Adaptive Conditional Quantile Neural Processes
Jun 08, 2023Peiman Mohseni, Nick Duffield, Bani Mallick, Arman Hasanzadeh
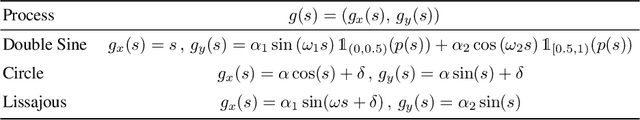
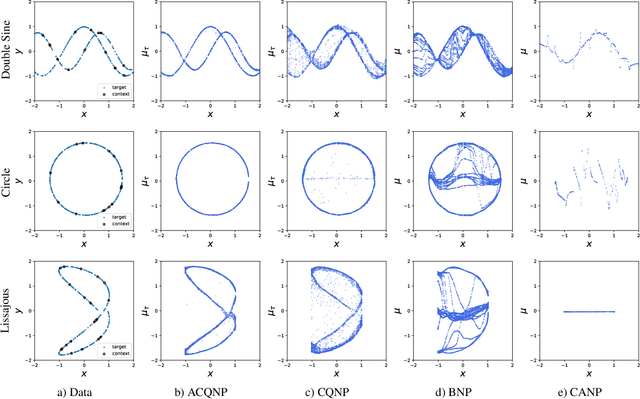
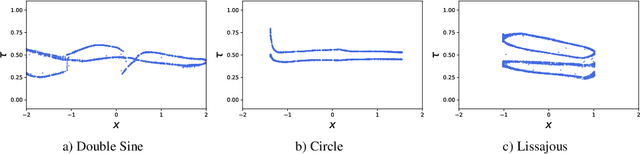
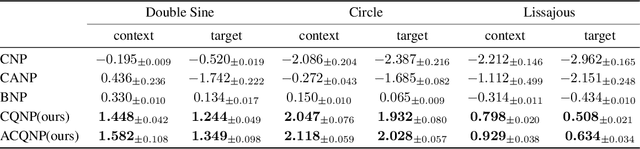
Neural processes are a family of probabilistic models that inherit the flexibility of neural networks to parameterize stochastic processes. Despite providing well-calibrated predictions, especially in regression problems, and quick adaptation to new tasks, the Gaussian assumption that is commonly used to represent the predictive likelihood fails to capture more complicated distributions such as multimodal ones. To overcome this limitation, we propose Conditional Quantile Neural Processes (CQNPs), a new member of the neural processes family, which exploits the attractive properties of quantile regression in modeling the distributions irrespective of their form. By introducing an extension of quantile regression where the model learns to focus on estimating informative quantiles, we show that the sampling efficiency and prediction accuracy can be further enhanced. Our experiments with real and synthetic datasets demonstrate substantial improvements in predictive performance compared to the baselines, and better modeling of heterogeneous distributions' characteristics such as multimodality.
MoReL: Multi-omics Relational Learning
Mar 15, 2022Arman Hasanzadeh, Ehsan Hajiramezanali, Nick Duffield, Xiaoning Qian
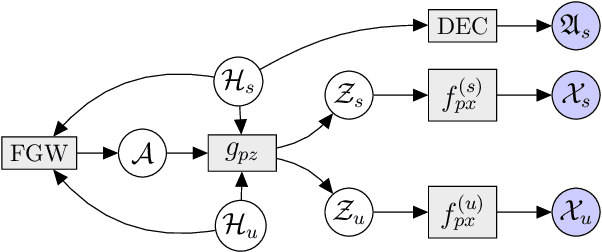
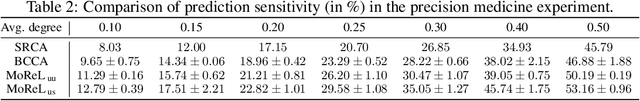
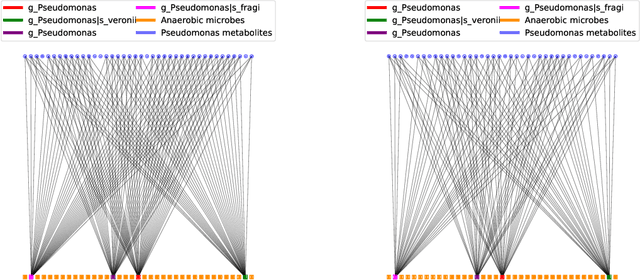

Multi-omics data analysis has the potential to discover hidden molecular interactions, revealing potential regulatory and/or signal transduction pathways for cellular processes of interest when studying life and disease systems. One of critical challenges when dealing with real-world multi-omics data is that they may manifest heterogeneous structures and data quality as often existing data may be collected from different subjects under different conditions for each type of omics data. We propose a novel deep Bayesian generative model to efficiently infer a multi-partite graph encoding molecular interactions across such heterogeneous views, using a fused Gromov-Wasserstein (FGW) regularization between latent representations of corresponding views for integrative analysis. With such an optimal transport regularization in the deep Bayesian generative model, it not only allows incorporating view-specific side information, either with graph-structured or unstructured data in different views, but also increases the model flexibility with the distribution-based regularization. This allows efficient alignment of heterogeneous latent variable distributions to derive reliable interaction predictions compared to the existing point-based graph embedding methods. Our experiments on several real-world datasets demonstrate enhanced performance of MoReL in inferring meaningful interactions compared to existing baselines.
Bayesian Graph Contrastive Learning
Jan 22, 2022Arman Hasanzadeh, Mohammadreza Armandpour, Ehsan Hajiramezanali, Mingyuan Zhou, Nick Duffield, Krishna Narayanan
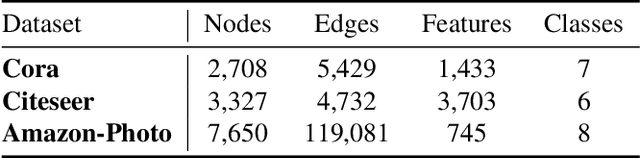

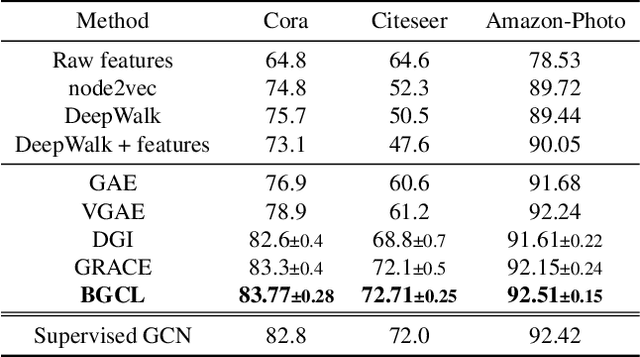
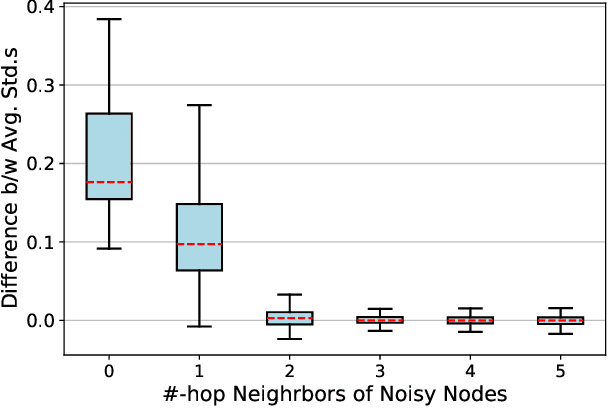
Contrastive learning has become a key component of self-supervised learning approaches for graph-structured data. However, despite their success, existing graph contrastive learning methods are incapable of uncertainty quantification for node representations or their downstream tasks, limiting their application in high-stakes domains. In this paper, we propose a novel Bayesian perspective of graph contrastive learning methods showing random augmentations leads to stochastic encoders. As a result, our proposed method represents each node by a distribution in the latent space in contrast to existing techniques which embed each node to a deterministic vector. By learning distributional representations, we provide uncertainty estimates in downstream graph analytics tasks and increase the expressive power of the predictive model. In addition, we propose a Bayesian framework to infer the probability of perturbations in each view of the contrastive model, eliminating the need for a computationally expensive search for hyperparameter tuning. We empirically show a considerable improvement in performance compared to existing state-of-the-art methods on several benchmark datasets.
BayReL: Bayesian Relational Learning for Multi-omics Data Integration
Oct 22, 2020Ehsan Hajiramezanali, Arman Hasanzadeh, Nick Duffield, Krishna R Narayanan, Xiaoning Qian
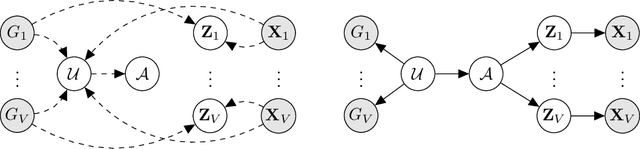
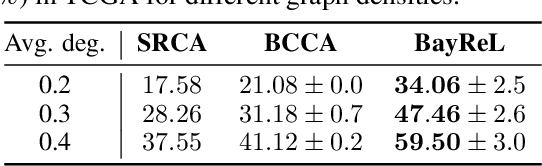
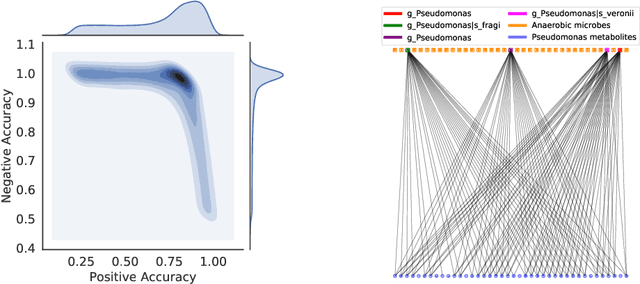
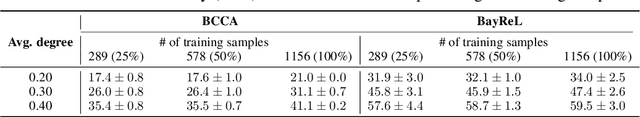
High-throughput molecular profiling technologies have produced high-dimensional multi-omics data, enabling systematic understanding of living systems at the genome scale. Studying molecular interactions across different data types helps reveal signal transduction mechanisms across different classes of molecules. In this paper, we develop a novel Bayesian representation learning method that infers the relational interactions across multi-omics data types. Our method, Bayesian Relational Learning (BayReL) for multi-omics data integration, takes advantage of a priori known relationships among the same class of molecules, modeled as a graph at each corresponding view, to learn view-specific latent variables as well as a multi-partite graph that encodes the interactions across views. Our experiments on several real-world datasets demonstrate enhanced performance of BayReL in inferring meaningful interactions compared to existing baselines.
Bayesian Graph Neural Networks with Adaptive Connection Sampling
Jun 30, 2020Arman Hasanzadeh, Ehsan Hajiramezanali, Shahin Boluki, Mingyuan Zhou, Nick Duffield, Krishna Narayanan, Xiaoning Qian
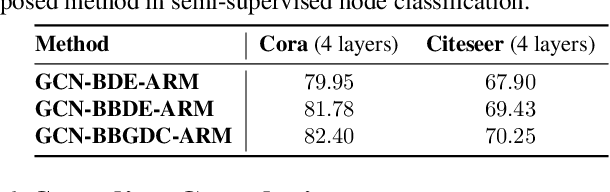
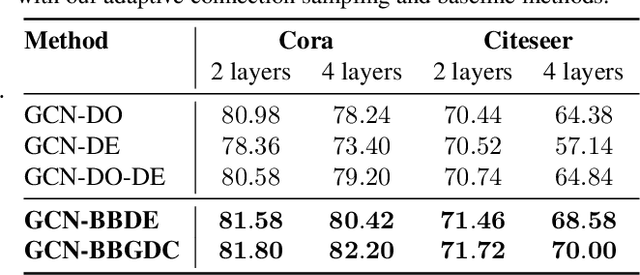
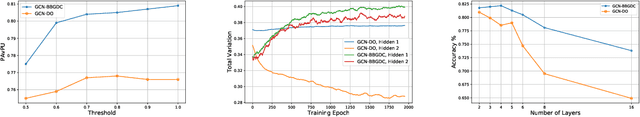

We propose a unified framework for adaptive connection sampling in graph neural networks (GNNs) that generalizes existing stochastic regularization methods for training GNNs. The proposed framework not only alleviates over-smoothing and over-fitting tendencies of deep GNNs, but also enables learning with uncertainty in graph analytic tasks with GNNs. Instead of using fixed sampling rates or hand-tuning them as model hyperparameters in existing stochastic regularization methods, our adaptive connection sampling can be trained jointly with GNN model parameters in both global and local fashions. GNN training with adaptive connection sampling is shown to be mathematically equivalent to an efficient approximation of training Bayesian GNNs. Experimental results with ablation studies on benchmark datasets validate that adaptively learning the sampling rate given graph training data is the key to boost the performance of GNNs in semi-supervised node classification, less prone to over-smoothing and over-fitting with more robust prediction.
Semi-Implicit Stochastic Recurrent Neural Networks
Oct 28, 2019Ehsan Hajiramezanali, Arman Hasanzadeh, Nick Duffield, Krishna Narayanan, Mingyuan Zhou, Xiaoning Qian
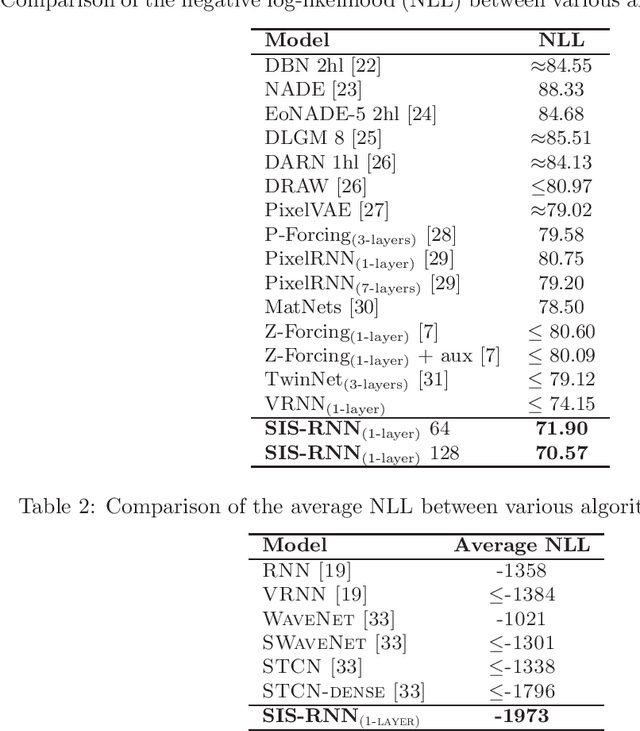
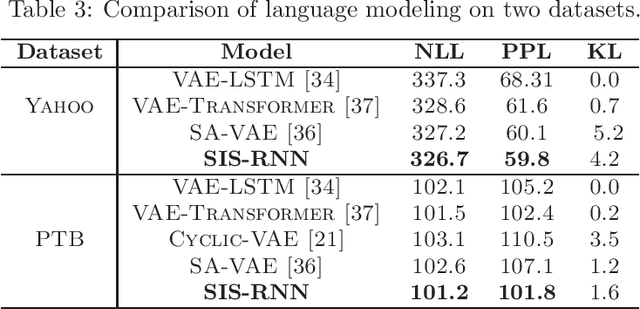
Stochastic recurrent neural networks with latent random variables of complex dependency structures have shown to be more successful in modeling sequential data than deterministic deep models. However, the majority of existing methods have limited expressive power due to the Gaussian assumption of latent variables. In this paper, we advocate learning implicit latent representations using semi-implicit variational inference to further increase model flexibility. Semi-implicit stochastic recurrent neural network(SIS-RNN) is developed to enrich inferred model posteriors that may have no analytic density functions, as long as independent random samples can be generated via reparameterization. Extensive experiments in different tasks on real-world datasets show that SIS-RNN outperforms the existing methods.
Temporal Network Sampling
Oct 18, 2019Nesreen K. Ahmed, Nick Duffield, Ryan A. Rossi
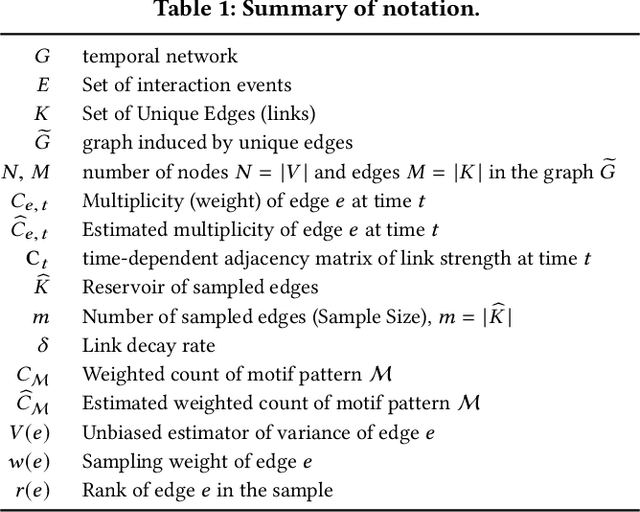
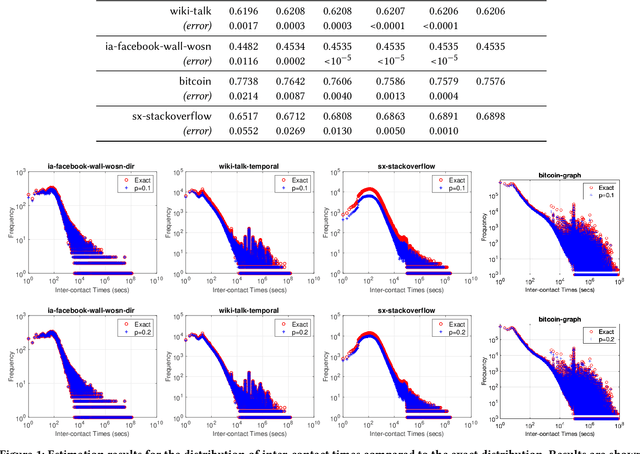
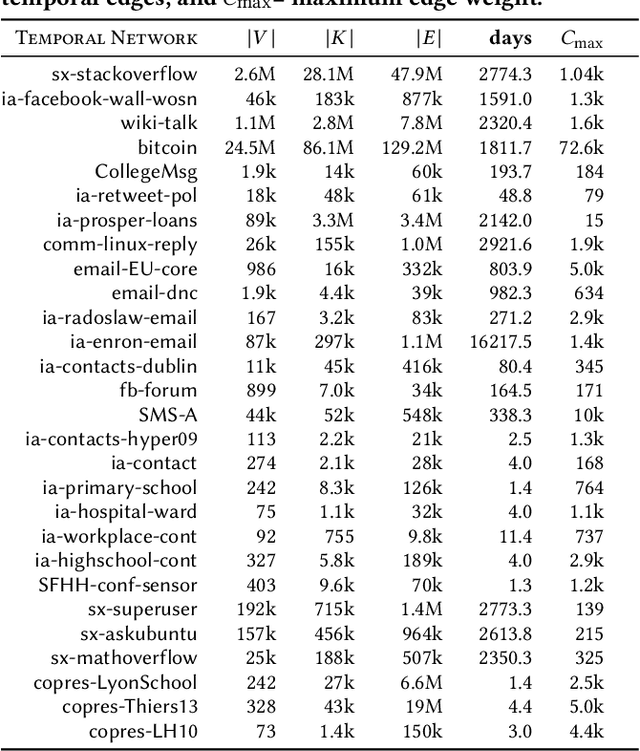

Temporal networks representing a stream of timestamped edges are seemingly ubiquitous in the real-world. However, the massive size and continuous nature of these networks make them fundamentally challenging to analyze and leverage for descriptive and predictive modeling tasks. In this work, we propose a general framework for temporal network sampling with unbiased estimation. We develop online, single-pass sampling algorithms and unbiased estimators for temporal network sampling. The proposed algorithms enable fast, accurate, and memory-efficient statistical estimation of temporal network patterns and properties. In addition, we propose a temporally decaying sampling algorithm with unbiased estimators for studying networks that evolve in continuous time, where the strength of links is a function of time, and the motif patterns are temporally-weighted. In contrast to the prior notion of a $\bigtriangleup t$-temporal motif, the proposed formulation and algorithms for counting temporally weighted motifs are useful for forecasting tasks in networks such as predicting future links, or a future time-series variable of nodes and links. Finally, extensive experiments on a variety of temporal networks from different domains demonstrate the effectiveness of the proposed algorithms.
Variational Graph Recurrent Neural Networks
Sep 09, 2019Ehsan Hajiramezanali, Arman Hasanzadeh, Nick Duffield, Krishna R Narayanan, Mingyuan Zhou, Xiaoning Qian
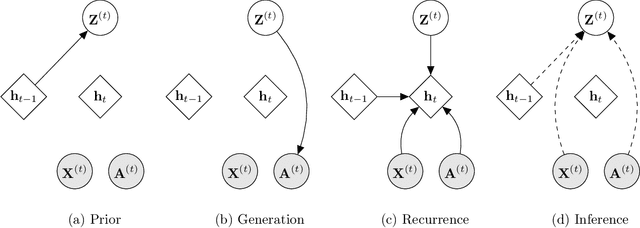
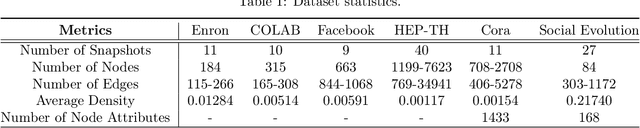
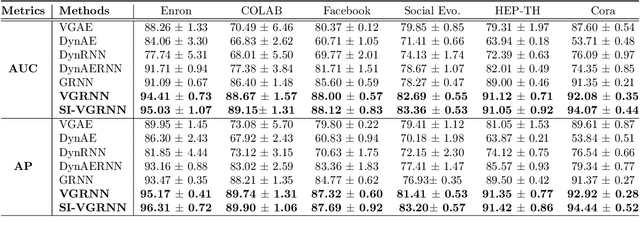
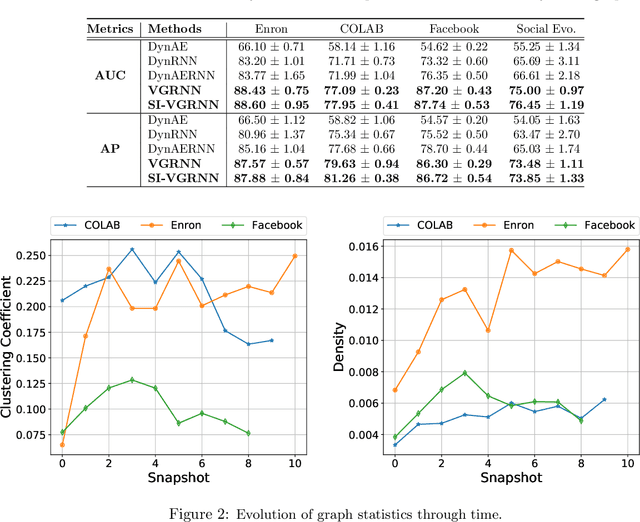
Representation learning over graph structured data has been mostly studied in static graph settings while efforts for modeling dynamic graphs are still scant. In this paper, we develop a novel hierarchical variational model that introduces additional latent random variables to jointly model the hidden states of a graph recurrent neural network (GRNN) to capture both topology and node attribute changes in dynamic graphs. We argue that the use of high-level latent random variables in this variational GRNN (VGRNN) can better capture potential variability observed in dynamic graphs as well as the uncertainty of node latent representation. With semi-implicit variational inference developed for this new VGRNN architecture (SI-VGRNN), we show that flexible non-Gaussian latent representations can further help dynamic graph analytic tasks. Our experiments with multiple real-world dynamic graph datasets demonstrate that SI-VGRNN and VGRNN consistently outperform the existing baseline and state-of-the-art methods by a significant margin in dynamic link prediction.
Semi-Implicit Graph Variational Auto-Encoders
Sep 07, 2019Arman Hasanzadeh, Ehsan Hajiramezanali, Nick Duffield, Krishna R. Narayanan, Mingyuan Zhou, Xiaoning Qian
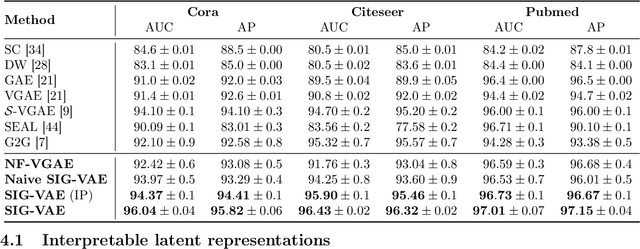
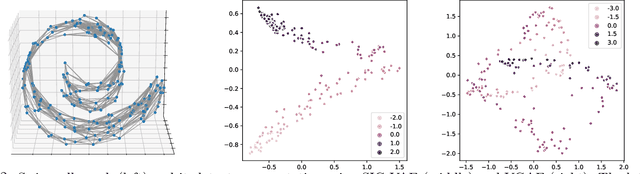
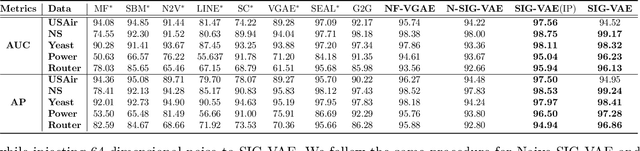
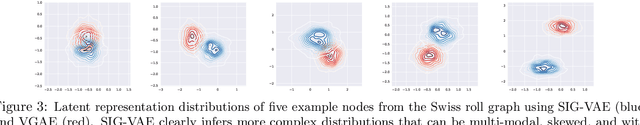
Semi-implicit graph variational auto-encoder (SIG-VAE) is proposed to expand the flexibility of variational graph auto-encoders (VGAE) to model graph data. SIG-VAE employs a hierarchical variational framework to enable neighboring node sharing for better generative modeling of graph dependency structure, together with a Bernoulli-Poisson link decoder. Not only does this hierarchical construction provide a more flexible generative graph model to better capture real-world graph properties, but also does SIG-VAE naturally lead to semi-implicit hierarchical variational inference that allows faithful modeling of implicit posteriors of given graph data, which may exhibit heavy tails, multiple modes, skewness, and rich dependency structures. Compared to VGAE, the derived graph latent representations by SIG-VAE are more interpretable, due to more expressive generative model and more faithful inference enabled by the flexible semi-implicit construction. Extensive experiments with a variety of graph data show that SIG-VAE significantly outperforms state-of-the-art methods on several different graph analytic tasks.
Network Shrinkage Estimation
Aug 02, 2019Nesreen K. Ahmed, Nick Duffield
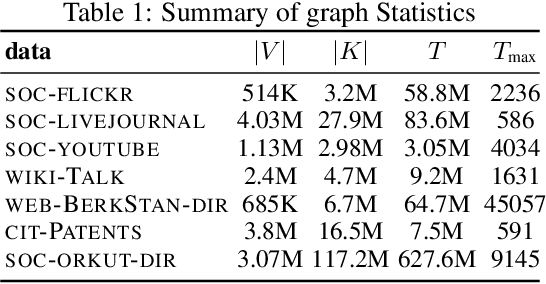
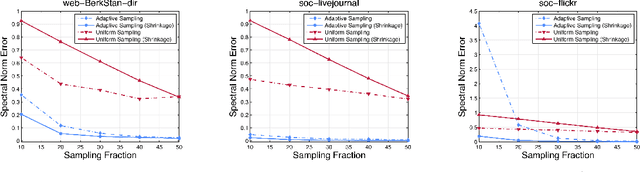
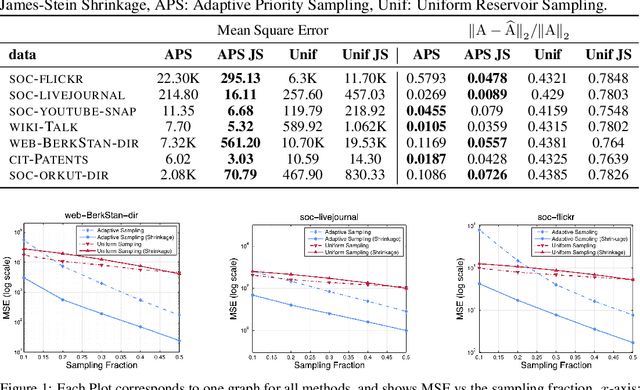
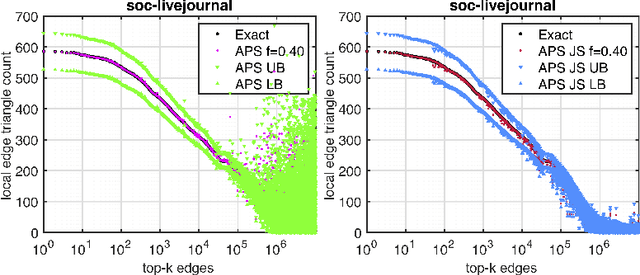
Networks are a natural representation of complex systems across the sciences, and higher-order dependencies are central to the understanding and modeling of these systems. However, in many practical applications such as online social networks, networks are massive, dynamic, and naturally streaming, where pairwise interactions become available one at a time in some arbitrary order. The massive size and streaming nature of these networks allow only partial observation, since it is infeasible to analyze the entire network. Under such scenarios, it is challenging to study the higher-order structural and connectivity patterns of streaming networks. In this work, we consider the fundamental problem of estimating the higher-order dependencies using adaptive sampling. We propose a novel adaptive, single-pass sampling framework and unbiased estimators for higher-order network analysis of large streaming networks. Our algorithms exploit adaptive techniques to identify edges that are highly informative for efficiently estimating the higher-order structure of streaming networks from small sample data. We also introduce a novel James-Stein-type shrinkage estimator to minimize the estimation error. Our approach is fully analytic with theoretical guarantees, computationally efficient, and can be incrementally updated in a streaming setting. Numerical experiments on large networks show that our approach is superior to baseline methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge