Rethinking the Encoding of Satellite Image Time Series
May 03, 2023Xin Cai, Yaxin Bi, Peter Nicholl, Roy Sterritt
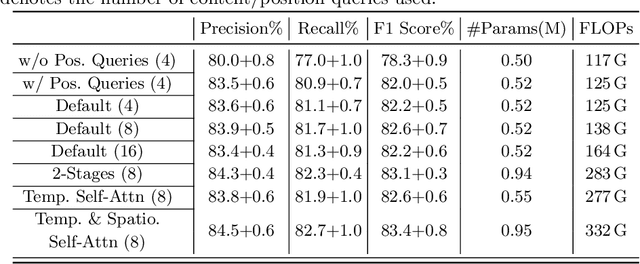
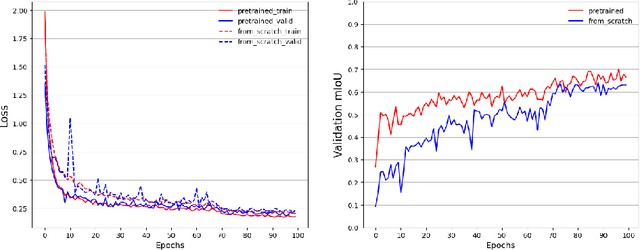
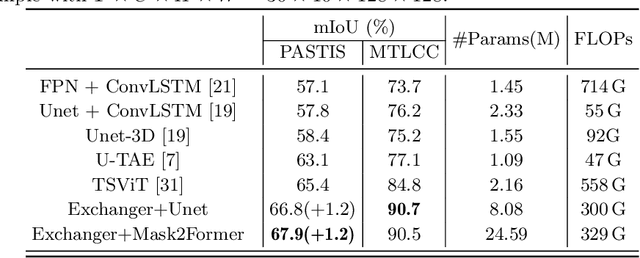
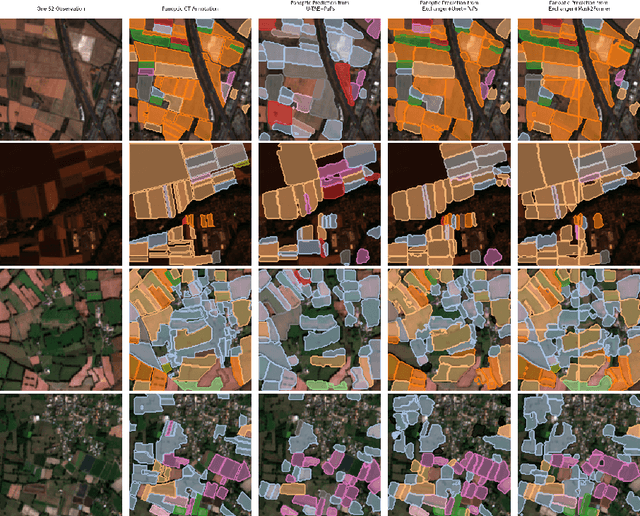
Representation learning of Satellite Image Time Series (SITS) presents its unique challenges, such as prohibitive computation burden caused by high spatiotemporal resolutions, irregular acquisition times, and complex spatiotemporal interactions, leading to highly-specialized neural network architectures for SITS analysis. Despite the promising results achieved by some pioneering work, we argue that satisfactory representation learning paradigms have not yet been established for SITS analysis, causing an isolated island where transferring successful paradigms or the latest advances from Computer Vision (CV) to SITS is arduous. In this paper, we develop a unique perspective of SITS processing as a direct set prediction problem, inspired by the recent trend in adopting query-based transformer decoders to streamline the object detection or image segmentation pipeline, and further propose to decompose the representation learning process of SITS into three explicit steps: collect--update--distribute, which is computationally efficient and suits for irregularly-sampled and asynchronous temporal observations. Facilitated by the unique reformulation and effective feature extraction framework proposed, our models pre-trained on pixel-set format input and then fine-tuned on downstream dense prediction tasks by simply appending a commonly-used segmentation network have attained new state-of-the-art (SoTA) results on PASTIS dataset compared to bespoke neural architectures such as U-TAE. Furthermore, the clear separation, conceptually and practically, between temporal and spatial components in the panoptic segmentation pipeline of SITS allows us to leverage the recent advances in CV, such as Mask2Former, a universal segmentation architecture, resulting in a noticeable 8.8 points increase in PQ compared to the best score reported so far.
Tampered VAE for Improved Satellite Image Time Series Classification
Mar 30, 2022Xin Cai, Yaxin Bi, Peter Nicholl
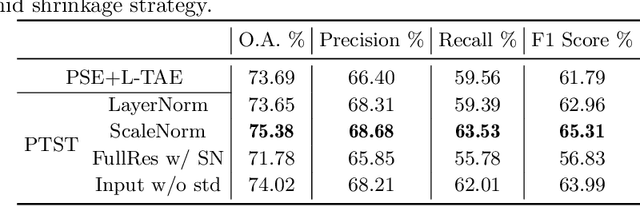
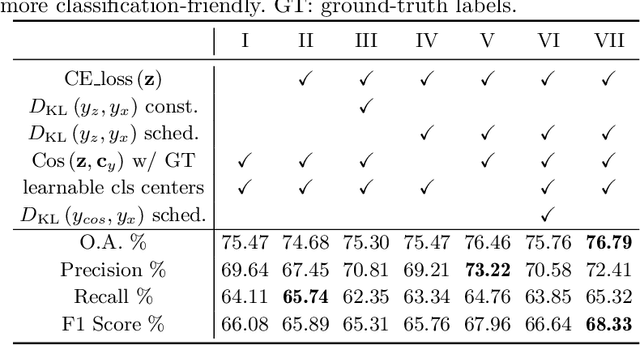
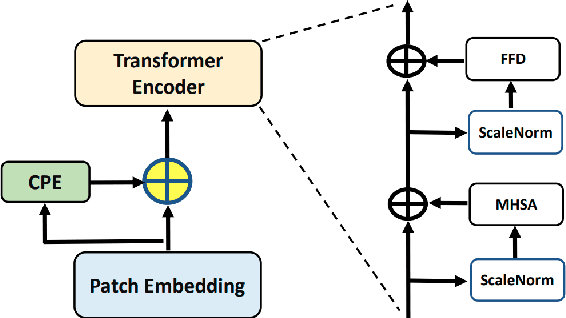
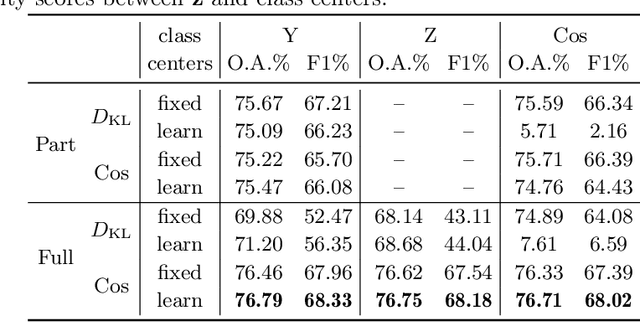
The unprecedented availability of spatial and temporal high-resolution satellite image time series (SITS) for crop type mapping is believed to necessitate deep learning architectures to accommodate challenges arising from both dimensions. Recent state-of-the-art deep learning models have shown promising results by stacking spatial and temporal encoders. However, we present a Pyramid Time-Series Transformer (PTST) that operates solely on the temporal dimension, i.e., neglecting the spatial dimension, can produce superior results with a drastic reduction in GPU memory consumption and easy extensibility. Furthermore, we augment it to perform semi-supervised learning by proposing a classification-friendly VAE framework that introduces clustering mechanisms into latent space and can promote linear separability therein. Consequently, a few principal axes of the latent space can explain the majority of variance in raw data. Meanwhile, the VAE framework with proposed tweaks can maintain competitive classification performance as its purely discriminative counterpart when only $40\%$ of labelled data is used. We hope the proposed framework can serve as a baseline for crop classification with SITS for its modularity and simplicity.
AlphaFold Accelerates Artificial Intelligence Powered Drug Discovery: Efficient Discovery of a Novel Cyclin-dependent Kinase 20 (CDK20) Small Molecule Inhibitor
Jan 21, 2022Feng Ren, Xiao Ding, Min Zheng, Mikhail Korzinkin, Xin Cai, Wei Zhu, Alexey Mantsyzov, Alex Aliper, Vladimir Aladinskiy, Zhongying Cao, Shanshan Kong, Xi Long, Bonnie Hei Man Liu, Yingtao Liu, Vladimir Naumov, Anastasia Shneyderman, Ivan V. Ozerov, Ju Wang, Frank W. Pun, Alan Aspuru-Guzik, Michael Levitt, Alex Zhavoronkov
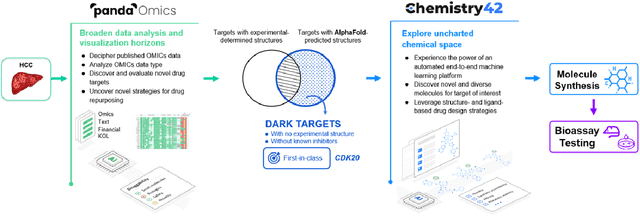
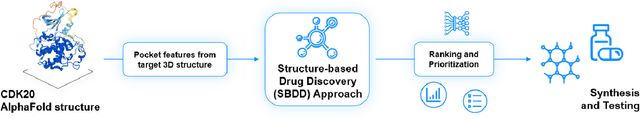
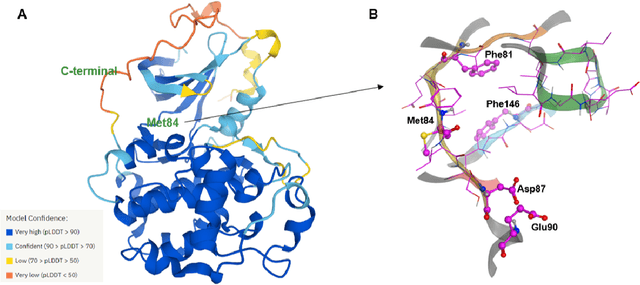
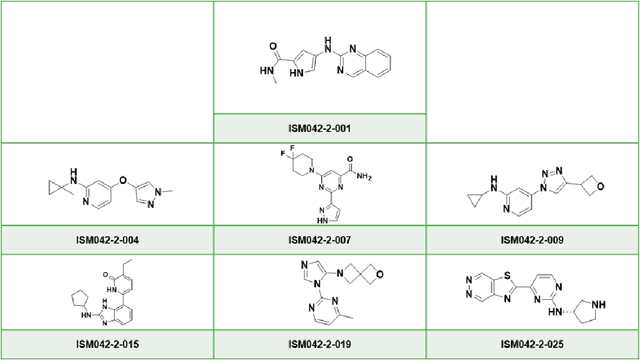
The AlphaFold computer program predicted protein structures for the whole human genome, which has been considered as a remarkable breakthrough both in artificial intelligence (AI) application and structural biology. Despite the varying confidence level, these predicted structures still could significantly contribute to the structure-based drug design of novel targets, especially the ones with no or limited structural information. In this work, we successfully applied AlphaFold in our end-to-end AI-powered drug discovery engines constituted of a biocomputational platform PandaOmics and a generative chemistry platform Chemistry42, to identify a first-in-class hit molecule of a novel target without an experimental structure starting from target selection towards hit identification in a cost- and time-efficient manner. PandaOmics provided the targets of interest and Chemistry42 generated the molecules based on the AlphaFold predicted structure, and the selected molecules were synthesized and tested in biological assays. Through this approach, we identified a small molecule hit compound for CDK20 with a Kd value of 8.9 +/- 1.6 uM (n = 4) within 30 days from target selection and after only synthesizing 7 compounds. To the best of our knowledge, this is the first reported small molecule targeting CDK20 and more importantly, this work is the first demonstration of AlphaFold application in the hit identification process in early drug discovery.
Gaze Estimation with an Ensemble of Four Architectures
Jul 05, 2021Xin Cai, Boyu Chen, Jiabei Zeng, Jiajun Zhang, Yunjia Sun, Xiao Wang, Zhilong Ji, Xiao Liu, Xilin Chen, Shiguang Shan
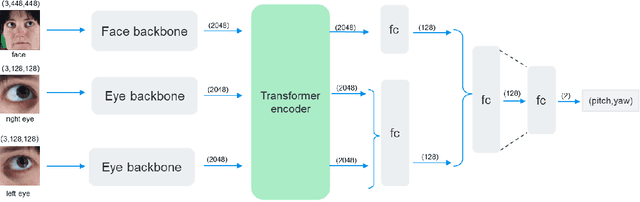
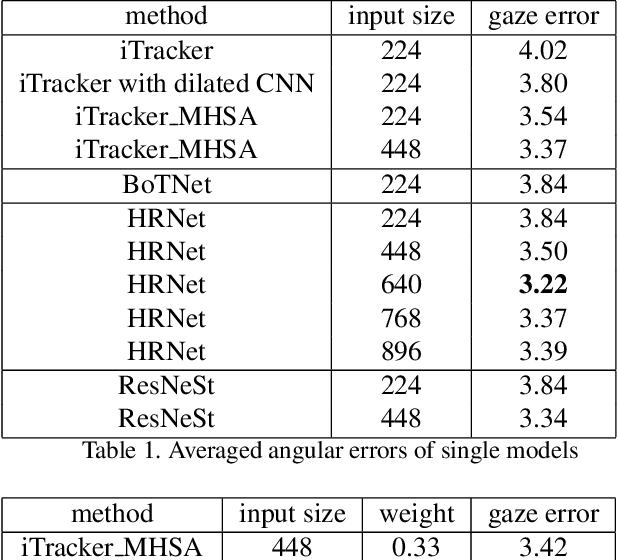
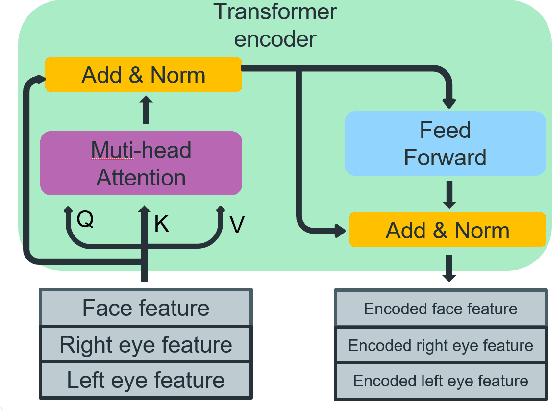
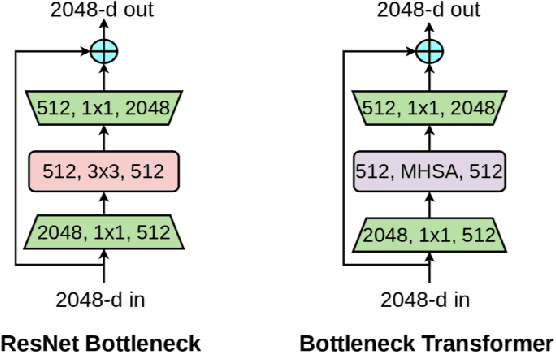
This paper presents a method for gaze estimation according to face images. We train several gaze estimators adopting four different network architectures, including an architecture designed for gaze estimation (i.e.,iTracker-MHSA) and three originally designed for general computer vision tasks(i.e., BoTNet, HRNet, ResNeSt). Then, we select the best six estimators and ensemble their predictions through a linear combination. The method ranks the first on the leader-board of ETH-XGaze Competition, achieving an average angular error of $3.11^{\circ}$ on the ETH-XGaze test set.
Pixel-Level Dense Prediction without Decoder
Sep 22, 2019Xin Cai, Yi-Fei Pu
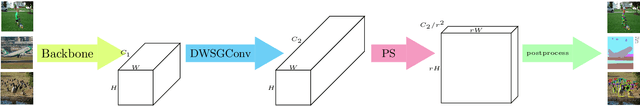
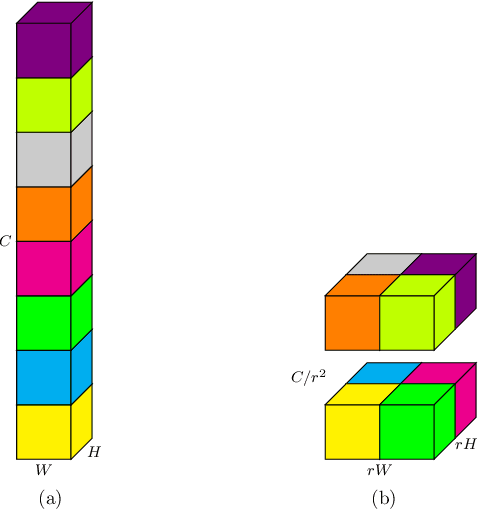
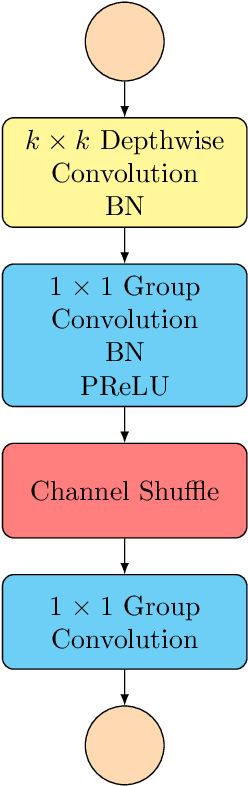

Pixel-level dense prediction tasks such as keypoint estimation are dominated by encoder-decoder structures, where the decoder as a vital component is complex and computationally intensive. In contrast, we propose a fully decoding-free pixel-level dense prediction network called FlatteNet, in which the high dimensional tensor outputted by the backbone network is directly flattened to fit the desired output resolution. The proposed FlatteNet is end-to-end differentiable. By removing the decoder unit, FlatteNet requires much fewer parameters and lower computational complexity. We empirically demonstrate the effectiveness of the proposed network through competitive results in human pose estimation on MPII, semantic segmentation on PASCAL-Context, and object detection on PASCAL VOC. We hope that the proposed FlatteNet can serve as a simple and strong alternative of current mainstream decoder-based pixel-level dense prediction networks.
An overlapping-free leaf segmentation method for plant point clouds
Aug 12, 2019Dawei Li, Yan Cao, Guoliang Shi, Xin Cai, Yang Chen, Sifan Wang, Siyuan Yan
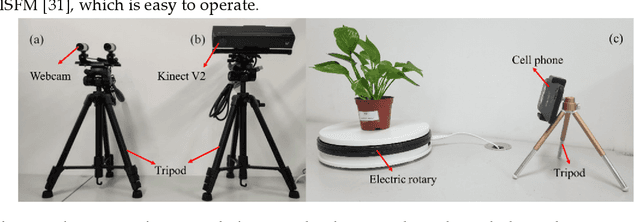
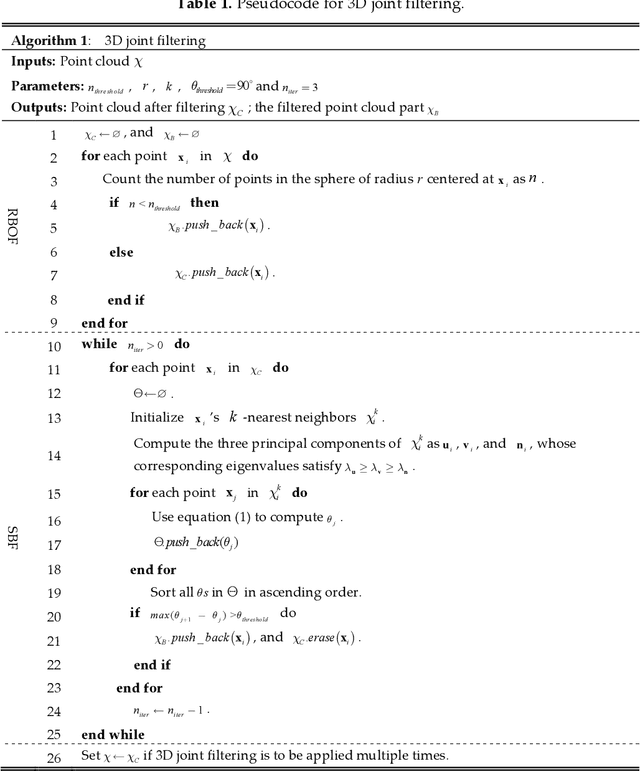
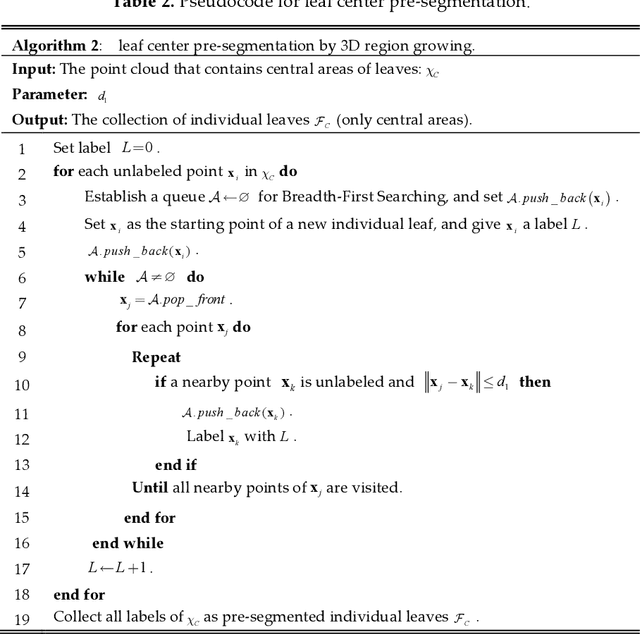


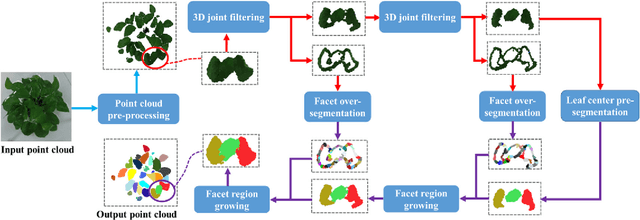

Automatic leaf segmentation, as well as identification and classification methods that built upon it, are able to provide immediate monitoring for plant growth status to guarantee the output. Although 3D plant point clouds contain abundant phenotypic features, plant leaves are usually distributed in clusters and are sometimes seriously overlapped in the canopy. Therefore, it is still a big challenge to automatically segment each individual leaf from a highly crowded plant canopy in 3D for plant phenotyping purposes. In this work, we propose an overlapping-free individual leaf segmentation method for plant point clouds using the 3D filtering and facet region growing. In order to separate leaves with different overlapping situations, we develop a new 3D joint filtering operator, which integrates a Radius-based Outlier Filter (RBOF) and a Surface Boundary Filter (SBF) to help to separate occluded leaves. By introducing the facet over-segmentation and facet-based region growing, the noise in segmentation is suppressed and labeled leaf centers can expand to their whole leaves, respectively. Our method can work on point clouds generated from three types of 3D imaging platforms, and also suitable for different kinds of plant species. In experiments, it obtains a point-level cover rate of 97% for Epipremnum aureum, 99% for Monstera deliciosa, 99% for Calathea makoyana, and 87% for Hedera nepalensis sample plants. At the leaf level, our method reaches an average Recall at 100.00%, a Precision at 99.33%, and an average F-measure at 99.66%, respectively. The proposed method can also facilitate the automatic traits estimation of each single leaf (such as the leaf area, length, and width), which has potential to become a highly effective tool for plant research and agricultural engineering.
An Integrated Image Filter for Enhancing Change Detection Results
Jul 02, 2019Dawei Li, Siyuan Yan, Xin Cai, Yan Cao, Sifan Wang
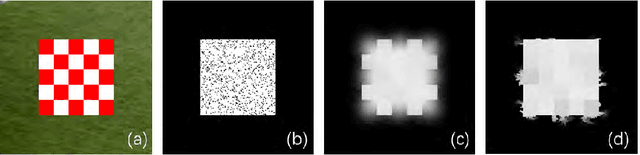
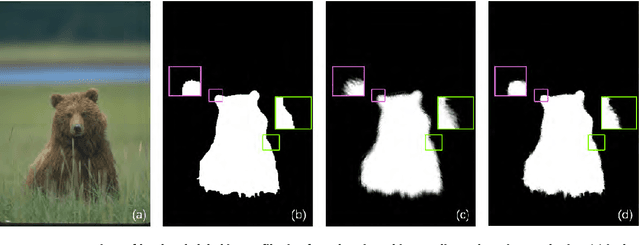

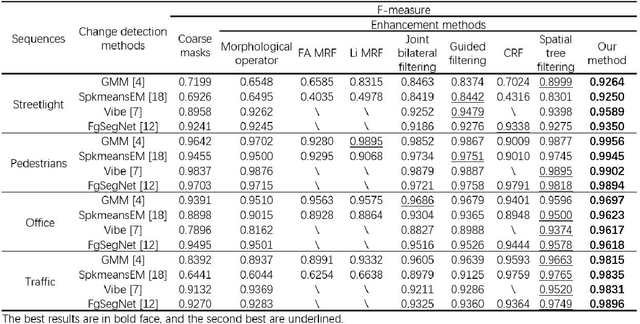

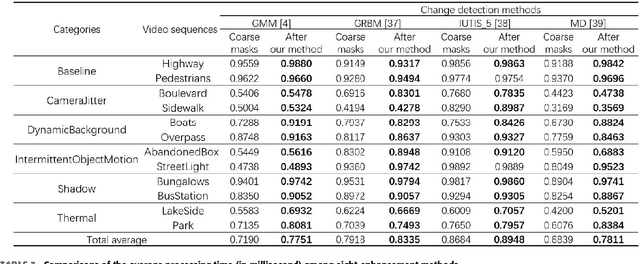
Change detection is a fundamental task in computer vision. Despite significant advances have been made, most of the change detection methods fail to work well in challenging scenes due to ubiquitous noise and interferences. Nowadays, post-processing methods (e.g. MRF, and CRF) aiming to enhance the binary change detection results still fall short of the requirements on universality for distinctive scenes, applicability for different types of detection methods, accuracy, and real-time performance. Inspired by the nature of image filtering, which separates noise from pixel observations and recovers the real structure of patches, we consider utilizing image filters to enhance the detection masks. In this paper, we present an integrated filter which comprises a weighted local guided image filter and a weighted spatiotemporal tree filter. The spatiotemporal tree filter leverages the global spatiotemporal information of adjacent video frames and meanwhile the guided filter carries out local window filtering of pixels, for enhancing the coarse change detection masks. The main contributions are three: (i) the proposed filter can make full use of the information of the same object in consecutive frames to improve its current detection mask by computations on a spatiotemporal minimum spanning tree; (ii) the integrated filter possesses both advantages of local filtering and global filtering; it not only has good edge-preserving property but also can handle heavily textured and colorful foreground regions; and (iii) Unlike some popular enhancement methods (MRF, and CRF) that require either a priori background probabilities or a posteriori foreground probabilities for every pixel to improve the coarse detection masks, our method is a versatile enhancement filter that can be applied after many different types of change detection methods, and is particularly suitable for video sequences.
Bayesian inverse regression for supervised dimension reduction with small datasets
Jun 19, 2019Xin Cai, Guang Lin, Jinglai Li
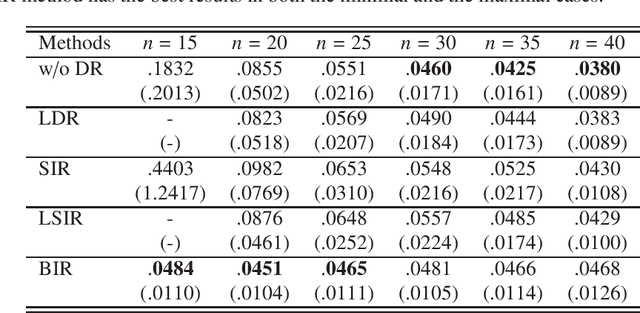
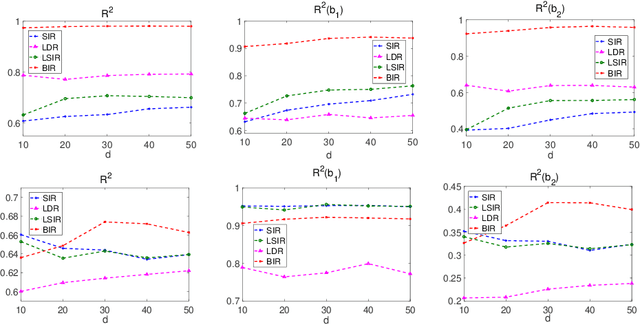

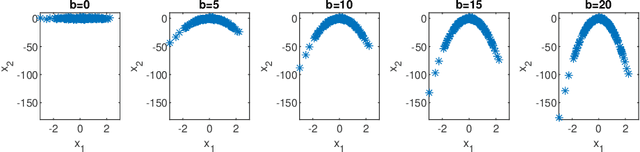
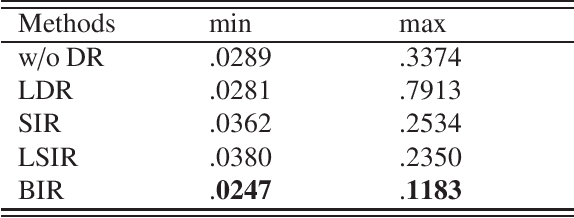
We consider supervised dimension reduction problems, namely to identify a low dimensional projection of the predictors $\-x$ which can retain the statistical relationship between $\-x$ and the response variable $y$. We follow the idea of the sliced inverse regression (SIR) class of methods, which is to use the statistical information of the conditional distribution $\pi(\-x|y)$ to identify the dimension reduction (DR) space and in particular we focus on the task of computing this conditional distribution. We propose a Bayesian framework to compute the conditional distribution where the likelihood function is obtained using the Gaussian process regression model. The conditional distribution $\pi(\-x|y)$ can then be obtained directly by assigning weights to the original data points. We then can perform DR by considering certain moment functions (e.g. the first moment) of the samples of the posterior distribution. With numerical examples, we demonstrate that the proposed method is especially effective for small data problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge