RegWSI: Whole Slide Image Registration using Combined Deep Feature- and Intensity-Based Methods: Winner of the ACROBAT 2023 Challenge
Apr 26, 2024Marek Wodzinski, Niccolò Marini, Manfredo Atzori, Henning Müller
The automatic registration of differently stained whole slide images (WSIs) is crucial for improving diagnosis and prognosis by fusing complementary information emerging from different visible structures. It is also useful to quickly transfer annotations between consecutive or restained slides, thus significantly reducing the annotation time and associated costs. Nevertheless, the slide preparation is different for each stain and the tissue undergoes complex and large deformations. Therefore, a robust, efficient, and accurate registration method is highly desired by the scientific community and hospitals specializing in digital pathology. We propose a two-step hybrid method consisting of (i) deep learning- and feature-based initial alignment algorithm, and (ii) intensity-based nonrigid registration using the instance optimization. The proposed method does not require any fine-tuning to a particular dataset and can be used directly for any desired tissue type and stain. The method scored 1st place in the ACROBAT 2023 challenge. We evaluated using three open datasets: (i) ANHIR, (ii) ACROBAT, and (iii) HyReCo, and performed several ablation studies concerning the resolution used for registration and the initial alignment robustness and stability. The method achieves the most accurate results for the ACROBAT dataset, the cell-level registration accuracy for the restained slides from the HyReCo dataset, and is among the best methods evaluated on the ANHIR dataset. The method does not require any fine-tuning to a new datasets and can be used out-of-the-box for other types of microscopic images. The method is incorporated into the DeeperHistReg framework, allowing others to directly use it to register, transform, and save the WSIs at any desired pyramid level. The proposed method is a significant contribution to the WSI registration, thus advancing the field of digital pathology.
DeeperHistReg: Robust Whole Slide Images Registration Framework
Apr 19, 2024Marek Wodzinski, Niccolò Marini, Manfredo Atzori, Henning Müller
DeeperHistReg is a software framework dedicated to registering whole slide images (WSIs) acquired using multiple stains. It allows one to perform the preprocessing, initial alignment, and nonrigid registration of WSIs acquired using multiple stains (e.g. hematoxylin \& eosin, immunochemistry). The framework implements several state-of-the-art registration algorithms and provides an interface to operate on arbitrary resolution of the WSIs (up to 200k x 200k). The framework is extensible and new algorithms can be easily integrated by other researchers. The framework is available both as a PyPI package and as a Docker container.
A comparative study on wearables and single-camera video for upper-limb out-of-thelab activity recognition with different deep learning architectures
Feb 04, 2024Mario Martínez-Zarzuela, David González-Ortega, Míriam Antón-Rodríguez, Francisco Javier Díaz-Pernas, Henning Müller, Cristina Simón-Martínez
The use of a wide range of computer vision solutions, and more recently high-end Inertial Measurement Units (IMU) have become increasingly popular for assessing human physical activity in clinical and research settings. Nevertheless, to increase the feasibility of patient tracking in out-of-the-lab settings, it is necessary to use a reduced number of devices for movement acquisition. Promising solutions in this context are IMU-based wearables and single camera systems. Additionally, the development of machine learning systems able to recognize and digest clinically relevant data in-the-wild is needed, and therefore determining the ideal input to those is crucial.
Benchmarking the CoW with the TopCoW Challenge: Topology-Aware Anatomical Segmentation of the Circle of Willis for CTA and MRA
Dec 29, 2023Kaiyuan Yang, Fabio Musio, Yihui Ma, Norman Juchler, Johannes C. Paetzold, Rami Al-Maskari, Luciano Höher, Hongwei Bran Li, Ibrahim Ethem Hamamci, Anjany Sekuboyina, Suprosanna Shit, Houjing Huang, Diana Waldmannstetter, Florian Kofler, Fernando Navarro, Martin Menten, Ivan Ezhov, Daniel Rueckert, Iris Vos, Ynte Ruigrok, Birgitta Velthuis, Hugo Kuijf, Julien Hämmerli, Catherine Wurster, Philippe Bijlenga, Laura Westphal, Jeroen Bisschop, Elisa Colombo, Hakim Baazaoui, Andrew Makmur, James Hallinan, Bene Wiestler, Jan S. Kirschke, Roland Wiest, Emmanuel Montagnon, Laurent Letourneau-Guillon, Adrian Galdran, Francesco Galati, Daniele Falcetta, Maria A. Zuluaga, Chaolong Lin, Haoran Zhao, Zehan Zhang, Sinyoung Ra, Jongyun Hwang, Hyunjin Park, Junqiang Chen, Marek Wodzinski, Henning Müller, Pengcheng Shi, Wei Liu, Ting Ma, Cansu Yalçin, Rachika E. Hamadache, Joaquim Salvi, Xavier Llado, Uma Maria Lal-Trehan Estrada, Valeriia Abramova, Luca Giancardo, Arnau Oliver, Jialu Liu, Haibin Huang, Yue Cui, Zehang Lin, Yusheng Liu, Shunzhi Zhu, Tatsat R. Patel, Vincent M. Tutino, Maysam Orouskhani, Huayu Wang, Mahmud Mossa-Basha, Chengcheng Zhu, Maximilian R. Rokuss, Yannick Kirchhoff, Nico Disch, Julius Holzschuh, Fabian Isensee, Klaus Maier-Hein, Yuki Sato, Sven Hirsch, Susanne Wegener, Bjoern Menze
The Circle of Willis (CoW) is an important network of arteries connecting major circulations of the brain. Its vascular architecture is believed to affect the risk, severity, and clinical outcome of serious neuro-vascular diseases. However, characterizing the highly variable CoW anatomy is still a manual and time-consuming expert task. The CoW is usually imaged by two angiographic imaging modalities, magnetic resonance angiography (MRA) and computed tomography angiography (CTA), but there exist limited public datasets with annotations on CoW anatomy, especially for CTA. Therefore we organized the TopCoW Challenge in 2023 with the release of an annotated CoW dataset and invited submissions worldwide for the CoW segmentation task, which attracted over 140 registered participants from four continents. TopCoW dataset was the first public dataset with voxel-level annotations for CoW's 13 vessel components, made possible by virtual-reality (VR) technology. It was also the first dataset with paired MRA and CTA from the same patients. TopCoW challenge aimed to tackle the CoW characterization problem as a multiclass anatomical segmentation task with an emphasis on topological metrics. The top performing teams managed to segment many CoW components to Dice scores around 90%, but with lower scores for communicating arteries and rare variants. There were also topological mistakes for predictions with high Dice scores. Additional topological analysis revealed further areas for improvement in detecting certain CoW components and matching CoW variant's topology accurately. TopCoW represented a first attempt at benchmarking the CoW anatomical segmentation task for MRA and CTA, both morphologically and topologically.
Automatic Aorta Segmentation with Heavily Augmented, High-Resolution 3-D ResUNet: Contribution to the SEG.A Challenge
Oct 24, 2023Marek Wodzinski, Henning Müller
Automatic aorta segmentation from 3-D medical volumes is an important yet difficult task. Several factors make the problem challenging, e.g. the possibility of aortic dissection or the difficulty with segmenting and annotating the small branches. This work presents a contribution by the MedGIFT team to the SEG.A challenge organized during the MICCAI 2023 conference. We propose a fully automated algorithm based on deep encoder-decoder architecture. The main assumption behind our work is that data preprocessing and augmentation are much more important than the deep architecture, especially in low data regimes. Therefore, the solution is based on a variant of traditional convolutional U-Net. The proposed solution achieved a Dice score above 0.9 for all testing cases with the highest stability among all participants. The method scored 1st, 4th, and 3rd in terms of the clinical evaluation, quantitative results, and volumetric meshing quality, respectively. We freely release the source code, pretrained model, and provide access to the algorithm on the Grand-Challenge platform.
Uncovering Unique Concept Vectors through Latent Space Decomposition
Jul 14, 2023Mara Graziani, Laura O' Mahony, An-Phi Nguyen, Henning Müller, Vincent Andrearczyk
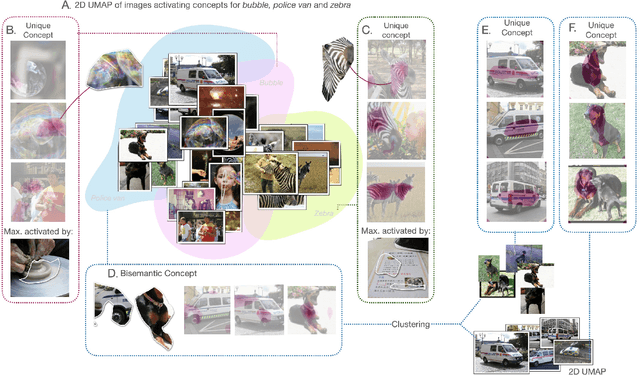
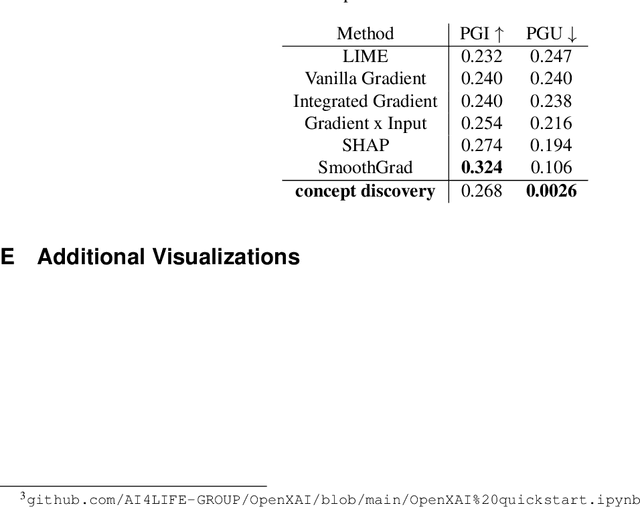
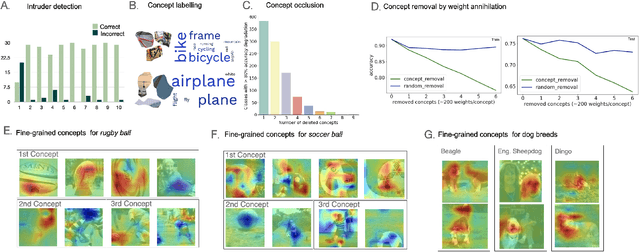

Interpreting the inner workings of deep learning models is crucial for establishing trust and ensuring model safety. Concept-based explanations have emerged as a superior approach that is more interpretable than feature attribution estimates such as pixel saliency. However, defining the concepts for the interpretability analysis biases the explanations by the user's expectations on the concepts. To address this, we propose a novel post-hoc unsupervised method that automatically uncovers the concepts learned by deep models during training. By decomposing the latent space of a layer in singular vectors and refining them by unsupervised clustering, we uncover concept vectors aligned with directions of high variance that are relevant to the model prediction, and that point to semantically distinct concepts. Our extensive experiments reveal that the majority of our concepts are readily understandable to humans, exhibit coherency, and bear relevance to the task at hand. Moreover, we showcase the practical utility of our method in dataset exploration, where our concept vectors successfully identify outlier training samples affected by various confounding factors. This novel exploration technique has remarkable versatility to data types and model architectures and it will facilitate the identification of biases and the discovery of sources of error within training data.
A Hierarchical Transformer Encoder to Improve Entire Neoplasm Segmentation on Whole Slide Image of Hepatocellular Carcinoma
Jul 11, 2023Zhuxian Guo, Qitong Wang, Henning Müller, Themis Palpanas, Nicolas Loménie, Camille Kurtz
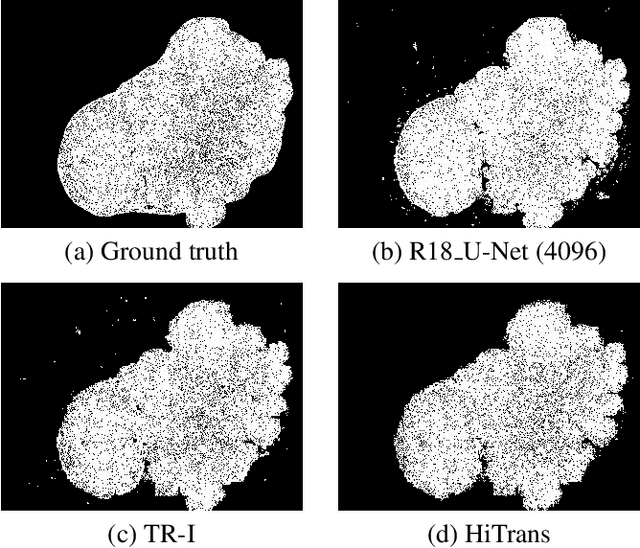

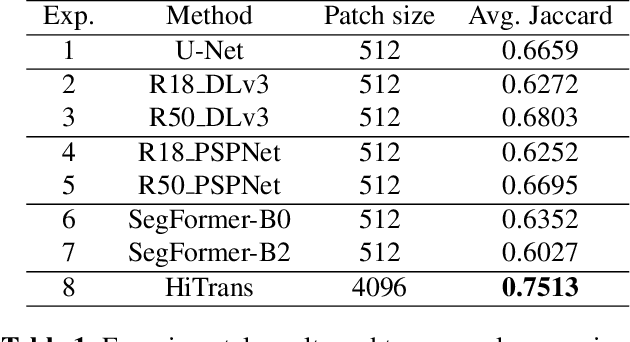
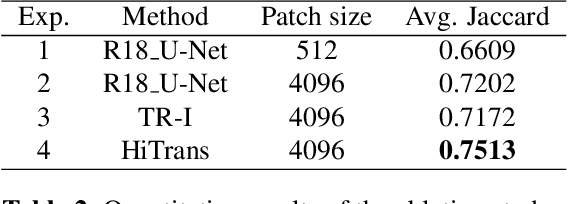

In digital histopathology, entire neoplasm segmentation on Whole Slide Image (WSI) of Hepatocellular Carcinoma (HCC) plays an important role, especially as a preprocessing filter to automatically exclude healthy tissue, in histological molecular correlations mining and other downstream histopathological tasks. The segmentation task remains challenging due to HCC's inherent high-heterogeneity and the lack of dependency learning in large field of view. In this article, we propose a novel deep learning architecture with a hierarchical Transformer encoder, HiTrans, to learn the global dependencies within expanded 4096$\times$4096 WSI patches. HiTrans is designed to encode and decode the patches with larger reception fields and the learned global dependencies, compared to the state-of-the-art Fully Convolutional Neural networks (FCNN). Empirical evaluations verified that HiTrans leads to better segmentation performance by taking into account regional and global dependency information.
The ACROBAT 2022 Challenge: Automatic Registration Of Breast Cancer Tissue
May 29, 2023Philippe Weitz, Masi Valkonen, Leslie Solorzano, Circe Carr, Kimmo Kartasalo, Constance Boissin, Sonja Koivukoski, Aino Kuusela, Dusan Rasic, Yanbo Feng, Sandra Sinius Pouplier, Abhinav Sharma, Kajsa Ledesma Eriksson, Stephanie Robertson, Christian Marzahl, Chandler D. Gatenbee, Alexander R. A. Anderson, Marek Wodzinski, Artur Jurgas, Niccolò Marini, Manfredo Atzori, Henning Müller, Daniel Budelmann, Nick Weiss, Stefan Heldmann, Johannes Lotz, Jelmer M. Wolterink, Bruno De Santi, Abhijeet Patil, Amit Sethi, Satoshi Kondo, Satoshi Kasai, Kousuke Hirasawa, Mahtab Farrokh, Neeraj Kumar, Russell Greiner, Leena Latonen, Anne-Vibeke Laenkholm, Johan Hartman, Pekka Ruusuvuori, Mattias Rantalainen
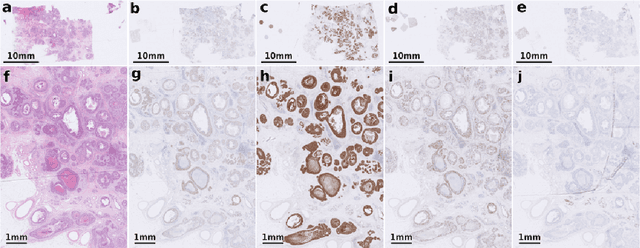
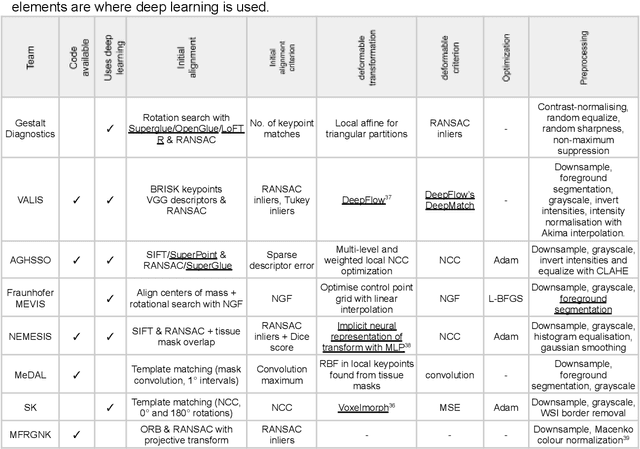
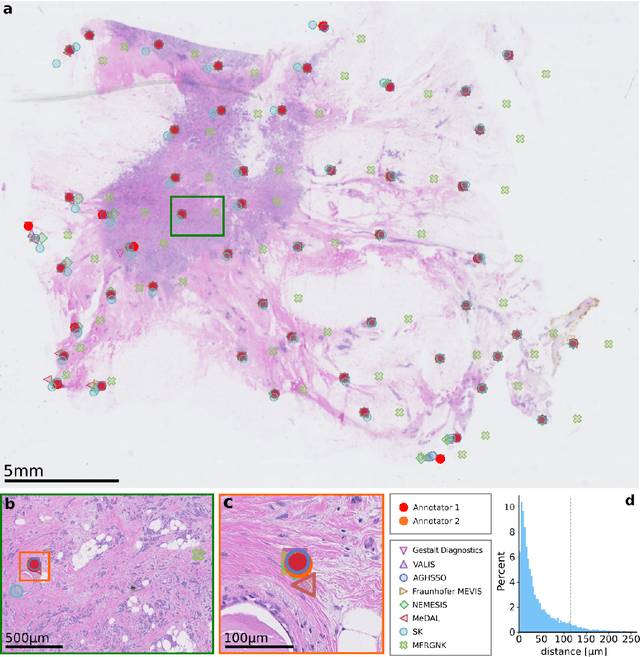
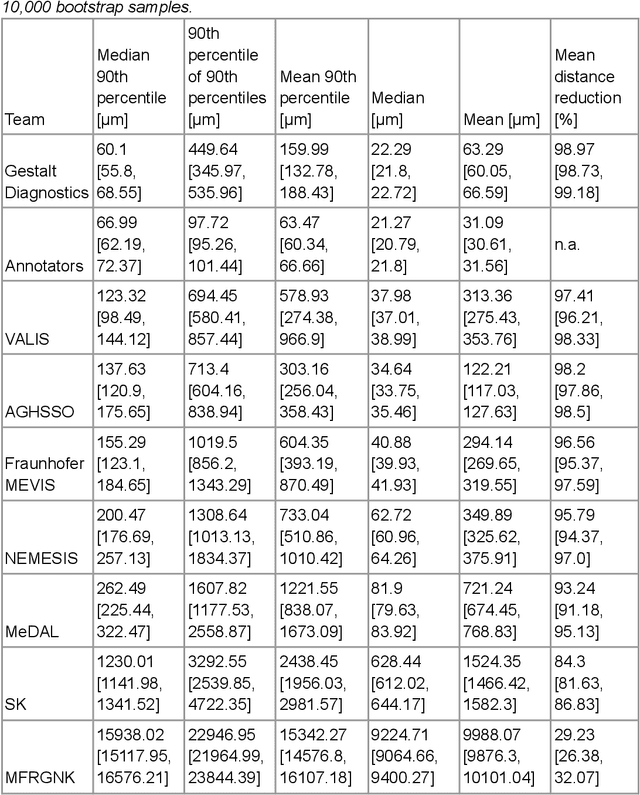
The alignment of tissue between histopathological whole-slide-images (WSI) is crucial for research and clinical applications. Advances in computing, deep learning, and availability of large WSI datasets have revolutionised WSI analysis. Therefore, the current state-of-the-art in WSI registration is unclear. To address this, we conducted the ACROBAT challenge, based on the largest WSI registration dataset to date, including 4,212 WSIs from 1,152 breast cancer patients. The challenge objective was to align WSIs of tissue that was stained with routine diagnostic immunohistochemistry to its H&E-stained counterpart. We compare the performance of eight WSI registration algorithms, including an investigation of the impact of different WSI properties and clinical covariates. We find that conceptually distinct WSI registration methods can lead to highly accurate registration performances and identify covariates that impact performances across methods. These results establish the current state-of-the-art in WSI registration and guide researchers in selecting and developing methods.
Evaluating Post-hoc Interpretability with Intrinsic Interpretability
May 04, 2023José Pereira Amorim, Pedro Henriques Abreu, João Santos, Henning Müller
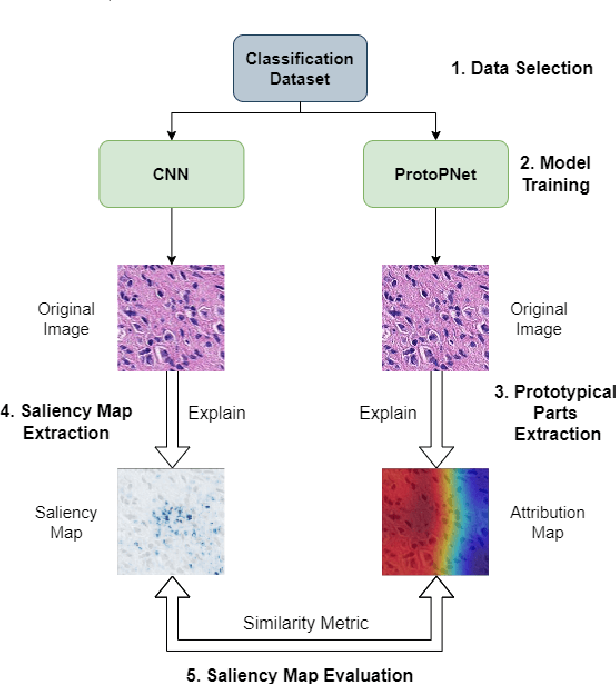

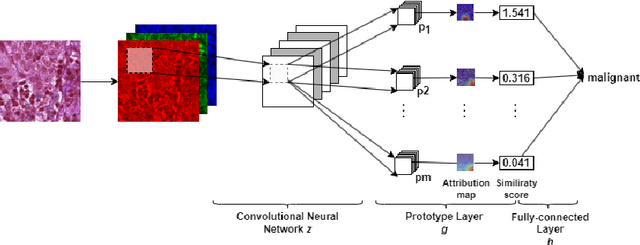

Despite Convolutional Neural Networks having reached human-level performance in some medical tasks, their clinical use has been hindered by their lack of interpretability. Two major interpretability strategies have been proposed to tackle this problem: post-hoc methods and intrinsic methods. Although there are several post-hoc methods to interpret DL models, there is significant variation between the explanations provided by each method, and it a difficult to validate them due to the lack of ground-truth. To address this challenge, we adapted the intrinsical interpretable ProtoPNet for the context of histopathology imaging and compared the attribution maps produced by it and the saliency maps made by post-hoc methods. To evaluate the similarity between saliency map methods and attribution maps we adapted 10 saliency metrics from the saliency model literature, and used the breast cancer metastases detection dataset PatchCamelyon with 327,680 patches of histopathological images of sentinel lymph node sections to validate the proposed approach. Overall, SmoothGrad and Occlusion were found to have a statistically bigger overlap with ProtoPNet while Deconvolution and Lime have been found to have the least.
VIDIMU. Multimodal video and IMU kinematic dataset on daily life activities using affordable devices
Mar 27, 2023Mario Martínez-Zarzuela, Javier González-Alonso, Míriam Antón-Rodríguez, Francisco J. Díaz-Pernas, Henning Müller, Cristina Simón-Martínez
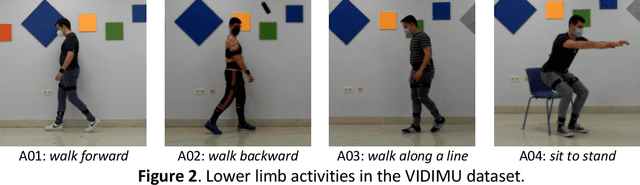

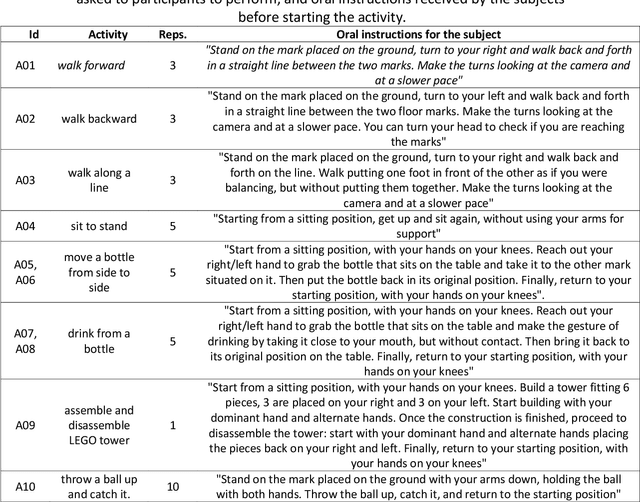

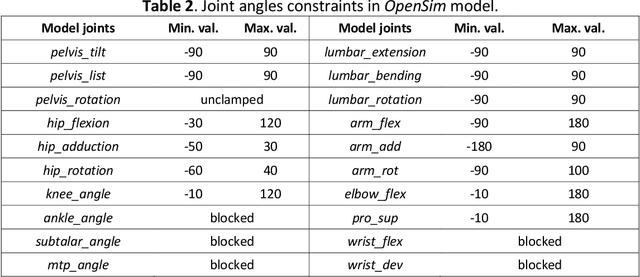
Human activity recognition and clinical biomechanics are challenging problems in physical telerehabilitation medicine. However, most publicly available datasets on human body movements cannot be used to study both problems in an out-of-the-lab movement acquisition setting. The objective of the VIDIMU dataset is to pave the way towards affordable patient tracking solutions for remote daily life activities recognition and kinematic analysis. The dataset includes 13 activities registered using a commodity camera and five inertial sensors. The video recordings were acquired in 54 subjects, of which 16 also had simultaneous recordings of inertial sensors. The novelty of VIDIMU lies in: i) the clinical relevance of the chosen movements, ii) the combined utilization of affordable video and custom sensors, and iii) the implementation of state-of-the-art tools for multimodal data processing of 3D body pose tracking and motion reconstruction in a musculoskeletal model from inertial data. The validation confirms that a minimally disturbing acquisition protocol, performed according to real-life conditions can provide a comprehensive picture of human joint angles during daily life activities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge