Single-Shared Network with Prior-Inspired Loss for Parameter-Efficient Multi-Modal Imaging Skin Lesion Classification
Mar 28, 2024Peng Tang, Tobias Lasser
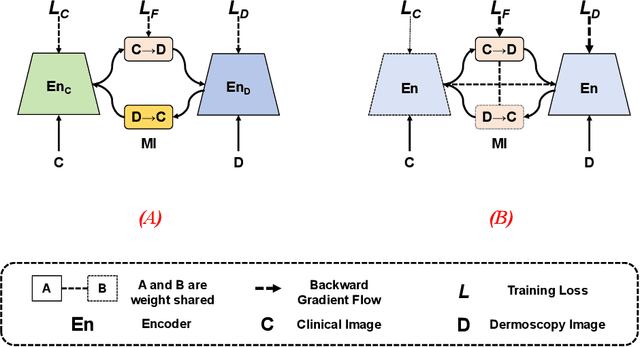
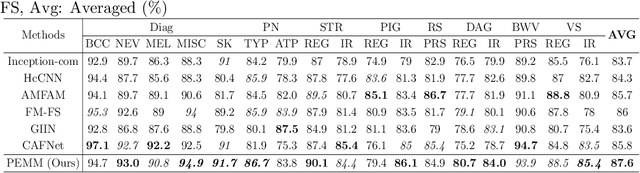
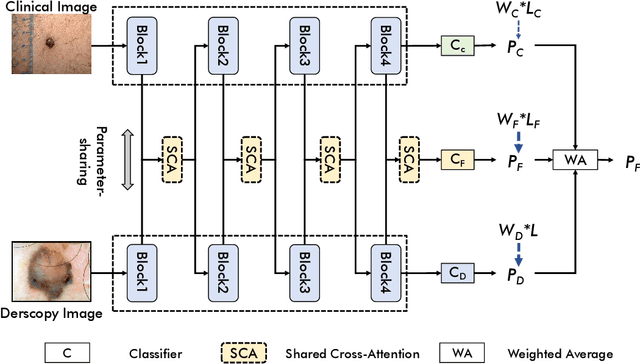
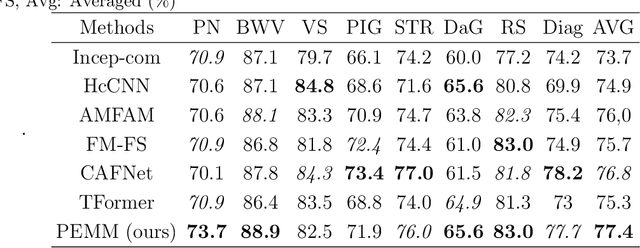
In this study, we introduce a multi-modal approach that efficiently integrates multi-scale clinical and dermoscopy features within a single network, thereby substantially reducing model parameters. The proposed method includes three novel fusion schemes. Firstly, unlike current methods that usually employ two individual models for for clinical and dermoscopy modalities, we verified that multimodal feature can be learned by sharing the parameters of encoder while leaving the individual modal-specific classifiers. Secondly, the shared cross-attention module can replace the individual one to efficiently interact between two modalities at multiple layers. Thirdly, different from current methods that equally optimize dermoscopy and clinical branches, inspired by prior knowledge that dermoscopy images play a more significant role than clinical images, we propose a novel biased loss. This loss guides the single-shared network to prioritize dermoscopy information over clinical information, implicitly learning a better joint feature representation for the modal-specific task. Extensive experiments on a well-recognized Seven-Point Checklist (SPC) dataset and a collected dataset demonstrate the effectiveness of our method on both CNN and Transformer structures. Furthermore, our method exhibits superiority in both accuracy and model parameters compared to currently advanced methods.
Synthesize Step-by-Step: Tools, Templates and LLMs as Data Generators for Reasoning-Based Chart VQA
Mar 28, 2024Zhuowan Li, Bhavan Jasani, Peng Tang, Shabnam Ghadar
Understanding data visualizations like charts and plots requires reasoning about both visual elements and numerics. Although strong in extractive questions, current chart visual question answering (chart VQA) models suffer on complex reasoning questions. In this work, we address the lack of reasoning ability by data augmentation. We leverage Large Language Models (LLMs), which have shown to have strong reasoning ability, as an automatic data annotator that generates question-answer annotations for chart images. The key innovation in our method lies in the Synthesize Step-by-Step strategy: our LLM-based data generator learns to decompose the complex question into step-by-step sub-questions (rationales), which are then used to derive the final answer using external tools, i.e. Python. This step-wise generation procedure is trained on synthetic data generated using a template-based QA generation pipeline. Experimental results highlight the significance of the proposed step-by-step generation. By training with the LLM-augmented data (LAMENDA), we significantly enhance the chart VQA models, achieving the state-of-the-art accuracy on the ChartQA and PlotQA datasets. In particular, our approach improves the accuracy of the previous state-of-the-art approach from 38% to 54% on the human-written questions in the ChartQA dataset, which needs strong reasoning. We hope our work underscores the potential of synthetic data and encourages further exploration of data augmentation using LLMs for reasoning-heavy tasks.
Federated Semi-supervised Learning for Medical Image Segmentation with intra-client and inter-client Consistency
Mar 19, 2024Yubin Zheng, Peng Tang, Tianjie Ju, Weidong Qiu, Bo Yan
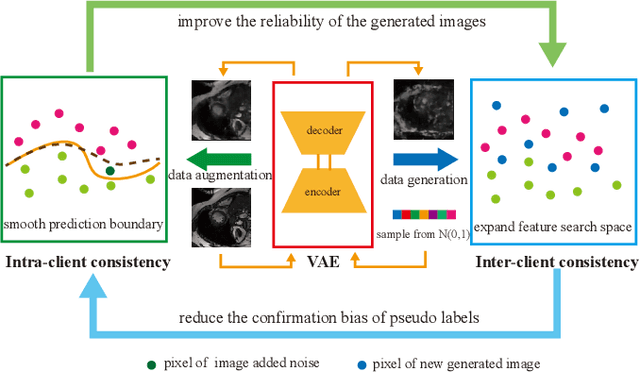
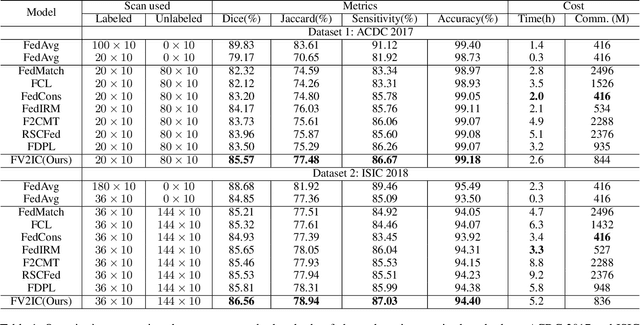
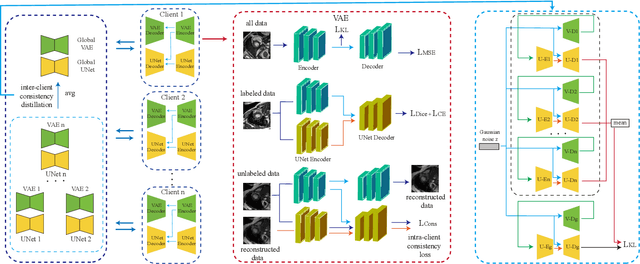
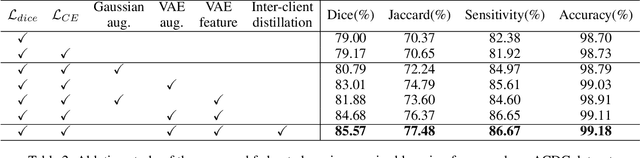
Medical image segmentation plays a vital role in clinic disease diagnosis and medical image analysis. However, labeling medical images for segmentation task is tough due to the indispensable domain expertise of radiologists. Furthermore, considering the privacy and sensitivity of medical images, it is impractical to build a centralized segmentation dataset from different medical institutions. Federated learning aims to train a shared model of isolated clients without local data exchange which aligns well with the scarcity and privacy characteristics of medical data. To solve the problem of labeling hard, many advanced semi-supervised methods have been proposed in a centralized data setting. As for federated learning, how to conduct semi-supervised learning under this distributed scenario is worth investigating. In this work, we propose a novel federated semi-supervised learning framework for medical image segmentation. The intra-client and inter-client consistency learning are introduced to smooth predictions at the data level and avoid confirmation bias of local models. They are achieved with the assistance of a Variational Autoencoder (VAE) trained collaboratively by clients. The added VAE model plays three roles: 1) extracting latent low-dimensional features of all labeled and unlabeled data; 2) performing a novel type of data augmentation in calculating intra-client consistency loss; 3) utilizing the generative ability of itself to conduct inter-client consistency distillation. The proposed framework is compared with other federated semi-supervised or self-supervised learning methods. The experimental results illustrate that our method outperforms the state-of-the-art method while avoiding a lot of computation and communication overhead.
SSDRec: Self-Augmented Sequence Denoising for Sequential Recommendation
Mar 07, 2024Chi Zhang, Qilong Han, Rui Chen, Xiangyu Zhao, Peng Tang, Hongtao Song
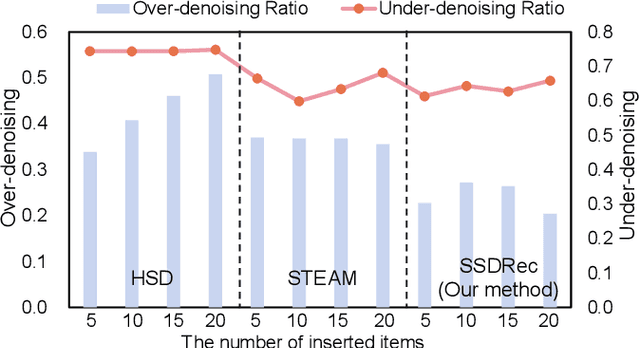
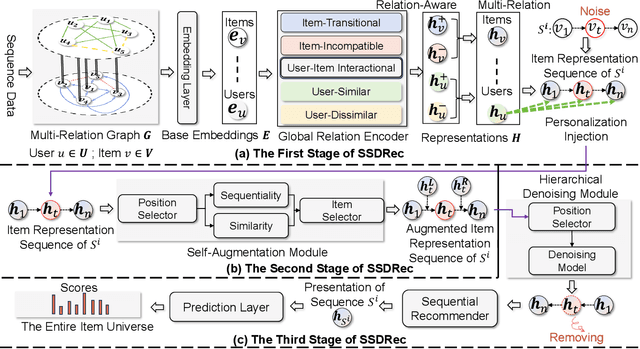
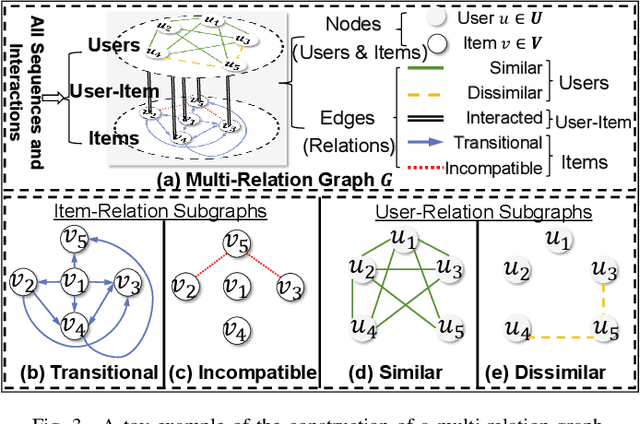
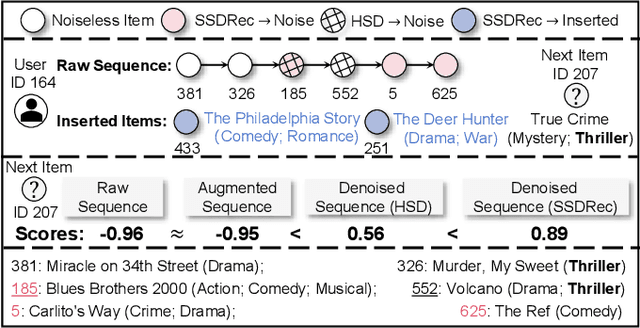
Traditional sequential recommendation methods assume that users' sequence data is clean enough to learn accurate sequence representations to reflect user preferences. In practice, users' sequences inevitably contain noise (e.g., accidental interactions), leading to incorrect reflections of user preferences. Consequently, some pioneer studies have explored modeling sequentiality and correlations in sequences to implicitly or explicitly reduce noise's influence. However, relying on only available intra-sequence information (i.e., sequentiality and correlations in a sequence) is insufficient and may result in over-denoising and under-denoising problems (OUPs), especially for short sequences. To improve reliability, we propose to augment sequences by inserting items before denoising. However, due to the data sparsity issue and computational costs, it is challenging to select proper items from the entire item universe to insert into proper positions in a target sequence. Motivated by the above observation, we propose a novel framework--Self-augmented Sequence Denoising for sequential Recommendation (SSDRec) with a three-stage learning paradigm to solve the above challenges. In the first stage, we empower SSDRec by a global relation encoder to learn multi-faceted inter-sequence relations in a data-driven manner. These relations serve as prior knowledge to guide subsequent stages. In the second stage, we devise a self-augmentation module to augment sequences to alleviate OUPs. Finally, we employ a hierarchical denoising module in the third stage to reduce the risk of false augmentations and pinpoint all noise in raw sequences. Extensive experiments on five real-world datasets demonstrate the superiority of \model over state-of-the-art denoising methods and its flexible applications to mainstream sequential recommendation models. The source code is available at https://github.com/zc-97/SSDRec.
Feature Norm Regularized Federated Learning: Transforming Skewed Distributions into Global Insights
Dec 12, 2023Ke Hu, WeiDong Qiu, Peng Tang
In the field of federated learning, addressing non-independent and identically distributed (non-i.i.d.) data remains a quintessential challenge for improving global model performance. This work introduces the Feature Norm Regularized Federated Learning (FNR-FL) algorithm, which uniquely incorporates class average feature norms to enhance model accuracy and convergence in non-i.i.d. scenarios. Our comprehensive analysis reveals that FNR-FL not only accelerates convergence but also significantly surpasses other contemporary federated learning algorithms in test accuracy, particularly under feature distribution skew scenarios. The novel modular design of FNR-FL facilitates seamless integration with existing federated learning frameworks, reinforcing its adaptability and potential for widespread application. We substantiate our claims through rigorous empirical evaluations, demonstrating FNR-FL's exceptional performance across various skewed data distributions. Relative to FedAvg, FNR-FL exhibits a substantial 66.24\% improvement in accuracy and a significant 11.40\% reduction in training time, underscoring its enhanced effectiveness and efficiency.
Joint-Individual Fusion Structure with Fusion Attention Module for Multi-Modal Skin Cancer Classification
Dec 07, 2023Peng Tang, Xintong Yan, Yang Nan, Xiaobin Hu, Bjoern H Menzee. Sebastian Krammer, Tobias Lasser
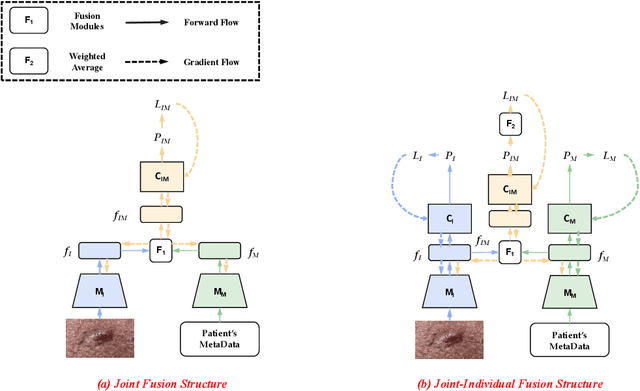
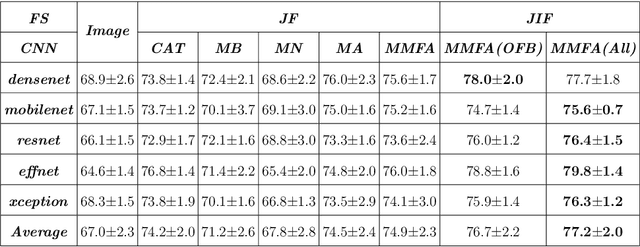
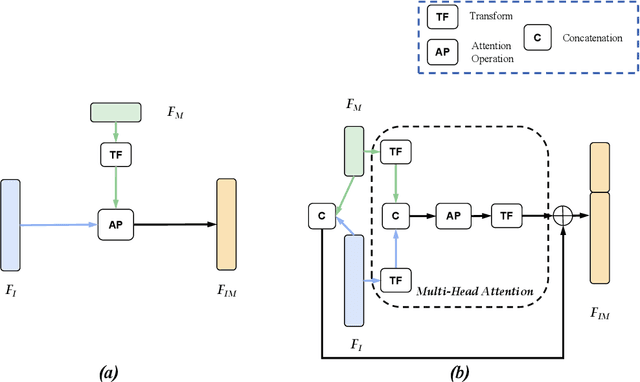
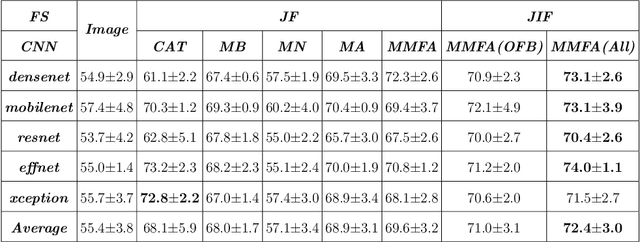
Most convolutional neural network (CNN) based methods for skin cancer classification obtain their results using only dermatological images. Although good classification results have been shown, more accurate results can be achieved by considering the patient's metadata, which is valuable clinical information for dermatologists. Current methods only use the simple joint fusion structure (FS) and fusion modules (FMs) for the multi-modal classification methods, there still is room to increase the accuracy by exploring more advanced FS and FM. Therefore, in this paper, we design a new fusion method that combines dermatological images (dermoscopy images or clinical images) and patient metadata for skin cancer classification from the perspectives of FS and FM. First, we propose a joint-individual fusion (JIF) structure that learns the shared features of multi-modality data and preserves specific features simultaneously. Second, we introduce a fusion attention (FA) module that enhances the most relevant image and metadata features based on both the self and mutual attention mechanism to support the decision-making pipeline. We compare the proposed JIF-MMFA method with other state-of-the-art fusion methods on three different public datasets. The results show that our JIF-MMFA method improves the classification results for all tested CNN backbones and performs better than the other fusion methods on the three public datasets, demonstrating our method's effectiveness and robustness
Multiple-Question Multiple-Answer Text-VQA
Nov 15, 2023Peng Tang, Srikar Appalaraju, R. Manmatha, Yusheng Xie, Vijay Mahadevan
We present Multiple-Question Multiple-Answer (MQMA), a novel approach to do text-VQA in encoder-decoder transformer models. The text-VQA task requires a model to answer a question by understanding multi-modal content: text (typically from OCR) and an associated image. To the best of our knowledge, almost all previous approaches for text-VQA process a single question and its associated content to predict a single answer. In order to answer multiple questions from the same image, each question and content are fed into the model multiple times. In contrast, our proposed MQMA approach takes multiple questions and content as input at the encoder and predicts multiple answers at the decoder in an auto-regressive manner at the same time. We make several novel architectural modifications to standard encoder-decoder transformers to support MQMA. We also propose a novel MQMA denoising pre-training task which is designed to teach the model to align and delineate multiple questions and content with associated answers. MQMA pre-trained model achieves state-of-the-art results on multiple text-VQA datasets, each with strong baselines. Specifically, on OCR-VQA (+2.5%), TextVQA (+1.4%), ST-VQA (+0.6%), DocVQA (+1.1%) absolute improvements over the previous state-of-the-art approaches.
DEED: Dynamic Early Exit on Decoder for Accelerating Encoder-Decoder Transformer Models
Nov 15, 2023Peng Tang, Pengkai Zhu, Tian Li, Srikar Appalaraju, Vijay Mahadevan, R. Manmatha
Encoder-decoder transformer models have achieved great success on various vision-language (VL) tasks, but they suffer from high inference latency. Typically, the decoder takes up most of the latency because of the auto-regressive decoding. To accelerate the inference, we propose an approach of performing Dynamic Early Exit on Decoder (DEED). We build a multi-exit encoder-decoder transformer model which is trained with deep supervision so that each of its decoder layers is capable of generating plausible predictions. In addition, we leverage simple yet practical techniques, including shared generation head and adaptation modules, to keep accuracy when exiting at shallow decoder layers. Based on the multi-exit model, we perform step-level dynamic early exit during inference, where the model may decide to use fewer decoder layers based on its confidence of the current layer at each individual decoding step. Considering different number of decoder layers may be used at different decoding steps, we compute deeper-layer decoder features of previous decoding steps just-in-time, which ensures the features from different decoding steps are semantically aligned. We evaluate our approach with two state-of-the-art encoder-decoder transformer models on various VL tasks. We show our approach can reduce overall inference latency by 30%-60% with comparable or even higher accuracy compared to baselines.
SR-R$^2$KAC: Improving Single Image Defocus Deblurring
Jul 30, 2023Peng Tang, Zhiqiang Xu, Pengfei Wei, Xiaobin Hu, Peilin Zhao, Xin Cao, Chunlai Zhou, Tobias Lasser
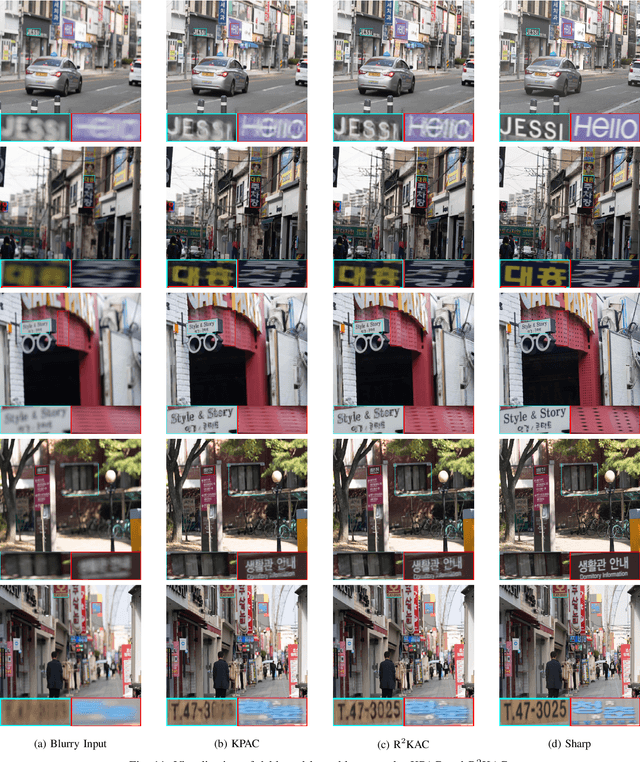
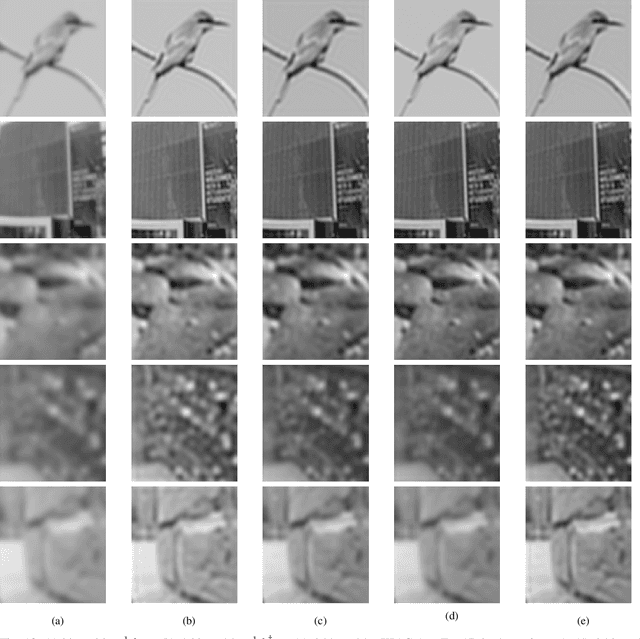

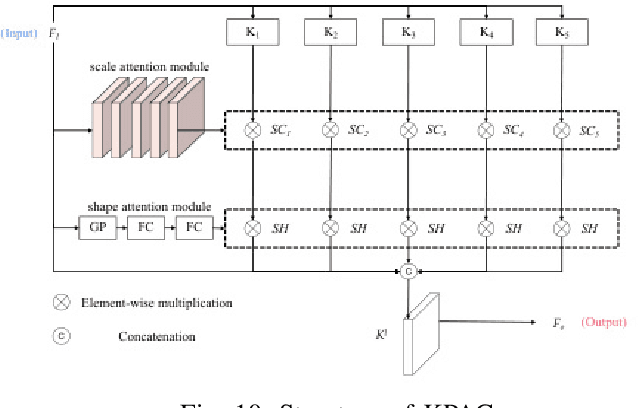


We propose an efficient deep learning method for single image defocus deblurring (SIDD) by further exploring inverse kernel properties. Although the current inverse kernel method, i.e., kernel-sharing parallel atrous convolution (KPAC), can address spatially varying defocus blurs, it has difficulty in handling large blurs of this kind. To tackle this issue, we propose a Residual and Recursive Kernel-sharing Atrous Convolution (R$^2$KAC). R$^2$KAC builds on a significant observation of inverse kernels, that is, successive use of inverse-kernel-based deconvolutions with fixed size helps remove unexpected large blurs but produces ringing artifacts. Specifically, on top of kernel-sharing atrous convolutions used to simulate multi-scale inverse kernels, R$^2$KAC applies atrous convolutions recursively to simulate a large inverse kernel. Specifically, on top of kernel-sharing atrous convolutions, R$^2$KAC stacks atrous convolutions recursively to simulate a large inverse kernel. To further alleviate the contingent effect of recursive stacking, i.e., ringing artifacts, we add identity shortcuts between atrous convolutions to simulate residual deconvolutions. Lastly, a scale recurrent module is embedded in the R$^2$KAC network, leading to SR-R$^2$KAC, so that multi-scale information from coarse to fine is exploited to progressively remove the spatially varying defocus blurs. Extensive experimental results show that our method achieves the state-of-the-art performance.
Graph-Ensemble Learning Model for Multi-label Skin Lesion Classification using Dermoscopy and Clinical Images
Jul 04, 2023Peng Tang, Yang Nan, Tobias Lasser
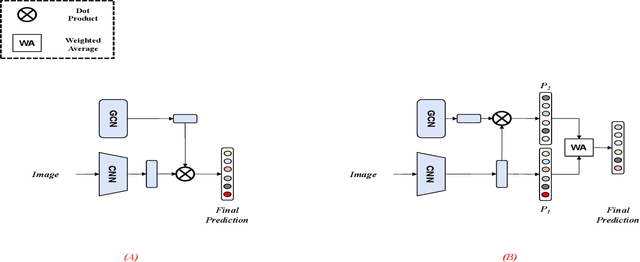
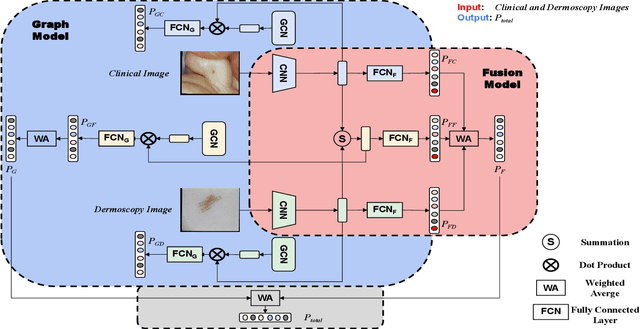
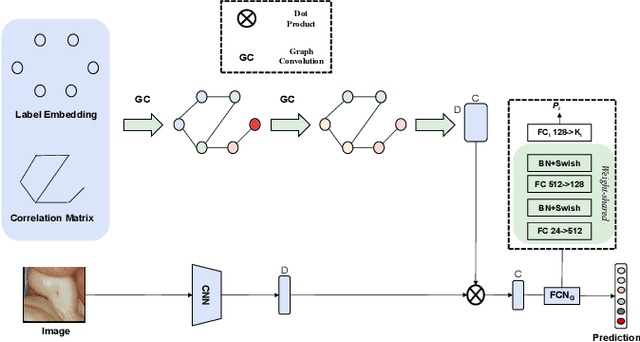
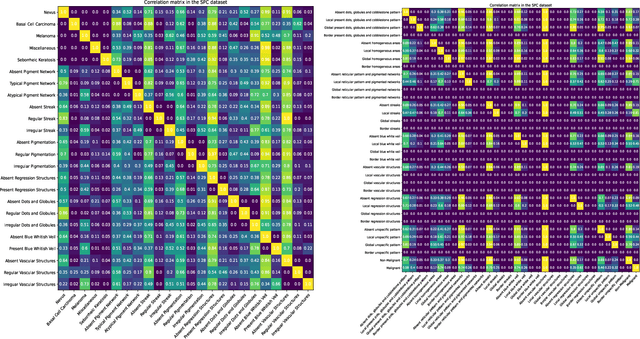
Many skin lesion analysis (SLA) methods recently focused on developing a multi-modal-based multi-label classification method due to two factors. The first is multi-modal data, i.e., clinical and dermoscopy images, which can provide complementary information to obtain more accurate results than single-modal data. The second one is that multi-label classification, i.e., seven-point checklist (SPC) criteria as an auxiliary classification task can not only boost the diagnostic accuracy of melanoma in the deep learning (DL) pipeline but also provide more useful functions to the clinical doctor as it is commonly used in clinical dermatologist's diagnosis. However, most methods only focus on designing a better module for multi-modal data fusion; few methods explore utilizing the label correlation between SPC and skin disease for performance improvement. This study fills the gap that introduces a Graph Convolution Network (GCN) to exploit prior co-occurrence between each category as a correlation matrix into the DL model for the multi-label classification. However, directly applying GCN degraded the performances in our experiments; we attribute this to the weak generalization ability of GCN in the scenario of insufficient statistical samples of medical data. We tackle this issue by proposing a Graph-Ensemble Learning Model (GELN) that views the prediction from GCN as complementary information of the predictions from the fusion model and adaptively fuses them by a weighted averaging scheme, which can utilize the valuable information from GCN while avoiding its negative influences as much as possible. To evaluate our method, we conduct experiments on public datasets. The results illustrate that our GELN can consistently improve the classification performance on different datasets and that the proposed method can achieve state-of-the-art performance in SPC and diagnosis classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge