Personalized Forgetting Mechanism with Concept-Driven Knowledge Tracing
Apr 18, 2024Shanshan Wang, Ying Hu, Xun Yang, Zhongzhou Zhang, Keyang Wang, Xingyi Zhang
Knowledge Tracing (KT) aims to trace changes in students' knowledge states throughout their entire learning process by analyzing their historical learning data and predicting their future learning performance. Existing forgetting curve theory based knowledge tracing models only consider the general forgetting caused by time intervals, ignoring the individualization of students and the causal relationship of the forgetting process. To address these problems, we propose a Concept-driven Personalized Forgetting knowledge tracing model (CPF) which integrates hierarchical relationships between knowledge concepts and incorporates students' personalized cognitive abilities. First, we integrate the students' personalized capabilities into both the learning and forgetting processes to explicitly distinguish students' individual learning gains and forgetting rates according to their cognitive abilities. Second, we take into account the hierarchical relationships between knowledge points and design a precursor-successor knowledge concept matrix to simulate the causal relationship in the forgetting process, while also integrating the potential impact of forgetting prior knowledge points on subsequent ones. The proposed personalized forgetting mechanism can not only be applied to the learning of specifc knowledge concepts but also the life-long learning process. Extensive experimental results on three public datasets show that our CPF outperforms current forgetting curve theory based methods in predicting student performance, demonstrating CPF can better simulate changes in students' knowledge status through the personalized forgetting mechanism.
CSR-dMRI: Continuous Super-Resolution of Diffusion MRI with Anatomical Structure-assisted Implicit Neural Representation Learning
Apr 04, 2024Ruoyou Wu, Jian Cheng, Cheng Li, Juan Zou, Jing Yang, Wenxin Fan, Shanshan Wang
Deep learning-based dMRI super-resolution methods can effectively enhance image resolution by leveraging the learning capabilities of neural networks on large datasets. However, these methods tend to learn a fixed scale mapping between low-resolution (LR) and high-resolution (HR) images, overlooking the need for radiologists to scale the images at arbitrary resolutions. Moreover, the pixel-wise loss in the image domain tends to generate over-smoothed results, losing fine textures and edge information. To address these issues, we propose a novel continuous super-resolution of dMRI with anatomical structure-assisted implicit neural representation learning method, called CSR-dMRI. Specifically, the CSR-dMRI model consists of two components. The first is the latent feature extractor, which primarily extracts latent space feature maps from LR dMRI and anatomical images while learning structural prior information from the anatomical images. The second is the implicit function network, which utilizes voxel coordinates and latent feature vectors to generate voxel intensities at corresponding positions. Additionally, a frequency-domain-based loss is introduced to preserve the structural and texture information, further enhancing the image quality. Extensive experiments on the publicly available HCP dataset validate the effectiveness of our approach. Furthermore, our method demonstrates superior generalization capability and can be applied to arbitrary-scale super-resolution, including non-integer scale factors, expanding its applicability beyond conventional approaches.
Automating Vessel Segmentation in the Heart and Brain: A Trend to Develop Multi-Modality and Label-Efficient Deep Learning Techniques
Apr 02, 2024Nazik Elsayed, Yousuf Babiker M. Osman, Cheng Li, Jiong Zhang, Shanshan Wang
Cardio-cerebrovascular diseases are the leading causes of mortality worldwide, whose accurate blood vessel segmentation is significant for both scientific research and clinical usage. However, segmenting cardio-cerebrovascular structures from medical images is very challenging due to the presence of thin or blurred vascular shapes, imbalanced distribution of vessel and non-vessel pixels, and interference from imaging artifacts. These difficulties make manual or semi-manual segmentation methods highly time-consuming, labor-intensive, and prone to errors with interobserver variability, where different experts may produce different segmentations from a variety of modalities. Consequently, there is a growing interest in developing automated algorithms. This paper provides an up-to-date survey of deep learning techniques, for cardio-cerebrovascular segmentation. It analyzes the research landscape, surveys recent approaches, and discusses challenges such as the scarcity of accurately annotated data and variability. This paper also illustrates the urgent needs for developing multi-modality label-efficient deep learning techniques. To the best of our knowledge, this paper is the first comprehensive survey of deep learning approaches that effectively segment vessels in both the heart and brain. It aims to advance automated segmentation techniques for cardio-cerebrovascular diseases, benefiting researchers and healthcare professionals.
Enhancing Out-of-Distribution Detection with Multitesting-based Layer-wise Feature Fusion
Mar 16, 2024Jiawei Li, Sitong Li, Shanshan Wang, Yicheng Zeng, Falong Tan, Chuanlong Xie
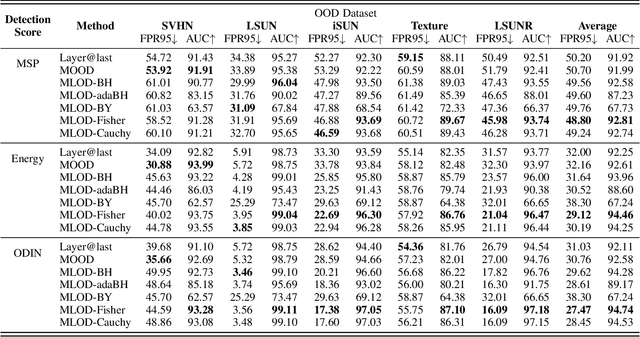

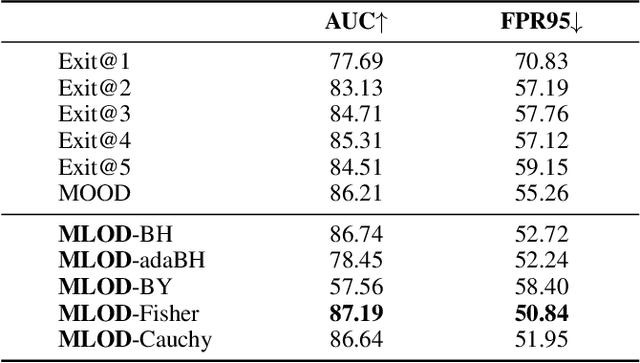
Deploying machine learning in open environments presents the challenge of encountering diverse test inputs that differ significantly from the training data. These out-of-distribution samples may exhibit shifts in local or global features compared to the training distribution. The machine learning (ML) community has responded with a number of methods aimed at distinguishing anomalous inputs from original training data. However, the majority of previous studies have primarily focused on the output layer or penultimate layer of pre-trained deep neural networks. In this paper, we propose a novel framework, Multitesting-based Layer-wise Out-of-Distribution (OOD) Detection (MLOD), to identify distributional shifts in test samples at different levels of features through rigorous multiple testing procedure. Our approach distinguishes itself from existing methods as it does not require modifying the structure or fine-tuning of the pre-trained classifier. Through extensive experiments, we demonstrate that our proposed framework can seamlessly integrate with any existing distance-based inspection method while efficiently utilizing feature extractors of varying depths. Our scheme effectively enhances the performance of out-of-distribution detection when compared to baseline methods. In particular, MLOD-Fisher achieves superior performance in general. When trained using KNN on CIFAR10, MLOD-Fisher significantly lowers the false positive rate (FPR) from 24.09% to 7.47% on average compared to merely utilizing the features of the last layer.
Swin-UMamba: Mamba-based UNet with ImageNet-based pretraining
Feb 05, 2024Jiarun Liu, Hao Yang, Hong-Yu Zhou, Yan Xi, Lequan Yu, Yizhou Yu, Yong Liang, Guangming Shi, Shaoting Zhang, Hairong Zheng, Shanshan Wang
Accurate medical image segmentation demands the integration of multi-scale information, spanning from local features to global dependencies. However, it is challenging for existing methods to model long-range global information, where convolutional neural networks (CNNs) are constrained by their local receptive fields, and vision transformers (ViTs) suffer from high quadratic complexity of their attention mechanism. Recently, Mamba-based models have gained great attention for their impressive ability in long sequence modeling. Several studies have demonstrated that these models can outperform popular vision models in various tasks, offering higher accuracy, lower memory consumption, and less computational burden. However, existing Mamba-based models are mostly trained from scratch and do not explore the power of pretraining, which has been proven to be quite effective for data-efficient medical image analysis. This paper introduces a novel Mamba-based model, Swin-UMamba, designed specifically for medical image segmentation tasks, leveraging the advantages of ImageNet-based pretraining. Our experimental results reveal the vital role of ImageNet-based training in enhancing the performance of Mamba-based models. Swin-UMamba demonstrates superior performance with a large margin compared to CNNs, ViTs, and latest Mamba-based models. Notably, on AbdomenMRI, Encoscopy, and Microscopy datasets, Swin-UMamba outperforms its closest counterpart U-Mamba by an average score of 3.58%. The code and models of Swin-UMamba are publicly available at: https://github.com/JiarunLiu/Swin-UMamba
Positive and negative sampling strategies for self-supervised learning on audio-video data
Feb 05, 2024Shanshan Wang, Soumya Tripathy, Toni Heittola, Annamaria Mesaros
In Self-Supervised Learning (SSL), Audio-Visual Correspondence (AVC) is a popular task to learn deep audio and video features from large unlabeled datasets. The key step in AVC is to randomly sample audio and video clips from the dataset and learn to minimize the feature distance between the positive pairs (corresponding audio-video pair) while maximizing the distance between the negative pairs (non-corresponding audio-video pairs). The learnt features are shown to be effective on various downstream tasks. However, these methods achieve subpar performance when the size of the dataset is rather small. In this paper, we investigate the effect of utilizing class label information in the AVC feature learning task. We modified various positive and negative data sampling techniques of SSL based on class label information to investigate the effect on the feature quality. We propose a new sampling approach which we call soft-positive sampling, where the positive pair for one audio sample is not from the exact corresponding video, but from a video of the same class. Experimental results suggest that when the dataset size is small in SSL setup, features learnt through the soft-positive sampling method significantly outperform those from the traditional SSL sampling approaches. This trend holds in both in-domain and out-of-domain downstream tasks, and even outperforms supervised classification. Finally, experiments show that class label information can easily be obtained using a publicly available classifier network and then can be used to boost the SSL performance without adding extra data annotation burden.
Knowledge-driven deep learning for fast MR imaging: undersampled MR image reconstruction from supervised to un-supervised learning
Feb 05, 2024Shanshan Wang, Ruoyou Wu, Sen Jia, Alou Diakite, Cheng Li, Qiegen Liu, Leslie Ying
Deep learning (DL) has emerged as a leading approach in accelerating MR imaging. It employs deep neural networks to extract knowledge from available datasets and then applies the trained networks to reconstruct accurate images from limited measurements. Unlike natural image restoration problems, MR imaging involves physics-based imaging processes, unique data properties, and diverse imaging tasks. This domain knowledge needs to be integrated with data-driven approaches. Our review will introduce the significant challenges faced by such knowledge-driven DL approaches in the context of fast MR imaging along with several notable solutions, which include learning neural networks and addressing different imaging application scenarios. The traits and trends of these techniques have also been given which have shifted from supervised learning to semi-supervised learning, and finally, to unsupervised learning methods. In addition, MR vendors' choices of DL reconstruction have been provided along with some discussions on open questions and future directions, which are critical for the reliable imaging systems.
Enhancing the vision-language foundation model with key semantic knowledge-emphasized report refinement
Jan 21, 2024Cheng Li, Weijian Huang, Hao Yang, Jiarun Liu, Shanshan Wang
Recently, vision-language representation learning has made remarkable advancements in building up medical foundation models, holding immense potential for transforming the landscape of clinical research and medical care. The underlying hypothesis is that the rich knowledge embedded in radiology reports can effectively assist and guide the learning process, reducing the need for additional labels. However, these reports tend to be complex and sometimes even consist of redundant descriptions that make the representation learning too challenging to capture the key semantic information. This paper develops a novel iterative vision-language representation learning framework by proposing a key semantic knowledge-emphasized report refinement method. Particularly, raw radiology reports are refined to highlight the key information according to a constructed clinical dictionary and two model-optimized knowledge-enhancement metrics. The iterative framework is designed to progressively learn, starting from gaining a general understanding of the patient's condition based on raw reports and gradually refines and extracts critical information essential to the fine-grained analysis tasks. The effectiveness of the proposed framework is validated on various downstream medical image analysis tasks, including disease classification, region-of-interest segmentation, and phrase grounding. Our framework surpasses seven state-of-the-art methods in both fine-tuning and zero-shot settings, demonstrating its encouraging potential for different clinical applications.
Generalizable vision-language pre-training for annotation-free pathology localization
Jan 04, 2024Hao Yang, Hong-Yu Zhou, Cheng Li, Weijian Huang, Jiarun Liu, Shanshan Wang
Locating pathologies automatically from medical images aids the understanding of the emergence and progression of diseases, and such an ability can significantly benefit clinical diagnostics. However, existing deep learning models heavily rely on expert annotations and lack generalization capabilities in open clinical environments. In this study, we present a generalizable vision-language pre-training model for Annotation-Free pathology Localization (AFLoc). The core strength of AFLoc lies in its image annotation-free multi-level semantic structure-based contrastive learning, which comprehensively aligns multi-granularity medical concepts from reports with abundant image features, to adapt to the diverse expressions of observed and emerging unseen pathologies. We conducted extensive experimental validation across 4 distinct external datasets, encompassing 11 types of chest pathologies, to verify its generalization ability. The results demonstrate that AFLoc surpasses 6 state-of-the-art methods and even outperforms the human benchmark in locating 5 different pathologies, underscoring its suitability for complex clinical environments.
AID-DTI: Accelerating High-fidelity Diffusion Tensor Imaging with Detail-Preserving Model-based Deep Learning
Jan 03, 2024Wenxin Fan, Jian Cheng, Cheng Li, Xinrui Ma, Jing Yang, Juan Zou, Ruoyou Wu, Qiegen Liu, Shanshan Wang
Deep learning has shown great potential in accelerating diffusion tensor imaging (DTI). Nevertheless, existing methods tend to suffer from Rician noise and detail loss in reconstructing the DTI-derived parametric maps especially when sparsely sampled q-space data are used. This paper proposes a novel method, AID-DTI (Accelerating hIgh fiDelity Diffusion Tensor Imaging), to facilitate fast and accurate DTI with only six measurements. AID-DTI is equipped with a newly designed Singular Value Decomposition (SVD)-based regularizer, which can effectively capture fine details while suppressing noise during network training. Experimental results on Human Connectome Project (HCP) data consistently demonstrate that the proposed method estimates DTI parameter maps with fine-grained details and outperforms three state-of-the-art methods both quantitatively and qualitatively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge