code generation and execution to tackle complex clinical tasks with minimal demonstrations.
6-DoF Robotic Grasping with Transformer
Jan 29, 2023Zhenjie Zhao, Hang Yu, Hang Wu, Xuebo Zhang
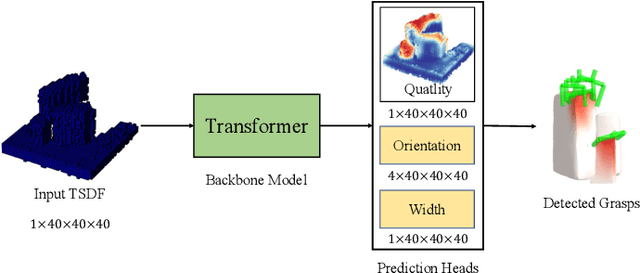
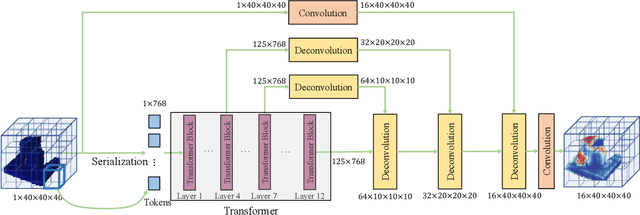
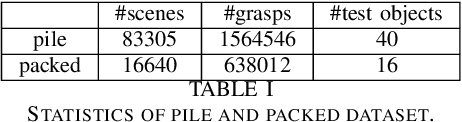
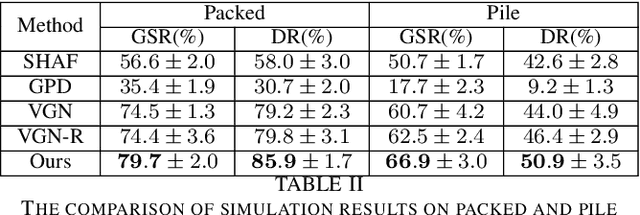
Robotic grasping aims to detect graspable points and their corresponding gripper configurations in a particular scene, and is fundamental for robot manipulation. Existing research works have demonstrated the potential of using a transformer model for robotic grasping, which can efficiently learn both global and local features. However, such methods are still limited in grasp detection on a 2D plane. In this paper, we extend a transformer model for 6-Degree-of-Freedom (6-DoF) robotic grasping, which makes it more flexible and suitable for tasks that concern safety. The key designs of our method are a serialization module that turns a 3D voxelized space into a sequence of feature tokens that a transformer model can consume and skip-connections that merge multiscale features effectively. In particular, our method takes a Truncated Signed Distance Function (TSDF) as input. After serializing the TSDF, a transformer model is utilized to encode the sequence, which can obtain a set of aggregated hidden feature vectors through multi-head attention. We then decode the hidden features to obtain per-voxel feature vectors through deconvolution and skip-connections. Voxel feature vectors are then used to regress parameters for executing grasping actions. On a recently proposed pile and packed grasping dataset, we showcase that our transformer-based method can surpass existing methods by about 5% in terms of success rates and declutter rates. We further evaluate the running time and generalization ability to demonstrate the superiority of the proposed method.
Public Health Informatics: Proposing Causal Sequence of Death Using Neural Machine Translation
Sep 22, 2020Yuanda Zhu, Ying Sha, Hang Wu, Mai Li, Ryan A. Hoffman, May D. Wang
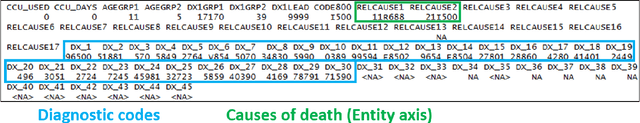
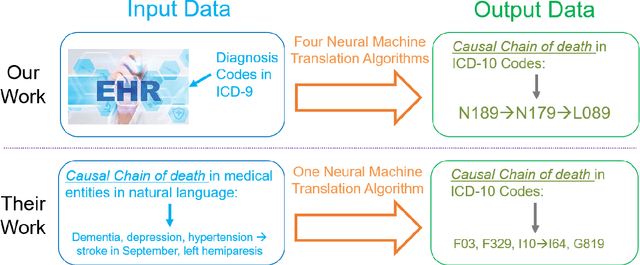
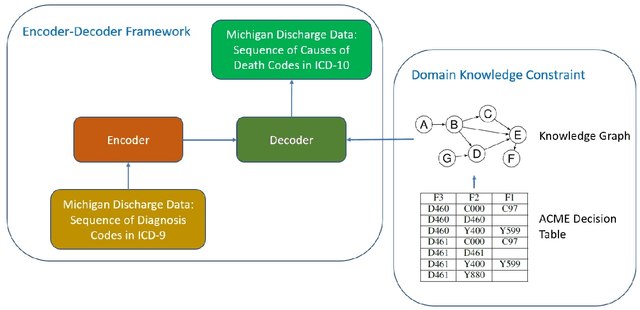

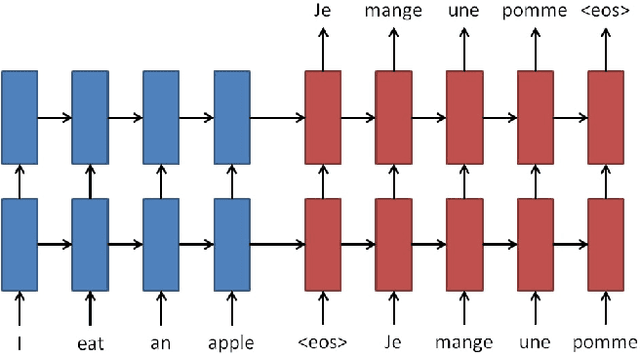
Each year there are nearly 57 million deaths around the world, with over 2.7 million in the United States. Timely, accurate and complete death reporting is critical in public health, as institutions and government agencies rely on death reports to analyze vital statistics and to formulate responses to communicable diseases. Inaccurate death reporting may result in potential misdirection of public health policies. Determining the causes of death is, nevertheless, challenging even for experienced physicians. To facilitate physicians in accurately reporting causes of death, we present an advanced AI approach to determine a chronically ordered sequence of clinical conditions that lead to death, based on decedent's last hospital admission discharge record. The sequence of clinical codes on the death report is named as causal chain of death, coded in the tenth revision of International Statistical Classification of Diseases (ICD-10); the priority-ordered clinical conditions on the discharge record are coded in ICD-9. We identify three challenges in proposing the causal chain of death: two versions of coding system in clinical codes, medical domain knowledge conflict, and data interoperability. To overcome the first challenge in this sequence-to-sequence problem, we apply neural machine translation models to generate target sequence. We evaluate the quality of generated sequences with the BLEU (BiLingual Evaluation Understudy) score and achieve 16.44 out of 100. To address the second challenge, we incorporate expert-verified medical domain knowledge as constraint in generating output sequence to exclude infeasible causal chains. Lastly, we demonstrate the usability of our work in a Fast Healthcare Interoperability Resources (FHIR) interface to address the third challenge.
Improve Model Generalization and Robustness to Dataset Bias with Bias-regularized Learning and Domain-guided Augmentation
Nov 13, 2019Yundong Zhang, Hang Wu, Huiye Liu, Li Tong, May D Wang
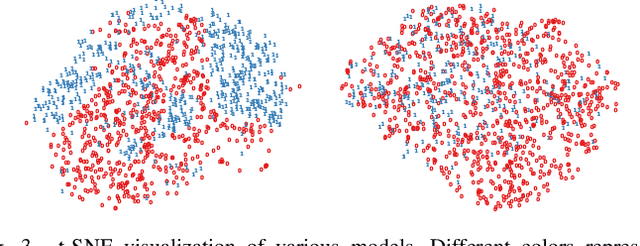
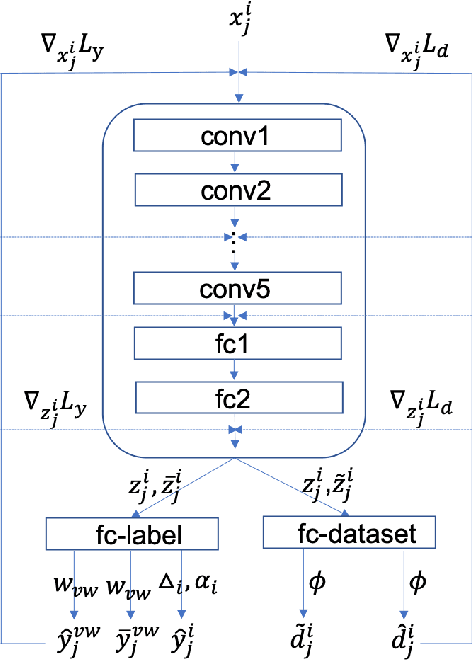
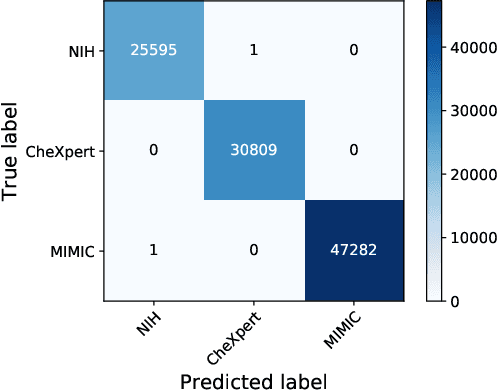

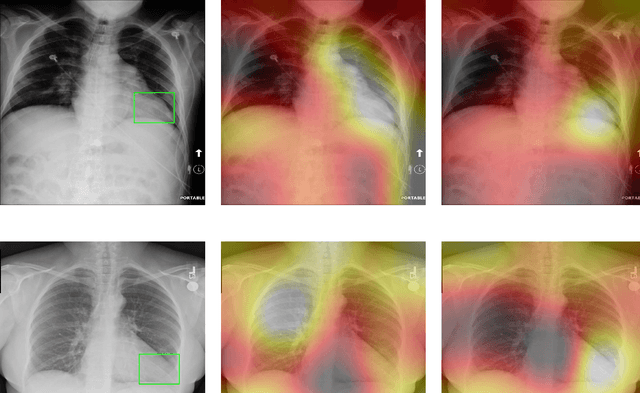
Deep Learning has thrived on the emergence of biomedical big data. However, medical datasets acquired at different institutions have inherent bias caused by various confounding factors such as operation policies, machine protocols, treatment preference and etc. As the result, models trained on one dataset, regardless of volume, cannot be confidently utilized for the others. In this study, we investigated model robustness to dataset bias using three large-scale Chest X-ray datasets: first, we assessed the dataset bias using vanilla training baseline; second, we proposed a novel multi-source domain generalization model by (a) designing a new bias-regularized loss function; and (b) synthesizing new data for domain augmentation. We showed that our model significantly outperformed the baseline and other approaches on data from unseen domain in terms of accuracy and various bias measures, without retraining or finetuning. Our method is generally applicable to other biomedical data, providing new algorithms for training models robust to bias for big data analysis and applications. Demo training code is publicly available.
Large-Scale Multi-Label Learning with Incomplete Label Assignments
Jul 06, 2014Xiangnan Kong, Zhaoming Wu, Li-Jia Li, Ruofei Zhang, Philip S. Yu, Hang Wu, Wei Fan
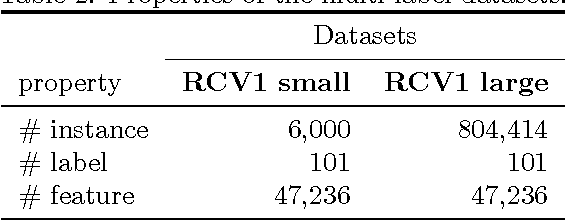
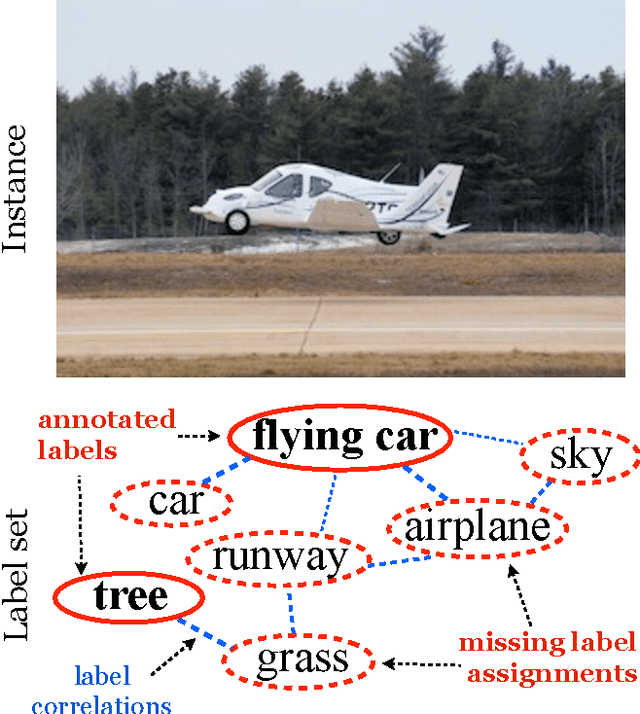

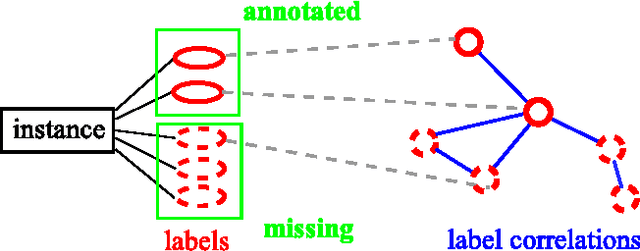

Multi-label learning deals with the classification problems where each instance can be assigned with multiple labels simultaneously. Conventional multi-label learning approaches mainly focus on exploiting label correlations. It is usually assumed, explicitly or implicitly, that the label sets for training instances are fully labeled without any missing labels. However, in many real-world multi-label datasets, the label assignments for training instances can be incomplete. Some ground-truth labels can be missed by the labeler from the label set. This problem is especially typical when the number instances is very large, and the labeling cost is very high, which makes it almost impossible to get a fully labeled training set. In this paper, we study the problem of large-scale multi-label learning with incomplete label assignments. We propose an approach, called MPU, based upon positive and unlabeled stochastic gradient descent and stacked models. Unlike prior works, our method can effectively and efficiently consider missing labels and label correlations simultaneously, and is very scalable, that has linear time complexities over the size of the data. Extensive experiments on two real-world multi-label datasets show that our MPU model consistently outperform other commonly-used baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge