Revisiting Cephalometric Landmark Detection from the view of Human Pose Estimation with Lightweight Super-Resolution Head
Sep 29, 2023Qian Wu, Si Yong Yeo, Yufei Chen, Jun Liu
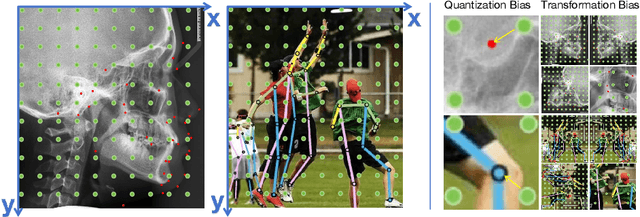

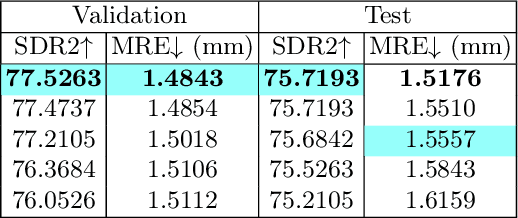
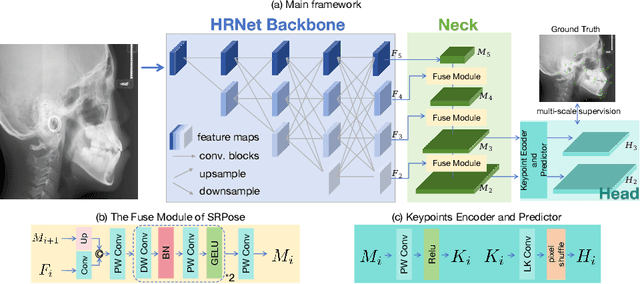
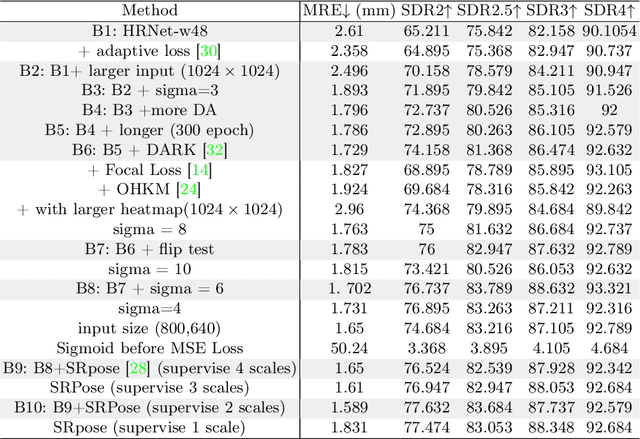
Accurate localization of cephalometric landmarks holds great importance in the fields of orthodontics and orthognathics due to its potential for automating key point labeling. In the context of landmark detection, particularly in cephalometrics, it has been observed that existing methods often lack standardized pipelines and well-designed bias reduction processes, which significantly impact their performance. In this paper, we revisit a related task, human pose estimation (HPE), which shares numerous similarities with cephalometric landmark detection (CLD), and emphasize the potential for transferring techniques from the former field to benefit the latter. Motivated by this insight, we have developed a robust and adaptable benchmark based on the well-established HPE codebase known as MMPose. This benchmark can serve as a dependable baseline for achieving exceptional CLD performance. Furthermore, we introduce an upscaling design within the framework to further enhance performance. This enhancement involves the incorporation of a lightweight and efficient super-resolution module, which generates heatmap predictions on high-resolution features and leads to further performance refinement, benefiting from its ability to reduce quantization bias. In the MICCAI CLDetection2023 challenge, our method achieves 1st place ranking on three metrics and 3rd place on the remaining one. The code for our method is available at https://github.com/5k5000/CLdetection2023.
Animal Kingdom: A Large and Diverse Dataset for Animal Behavior Understanding
Apr 18, 2022Xun Long Ng, Kian Eng Ong, Qichen Zheng, Yun Ni, Si Yong Yeo, Jun Liu
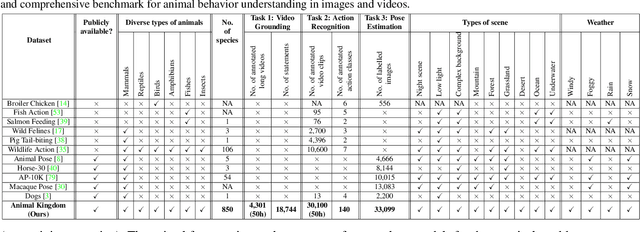
Understanding animals' behaviors is significant for a wide range of applications. However, existing animal behavior datasets have limitations in multiple aspects, including limited numbers of animal classes, data samples and provided tasks, and also limited variations in environmental conditions and viewpoints. To address these limitations, we create a large and diverse dataset, Animal Kingdom, that provides multiple annotated tasks to enable a more thorough understanding of natural animal behaviors. The wild animal footages used in our dataset record different times of the day in extensive range of environments containing variations in backgrounds, viewpoints, illumination and weather conditions. More specifically, our dataset contains 50 hours of annotated videos to localize relevant animal behavior segments in long videos for the video grounding task, 30K video sequences for the fine-grained multi-label action recognition task, and 33K frames for the pose estimation task, which correspond to a diverse range of animals with 850 species across 6 major animal classes. Such a challenging and comprehensive dataset shall be able to facilitate the community to develop, adapt, and evaluate various types of advanced methods for animal behavior analysis. Moreover, we propose a Collaborative Action Recognition (CARe) model that learns general and specific features for action recognition with unseen new animals. This method achieves promising performance in our experiments. Our dataset can be found at https://sutdcv.github.io/Animal-Kingdom.
A Novel Multi-task Deep Learning Model for Skin Lesion Segmentation and Classification
Mar 03, 2017Xulei Yang, Zeng Zeng, Si Yong Yeo, Colin Tan, Hong Liang Tey, Yi Su
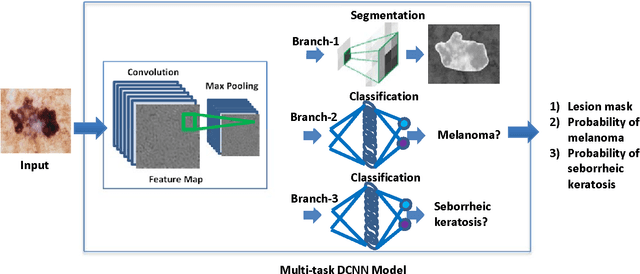

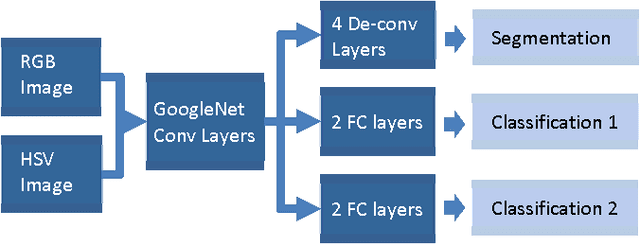

In this study, a multi-task deep neural network is proposed for skin lesion analysis. The proposed multi-task learning model solves different tasks (e.g., lesion segmentation and two independent binary lesion classifications) at the same time by exploiting commonalities and differences across tasks. This results in improved learning efficiency and potential prediction accuracy for the task-specific models, when compared to training the individual models separately. The proposed multi-task deep learning model is trained and evaluated on the dermoscopic image sets from the International Skin Imaging Collaboration (ISIC) 2017 Challenge - Skin Lesion Analysis towards Melanoma Detection, which consists of 2000 training samples and 150 evaluation samples. The experimental results show that the proposed multi-task deep learning model achieves promising performances on skin lesion segmentation and classification. The average value of Jaccard index for lesion segmentation is 0.724, while the average values of area under the receiver operating characteristic curve (AUC) on two individual lesion classifications are 0.880 and 0.972, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge