Graph Neural Networks for Protein-Protein Interactions -- A Short Survey
Apr 16, 2024Mingda Xu, Peisheng Qian, Ziyuan Zhao, Zeng Zeng, Jianguo Chen, Weide Liu, Xulei Yang
Protein-protein interactions (PPIs) play key roles in a broad range of biological processes. Numerous strategies have been proposed for predicting PPIs, and among them, graph-based methods have demonstrated promising outcomes owing to the inherent graph structure of PPI networks. This paper reviews various graph-based methodologies, and discusses their applications in PPI prediction. We classify these approaches into two primary groups based on their model structures. The first category employs Graph Neural Networks (GNN) or Graph Convolutional Networks (GCN), while the second category utilizes Graph Attention Networks (GAT), Graph Auto-Encoders and Graph-BERT. We highlight the distinctive methodologies of each approach in managing the graph-structured data inherent in PPI networks and anticipate future research directions in this domain.
Learning Intra-view and Cross-view Geometric Knowledge for Stereo Matching
Mar 06, 2024Rui Gong, Weide Liu, Zaiwang Gu, Xulei Yang, Jun Cheng
Geometric knowledge has been shown to be beneficial for the stereo matching task. However, prior attempts to integrate geometric insights into stereo matching algorithms have largely focused on geometric knowledge from single images while crucial cross-view factors such as occlusion and matching uniqueness have been overlooked. To address this gap, we propose a novel Intra-view and Cross-view Geometric knowledge learning Network (ICGNet), specifically crafted to assimilate both intra-view and cross-view geometric knowledge. ICGNet harnesses the power of interest points to serve as a channel for intra-view geometric understanding. Simultaneously, it employs the correspondences among these points to capture cross-view geometric relationships. This dual incorporation empowers the proposed ICGNet to leverage both intra-view and cross-view geometric knowledge in its learning process, substantially improving its ability to estimate disparities. Our extensive experiments demonstrate the superiority of the ICGNet over contemporary leading models.
Learn to Optimize Denoising Scores for 3D Generation: A Unified and Improved Diffusion Prior on NeRF and 3D Gaussian Splatting
Dec 08, 2023Xiaofeng Yang, Yiwen Chen, Cheng Chen, Chi Zhang, Yi Xu, Xulei Yang, Fayao Liu, Guosheng Lin
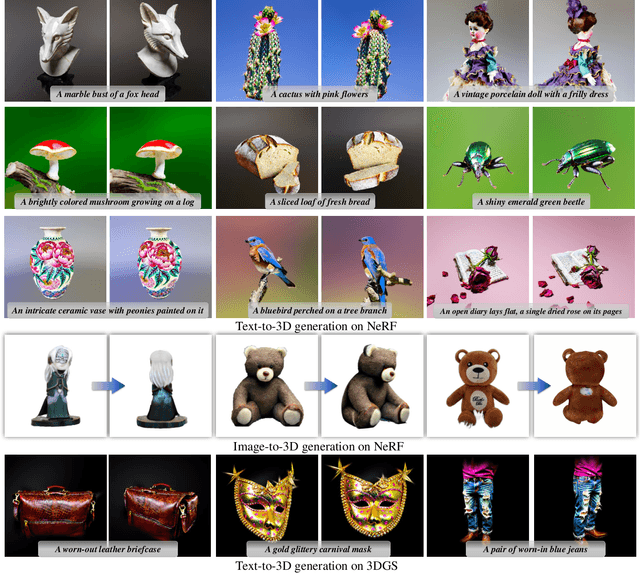
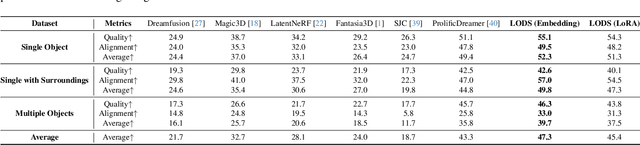
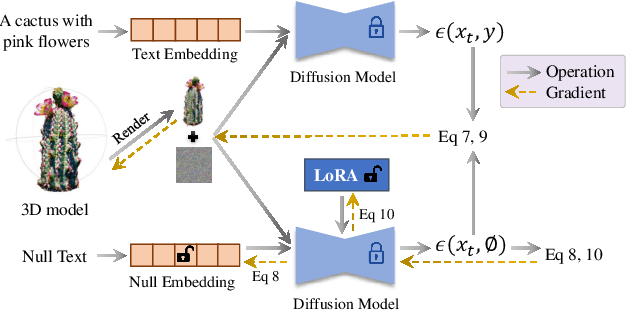
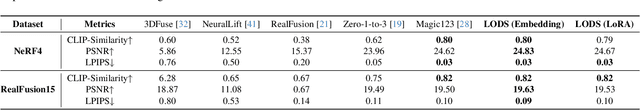
We propose a unified framework aimed at enhancing the diffusion priors for 3D generation tasks. Despite the critical importance of these tasks, existing methodologies often struggle to generate high-caliber results. We begin by examining the inherent limitations in previous diffusion priors. We identify a divergence between the diffusion priors and the training procedures of diffusion models that substantially impairs the quality of 3D generation. To address this issue, we propose a novel, unified framework that iteratively optimizes both the 3D model and the diffusion prior. Leveraging the different learnable parameters of the diffusion prior, our approach offers multiple configurations, affording various trade-offs between performance and implementation complexity. Notably, our experimental results demonstrate that our method markedly surpasses existing techniques, establishing new state-of-the-art in the realm of text-to-3D generation. Furthermore, our approach exhibits impressive performance on both NeRF and the newly introduced 3D Gaussian Splatting backbones. Additionally, our framework yields insightful contributions to the understanding of recent score distillation methods, such as the VSD and DDS loss.
SemiGNN-PPI: Self-Ensembling Multi-Graph Neural Network for Efficient and Generalizable Protein-Protein Interaction Prediction
May 15, 2023Ziyuan Zhao, Peisheng Qian, Xulei Yang, Zeng Zeng, Cuntai Guan, Wai Leong Tam, Xiaoli Li
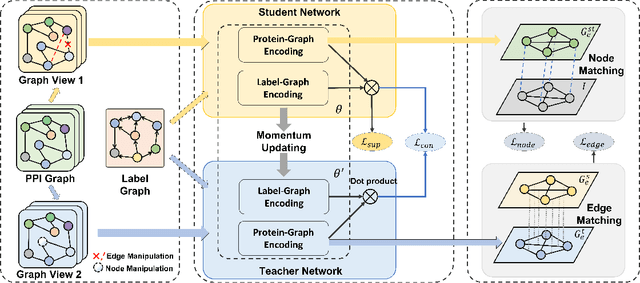
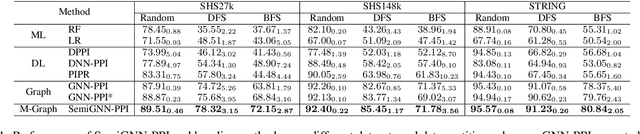
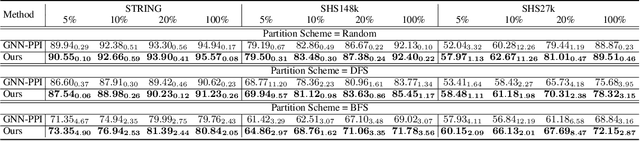
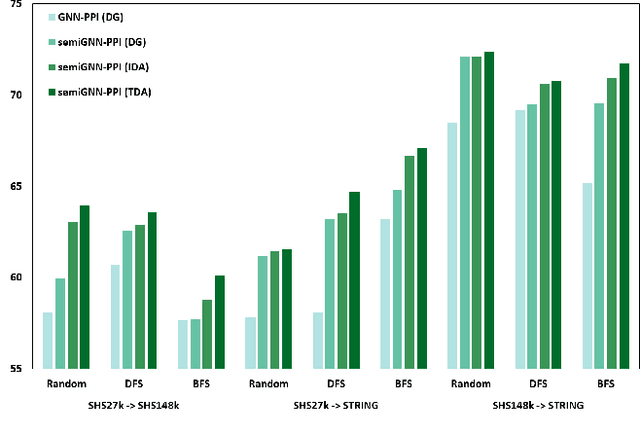
Protein-protein interactions (PPIs) are crucial in various biological processes and their study has significant implications for drug development and disease diagnosis. Existing deep learning methods suffer from significant performance degradation under complex real-world scenarios due to various factors, e.g., label scarcity and domain shift. In this paper, we propose a self-ensembling multigraph neural network (SemiGNN-PPI) that can effectively predict PPIs while being both efficient and generalizable. In SemiGNN-PPI, we not only model the protein correlations but explore the label dependencies by constructing and processing multiple graphs from the perspectives of both features and labels in the graph learning process. We further marry GNN with Mean Teacher to effectively leverage unlabeled graph-structured PPI data for self-ensemble graph learning. We also design multiple graph consistency constraints to align the student and teacher graphs in the feature embedding space, enabling the student model to better learn from the teacher model by incorporating more relationships. Extensive experiments on PPI datasets of different scales with different evaluation settings demonstrate that SemiGNN-PPI outperforms state-of-the-art PPI prediction methods, particularly in challenging scenarios such as training with limited annotations and testing on unseen data.
MS-MT: Multi-Scale Mean Teacher with Contrastive Unpaired Translation for Cross-Modality Vestibular Schwannoma and Cochlea Segmentation
Mar 28, 2023Ziyuan Zhao, Kaixin Xu, Huai Zhe Yeo, Xulei Yang, Cuntai Guan
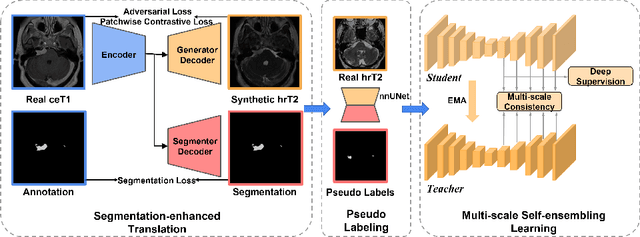

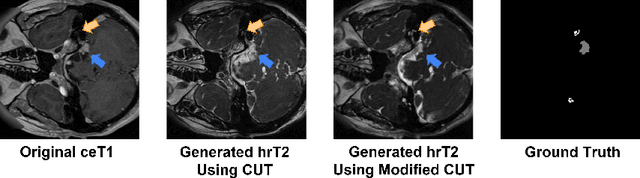
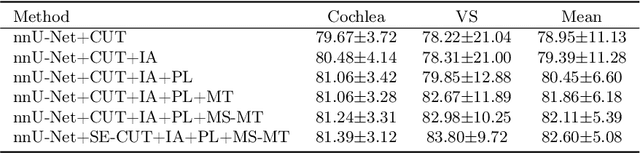
Domain shift has been a long-standing issue for medical image segmentation. Recently, unsupervised domain adaptation (UDA) methods have achieved promising cross-modality segmentation performance by distilling knowledge from a label-rich source domain to a target domain without labels. In this work, we propose a multi-scale self-ensembling based UDA framework for automatic segmentation of two key brain structures i.e., Vestibular Schwannoma (VS) and Cochlea on high-resolution T2 images. First, a segmentation-enhanced contrastive unpaired image translation module is designed for image-level domain adaptation from source T1 to target T2. Next, multi-scale deep supervision and consistency regularization are introduced to a mean teacher network for self-ensemble learning to further close the domain gap. Furthermore, self-training and intensity augmentation techniques are utilized to mitigate label scarcity and boost cross-modality segmentation performance. Our method demonstrates promising segmentation performance with a mean Dice score of 83.8% and 81.4% and an average asymmetric surface distance (ASSD) of 0.55 mm and 0.26 mm for the VS and Cochlea, respectively in the validation phase of the crossMoDA 2022 challenge.
MMGL: Multi-Scale Multi-View Global-Local Contrastive learning for Semi-supervised Cardiac Image Segmentation
Jul 05, 2022Ziyuan Zhao, Jinxuan Hu, Zeng Zeng, Xulei Yang, Peisheng Qian, Bharadwaj Veeravalli, Cuntai Guan
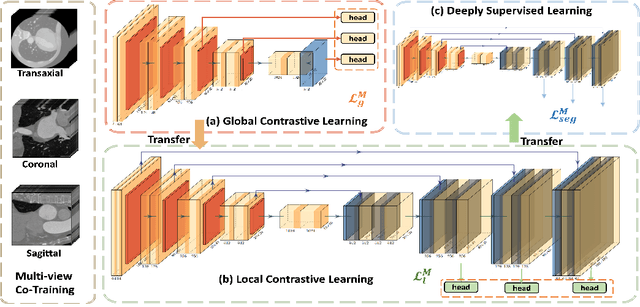
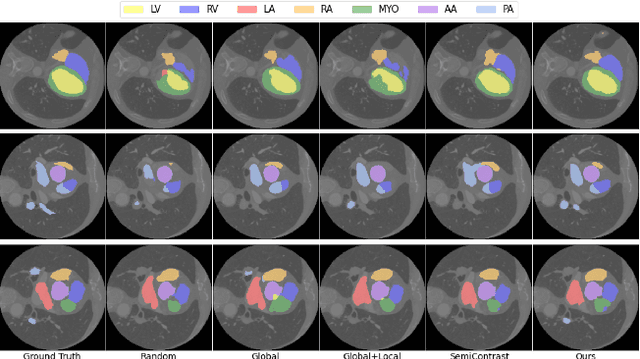
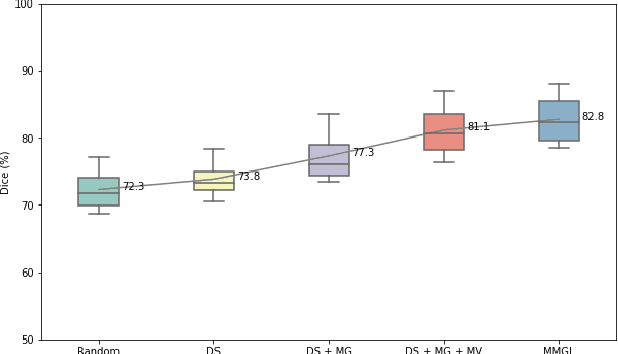
With large-scale well-labeled datasets, deep learning has shown significant success in medical image segmentation. However, it is challenging to acquire abundant annotations in clinical practice due to extensive expertise requirements and costly labeling efforts. Recently, contrastive learning has shown a strong capacity for visual representation learning on unlabeled data, achieving impressive performance rivaling supervised learning in many domains. In this work, we propose a novel multi-scale multi-view global-local contrastive learning (MMGL) framework to thoroughly explore global and local features from different scales and views for robust contrastive learning performance, thereby improving segmentation performance with limited annotations. Extensive experiments on the MM-WHS dataset demonstrate the effectiveness of MMGL framework on semi-supervised cardiac image segmentation, outperforming the state-of-the-art contrastive learning methods by a large margin.
Systematic Analysis and Removal of Circular Artifacts for StyleGAN
Mar 04, 2021Way Tan, Bihan Wen, Xulei Yang
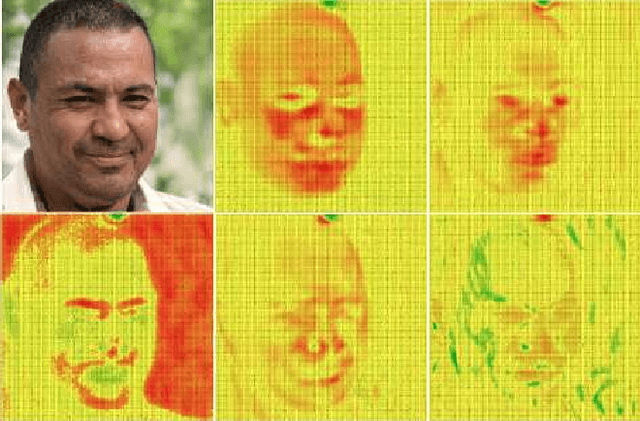

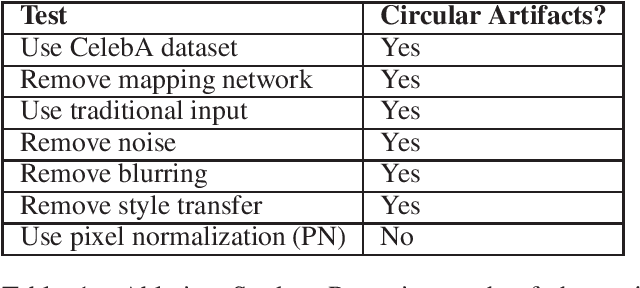
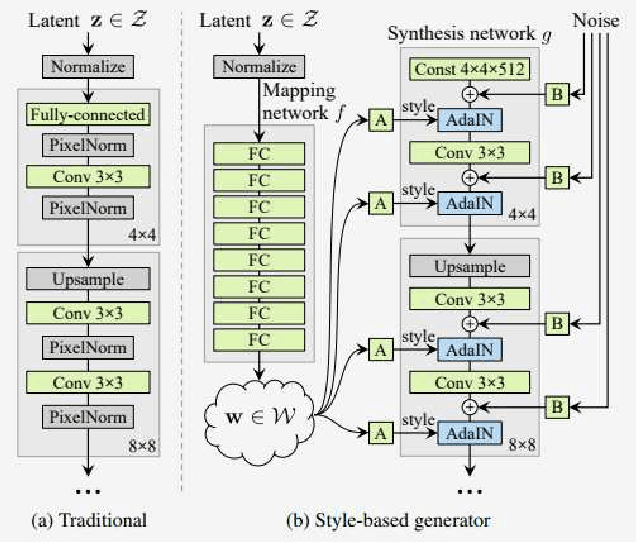

StyleGAN is one of the state-of-the-art image generators which is well-known for synthesizing high-resolution and hyper-realistic face images. Though images generated by vanilla StyleGAN model are visually appealing, they sometimes contain prominent circular artifacts which severely degrade the quality of generated images. In this work, we provide a systematic investigation on how those circular artifacts are formed by studying the functionalities of different stages of vanilla StyleGAN architecture, with both mechanism analysis and extensive experiments. The key modules of vanilla StyleGAN that promote such undesired artifacts are highlighted. Our investigation also explains why the artifacts are usually circular, relatively small and rarely split into 2 or more parts. Besides, we propose a simple yet effective solution to remove the prominent circular artifacts for vanilla StyleGAN, by applying a novel pixel-instance normalization (PIN) layer.
Semi-supervised and Unsupervised Methods for Heart Sounds Classification in Restricted Data Environments
Jun 04, 2020Balagopal Unnikrishnan, Pranshu Ranjan Singh, Xulei Yang, Matthew Chin Heng Chua
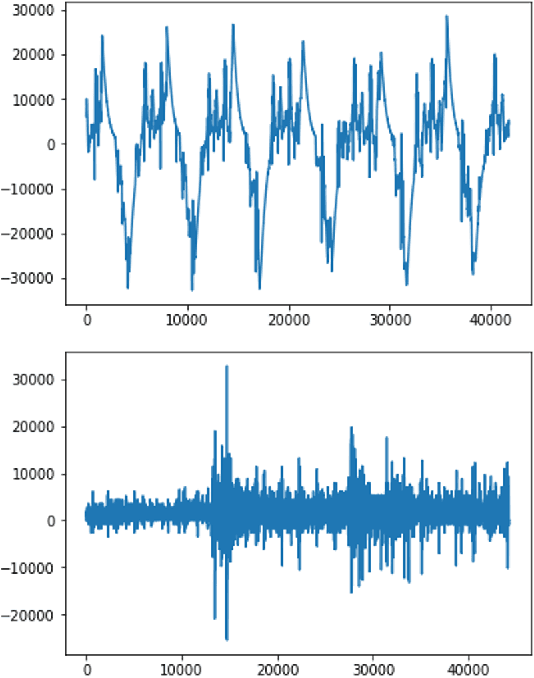
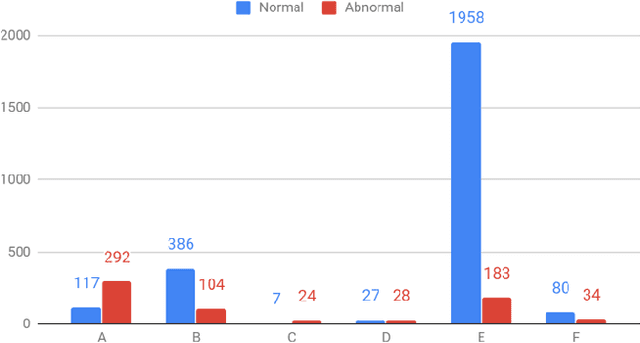
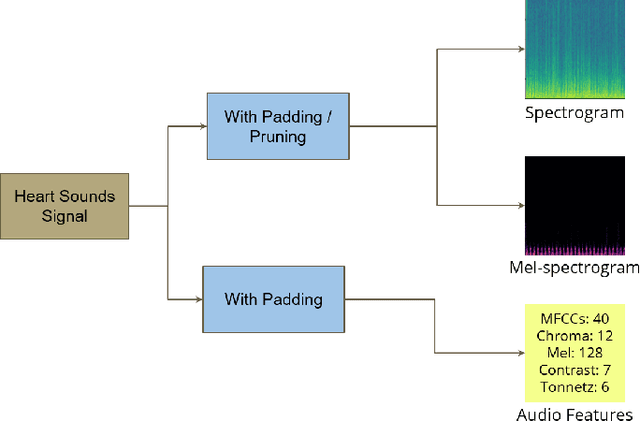
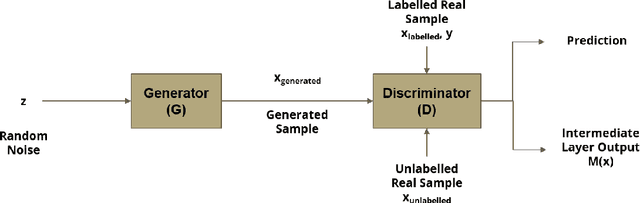
Automated heart sounds classification is a much-required diagnostic tool in the view of increasing incidences of heart related diseases worldwide. In this study, we conduct a comprehensive study of heart sounds classification by using various supervised, semi-supervised and unsupervised approaches on the PhysioNet/CinC 2016 Challenge dataset. Supervised approaches, including deep learning and machine learning methods, require large amounts of labelled data to train the models, which are challenging to obtain in most practical scenarios. In view of the need to reduce the labelling burden for clinical practices, where human labelling is both expensive and time-consuming, semi-supervised or even unsupervised approaches in restricted data setting are desirable. A GAN based semi-supervised method is therefore proposed, which allows the usage of unlabelled data samples to boost the learning of data distribution. It achieves a better performance in terms of AUROC over the supervised baseline when limited data samples exist. Furthermore, several unsupervised methods are explored as an alternative approach by considering the given problem as an anomaly detection scenario. In particular, the unsupervised feature extraction using 1D CNN Autoencoder coupled with one-class SVM obtains good performance without any data labelling. The potential of the proposed semi-supervised and unsupervised methods may lead to a workflow tool in the future for the creation of higher quality datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge