PathM3: A Multimodal Multi-Task Multiple Instance Learning Framework for Whole Slide Image Classification and Captioning
Mar 13, 2024Qifeng Zhou, Wenliang Zhong, Yuzhi Guo, Michael Xiao, Hehuan Ma, Junzhou Huang
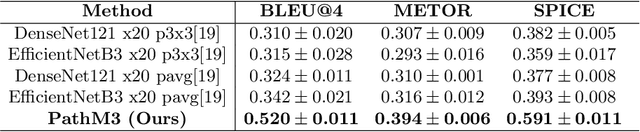
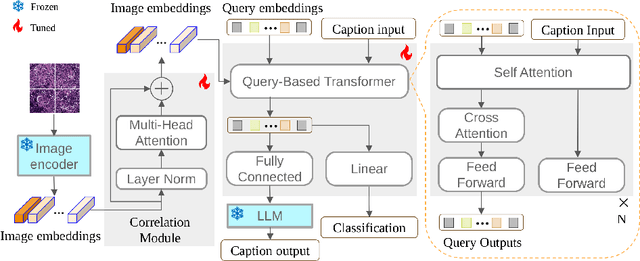
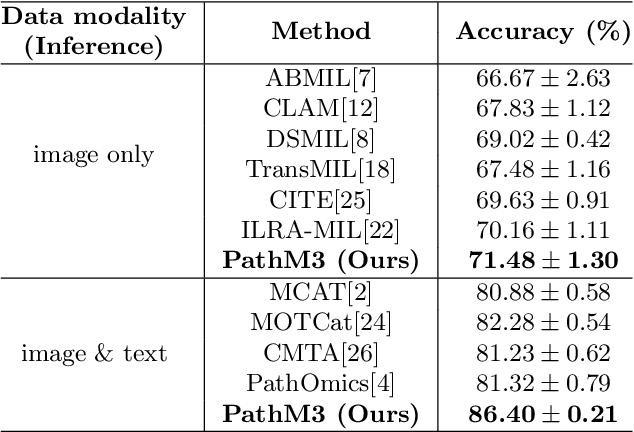

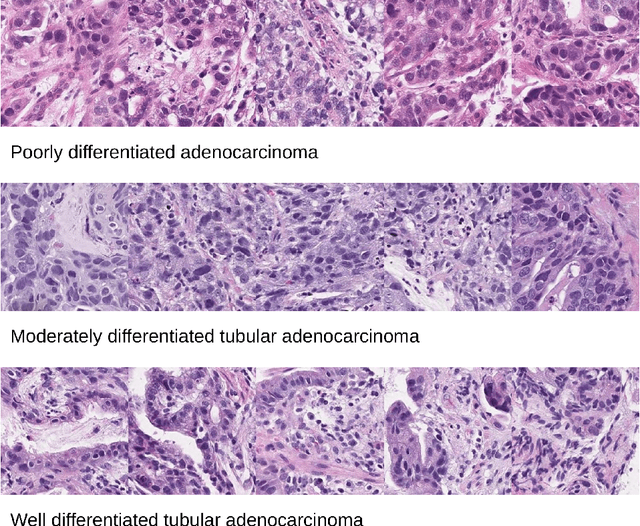
In the field of computational histopathology, both whole slide images (WSIs) and diagnostic captions provide valuable insights for making diagnostic decisions. However, aligning WSIs with diagnostic captions presents a significant challenge. This difficulty arises from two main factors: 1) Gigapixel WSIs are unsuitable for direct input into deep learning models, and the redundancy and correlation among the patches demand more attention; and 2) Authentic WSI diagnostic captions are extremely limited, making it difficult to train an effective model. To overcome these obstacles, we present PathM3, a multimodal, multi-task, multiple instance learning (MIL) framework for WSI classification and captioning. PathM3 adapts a query-based transformer to effectively align WSIs with diagnostic captions. Given that histopathology visual patterns are redundantly distributed across WSIs, we aggregate each patch feature with MIL method that considers the correlations among instances. Furthermore, our PathM3 overcomes data scarcity in WSI-level captions by leveraging limited WSI diagnostic caption data in the manner of multi-task joint learning. Extensive experiments with improved classification accuracy and caption generation demonstrate the effectiveness of our method on both WSI classification and captioning task.
Distribution-Free Fair Federated Learning with Small Samples
Feb 25, 2024Qichuan Yin, Junzhou Huang, Huaxiu Yao, Linjun Zhang
As federated learning gains increasing importance in real-world applications due to its capacity for decentralized data training, addressing fairness concerns across demographic groups becomes critically important. However, most existing machine learning algorithms for ensuring fairness are designed for centralized data environments and generally require large-sample and distributional assumptions, underscoring the urgent need for fairness techniques adapted for decentralized and heterogeneous systems with finite-sample and distribution-free guarantees. To address this issue, this paper introduces FedFaiREE, a post-processing algorithm developed specifically for distribution-free fair learning in decentralized settings with small samples. Our approach accounts for unique challenges in decentralized environments, such as client heterogeneity, communication costs, and small sample sizes. We provide rigorous theoretical guarantees for both fairness and accuracy, and our experimental results further provide robust empirical validation for our proposed method.
Segment Any Cell: A SAM-based Auto-prompting Fine-tuning Framework for Nuclei Segmentation
Jan 24, 2024Saiyang Na, Yuzhi Guo, Feng Jiang, Hehuan Ma, Junzhou Huang
In the rapidly evolving field of AI research, foundational models like BERT and GPT have significantly advanced language and vision tasks. The advent of pretrain-prompting models such as ChatGPT and Segmentation Anything Model (SAM) has further revolutionized image segmentation. However, their applications in specialized areas, particularly in nuclei segmentation within medical imaging, reveal a key challenge: the generation of high-quality, informative prompts is as crucial as applying state-of-the-art (SOTA) fine-tuning techniques on foundation models. To address this, we introduce Segment Any Cell (SAC), an innovative framework that enhances SAM specifically for nuclei segmentation. SAC integrates a Low-Rank Adaptation (LoRA) within the attention layer of the Transformer to improve the fine-tuning process, outperforming existing SOTA methods. It also introduces an innovative auto-prompt generator that produces effective prompts to guide segmentation, a critical factor in handling the complexities of nuclei segmentation in biomedical imaging. Our extensive experiments demonstrate the superiority of SAC in nuclei segmentation tasks, proving its effectiveness as a tool for pathologists and researchers. Our contributions include a novel prompt generation strategy, automated adaptability for diverse segmentation tasks, the innovative application of Low-Rank Attention Adaptation in SAM, and a versatile framework for semantic segmentation challenges.
MIVC: Multiple Instance Visual Component for Visual-Language Models
Dec 28, 2023Wenyi Wu, Qi Li, Wenliang Zhong, Junzhou Huang
Vision-language models have been widely explored across a wide range of tasks and achieve satisfactory performance. However, it's under-explored how to consolidate entity understanding through a varying number of images and to align it with the pre-trained language models for generative tasks. In this paper, we propose MIVC, a general multiple instance visual component to bridge the gap between various image inputs with off-the-shelf vision-language models by aggregating visual representations in a permutation-invariant fashion through a neural network. We show that MIVC could be plugged into the visual-language models to improve the model performance consistently on visual question answering, classification and captioning tasks on a public available e-commerce dataset with multiple images per product. Furthermore, we show that the component provides insight into the contribution of each image to the downstream tasks.
Structure-Aware DropEdge Towards Deep Graph Convolutional Networks
Jun 21, 2023Jiaqi Han, Wenbing Huang, Yu Rong, Tingyang Xu, Fuchun Sun, Junzhou Huang
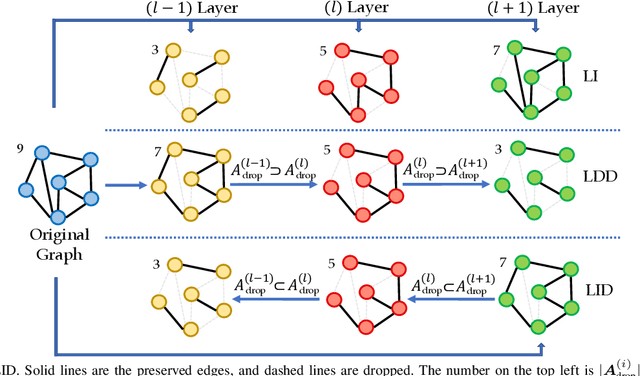
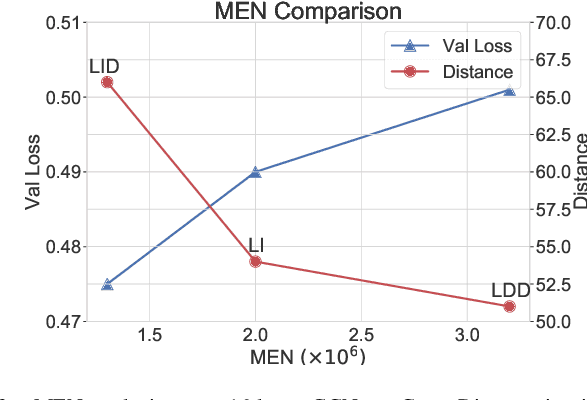
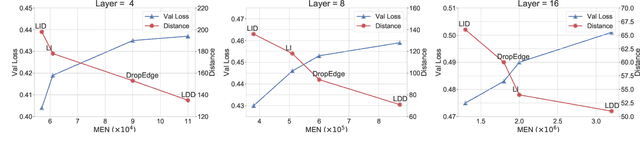
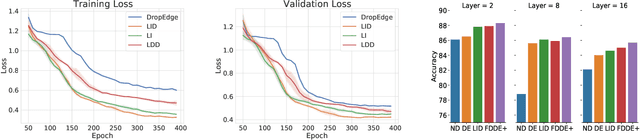
It has been discovered that Graph Convolutional Networks (GCNs) encounter a remarkable drop in performance when multiple layers are piled up. The main factor that accounts for why deep GCNs fail lies in over-smoothing, which isolates the network output from the input with the increase of network depth, weakening expressivity and trainability. In this paper, we start by investigating refined measures upon DropEdge -- an existing simple yet effective technique to relieve over-smoothing. We term our method as DropEdge++ for its two structure-aware samplers in contrast to DropEdge: layer-dependent sampler and feature-dependent sampler. Regarding the layer-dependent sampler, we interestingly find that increasingly sampling edges from the bottom layer yields superior performance than the decreasing counterpart as well as DropEdge. We theoretically reveal this phenomenon with Mean-Edge-Number (MEN), a metric closely related to over-smoothing. For the feature-dependent sampler, we associate the edge sampling probability with the feature similarity of node pairs, and prove that it further correlates the convergence subspace of the output layer with the input features. Extensive experiments on several node classification benchmarks, including both full- and semi- supervised tasks, illustrate the efficacy of DropEdge++ and its compatibility with a variety of backbones by achieving generally better performance over DropEdge and the no-drop version.
ChatGraph: Interpretable Text Classification by Converting ChatGPT Knowledge to Graphs
May 03, 2023Yucheng Shi, Hehuan Ma, Wenliang Zhong, Gengchen Mai, Xiang Li, Tianming Liu, Junzhou Huang
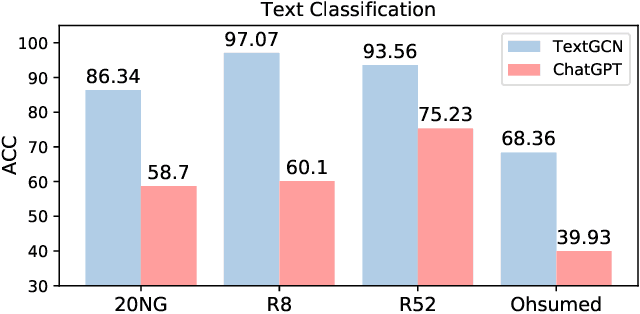
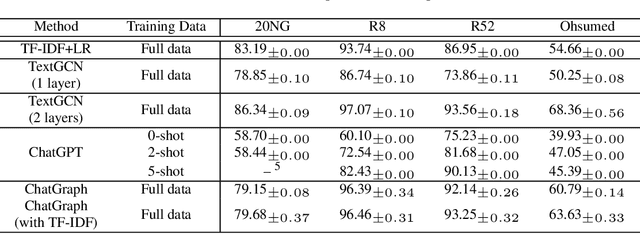
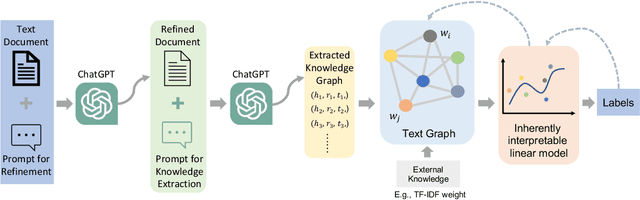

ChatGPT, as a recently launched large language model (LLM), has shown superior performance in various natural language processing (NLP) tasks. However, two major limitations hinder its potential applications: (1) the inflexibility of finetuning on downstream tasks and (2) the lack of interpretability in the decision-making process. To tackle these limitations, we propose a novel framework that leverages the power of ChatGPT for specific tasks, such as text classification, while improving its interpretability. The proposed framework conducts a knowledge graph extraction task to extract refined and structural knowledge from the raw data using ChatGPT. The rich knowledge is then converted into a graph, which is further used to train an interpretable linear classifier to make predictions. To evaluate the effectiveness of our proposed method, we conduct experiments on four datasets. The result shows that our method can significantly improve the performance compared to directly utilizing ChatGPT for text classification tasks. And our method provides a more transparent decision-making process compared with previous text classification methods.
Hierarchical Transformer for Survival Prediction Using Multimodality Whole Slide Images and Genomics
Nov 29, 2022Chunyuan Li, Xinliang Zhu, Jiawen Yao, Junzhou Huang
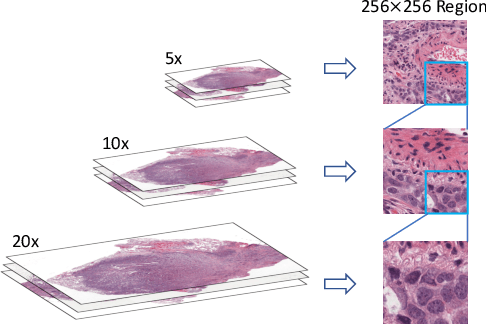
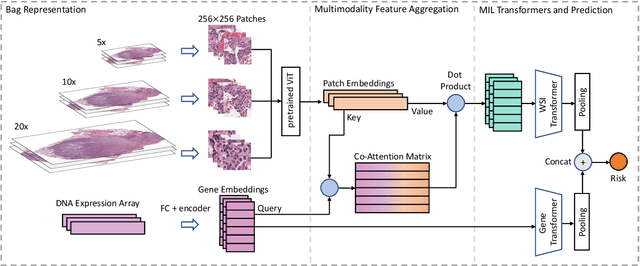
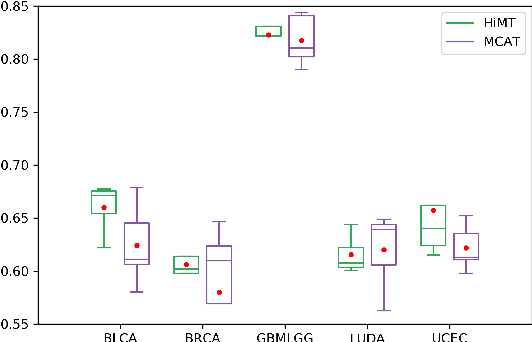

Learning good representation of giga-pixel level whole slide pathology images (WSI) for downstream tasks is critical. Previous studies employ multiple instance learning (MIL) to represent WSIs as bags of sampled patches because, for most occasions, only slide-level labels are available, and only a tiny region of the WSI is disease-positive area. However, WSI representation learning still remains an open problem due to: (1) patch sampling on a higher resolution may be incapable of depicting microenvironment information such as the relative position between the tumor cells and surrounding tissues, while patches at lower resolution lose the fine-grained detail; (2) extracting patches from giant WSI results in large bag size, which tremendously increases the computational cost. To solve the problems, this paper proposes a hierarchical-based multimodal transformer framework that learns a hierarchical mapping between pathology images and corresponding genes. Precisely, we randomly extract instant-level patch features from WSIs with different magnification. Then a co-attention mapping between imaging and genomics is learned to uncover the pairwise interaction and reduce the space complexity of imaging features. Such early fusion makes it computationally feasible to use MIL Transformer for the survival prediction task. Our architecture requires fewer GPU resources compared with benchmark methods while maintaining better WSI representation ability. We evaluate our approach on five cancer types from the Cancer Genome Atlas database and achieved an average c-index of $0.673$, outperforming the state-of-the-art multimodality methods.
Pareto-aware Neural Architecture Generation for Diverse Computational Budgets
Oct 14, 2022Yong Guo, Yaofo Chen, Yin Zheng, Qi Chen, Peilin Zhao, Jian Chen, Junzhou Huang, Mingkui Tan
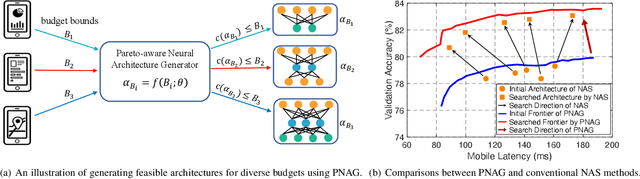
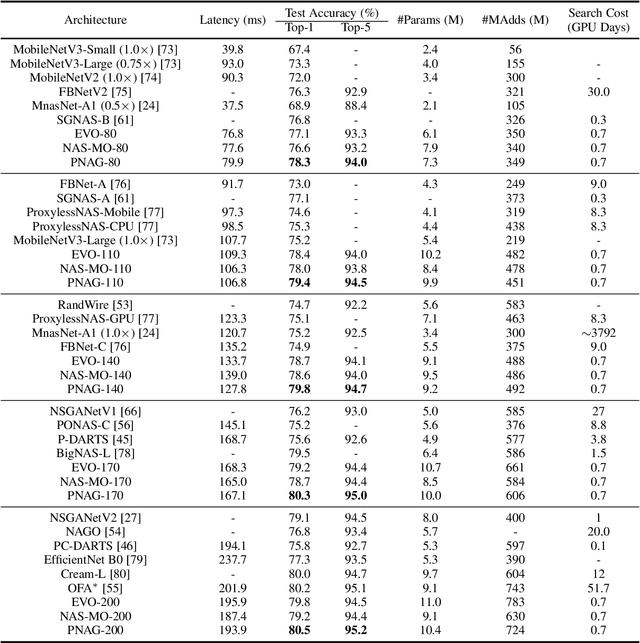
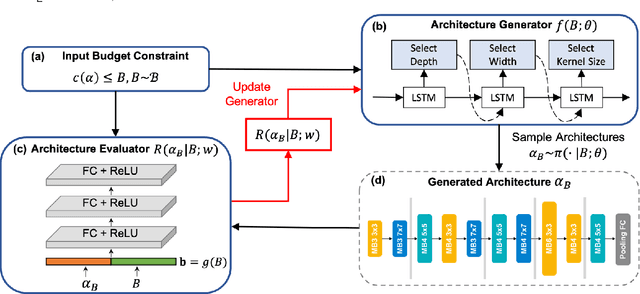
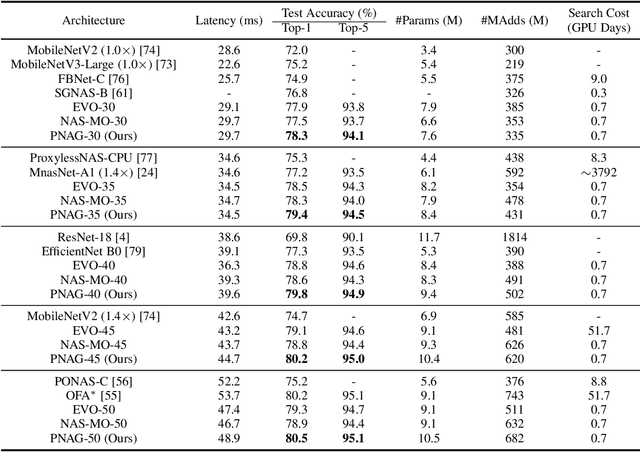
Designing feasible and effective architectures under diverse computational budgets, incurred by different applications/devices, is essential for deploying deep models in real-world applications. To achieve this goal, existing methods often perform an independent architecture search process for each target budget, which is very inefficient yet unnecessary. More critically, these independent search processes cannot share their learned knowledge (i.e., the distribution of good architectures) with each other and thus often result in limited search results. To address these issues, we propose a Pareto-aware Neural Architecture Generator (PNAG) which only needs to be trained once and dynamically produces the Pareto optimal architecture for any given budget via inference. To train our PNAG, we learn the whole Pareto frontier by jointly finding multiple Pareto optimal architectures under diverse budgets. Such a joint search algorithm not only greatly reduces the overall search cost but also improves the search results. Extensive experiments on three hardware platforms (i.e., mobile device, CPU, and GPU) show the superiority of our method over existing methods.
MARS: A Motif-based Autoregressive Model for Retrosynthesis Prediction
Sep 27, 2022Jiahan Liu, Chaochao Yan, Yang Yu, Chan Lu, Junzhou Huang, Le Ou-Yang, Peilin Zhao
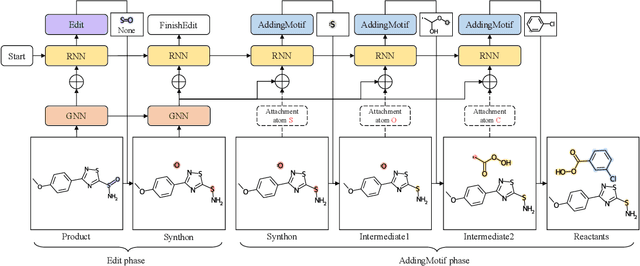
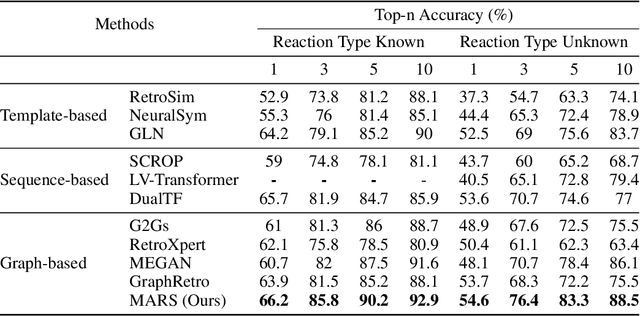
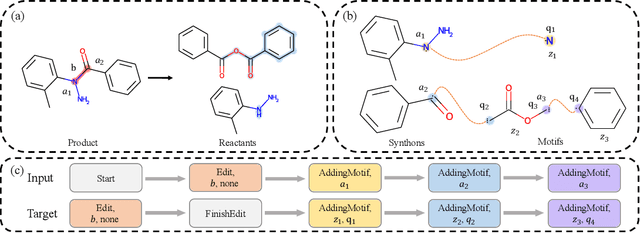
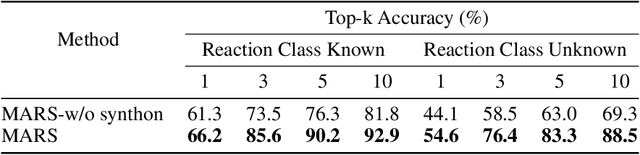
Retrosynthesis is a major task for drug discovery. It is formulated as a graph-generating problem by many existing approaches. Specifically, these methods firstly identify the reaction center, and break target molecule accordingly to generate synthons. Reactants are generated by either adding atoms sequentially to synthon graphs or directly adding proper leaving groups. However, both two strategies suffer since adding atoms results in a long prediction sequence which increases generation difficulty, while adding leaving groups can only consider the ones in the training set which results in poor generalization. In this paper, we propose a novel end-to-end graph generation model for retrosynthesis prediction, which sequentially identifies the reaction center, generates the synthons, and adds motifs to the synthons to generate reactants. Since chemically meaningful motifs are bigger than atoms and smaller than leaving groups, our method enjoys lower prediction complexity than adding atoms and better generalization than adding leaving groups. Experiments on a benchmark dataset show that the proposed model significantly outperforms previous state-of-the-art algorithms.
DeepNoise: Disentanglement of Experimental Noise from Real Biological Signals based on Fluorescent Microscopy Image Classification via Deep Learning
Sep 13, 2022Sen Yang, Tao Shen, Yuqi Fang, Xiyue Wang, Jun Zhang, Wei Yang, Junzhou Huang, Xiao Han
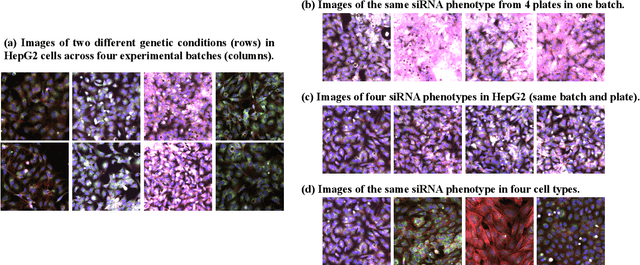
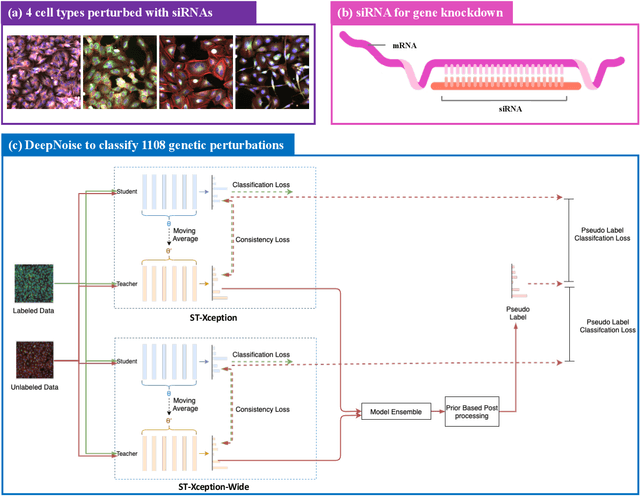
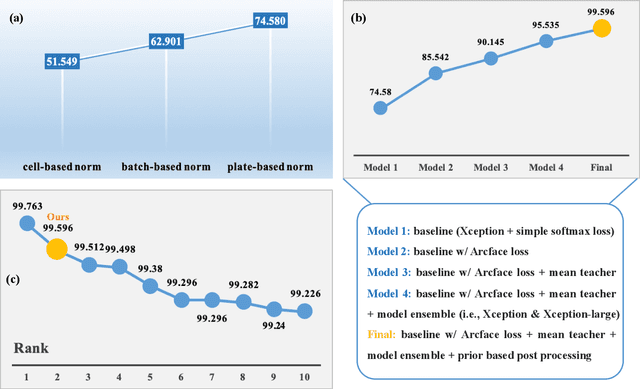
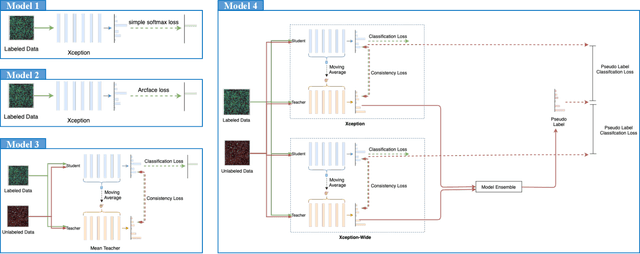
The high-content image-based assay is commonly leveraged for identifying the phenotypic impact of genetic perturbations in biology field. However, a persistent issue remains unsolved during experiments: the interferential technical noise caused by systematic errors (e.g., temperature, reagent concentration, and well location) is always mixed up with the real biological signals, leading to misinterpretation of any conclusion drawn. Here, we show a mean teacher based deep learning model (DeepNoise) that can disentangle biological signals from the experimental noise. Specifically, we aim to classify the phenotypic impact of 1,108 different genetic perturbations screened from 125,510 fluorescent microscopy images, which are totally unrecognizable by human eye. We validate our model by participating in the Recursion Cellular Image Classification Challenge, and our proposed method achieves an extremely high classification score (Acc: 99.596%), ranking the 2nd place among 866 participating groups. This promising result indicates the successful separation of biological and technical factors, which might help decrease the cost of treatment development and expedite the drug discovery process.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge