Toward Sustainable GenAI using Generation Directives for Carbon-Friendly Large Language Model Inference
Mar 19, 2024Baolin Li, Yankai Jiang, Vijay Gadepally, Devesh Tiwari
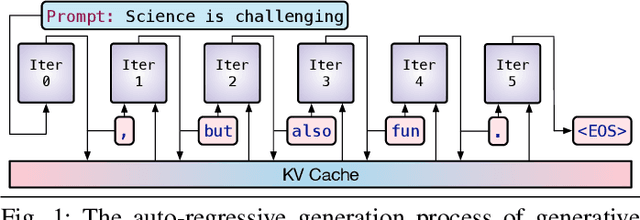

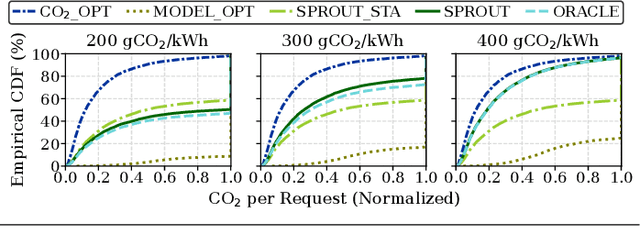
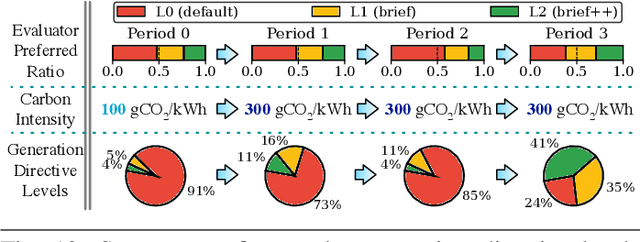
The rapid advancement of Generative Artificial Intelligence (GenAI) across diverse sectors raises significant environmental concerns, notably the carbon emissions from their cloud and high performance computing (HPC) infrastructure. This paper presents Sprout, an innovative framework designed to address these concerns by reducing the carbon footprint of generative Large Language Model (LLM) inference services. Sprout leverages the innovative concept of "generation directives" to guide the autoregressive generation process, thereby enhancing carbon efficiency. Our proposed method meticulously balances the need for ecological sustainability with the demand for high-quality generation outcomes. Employing a directive optimizer for the strategic assignment of generation directives to user prompts and an original offline quality evaluator, Sprout demonstrates a significant reduction in carbon emissions by over 40% in real-world evaluations using the Llama2 LLM and global electricity grid data. This research marks a critical step toward aligning AI technology with sustainable practices, highlighting the potential for mitigating environmental impacts in the rapidly expanding domain of generative artificial intelligence.
ZePT: Zero-Shot Pan-Tumor Segmentation via Query-Disentangling and Self-Prompting
Dec 07, 2023Yankai Jiang, Zhongzhen Huang, Rongzhao Zhang, Xiaofan Zhang, Shaoting Zhang
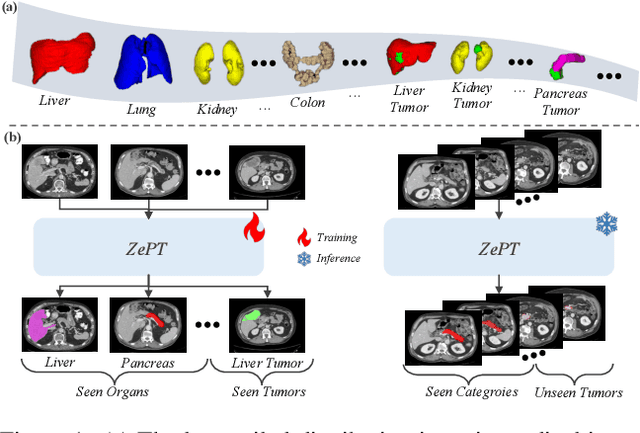
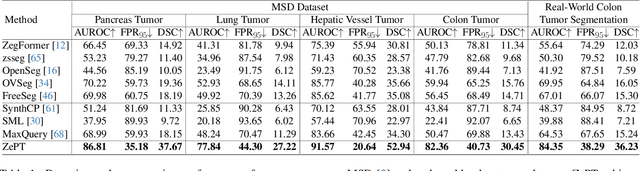
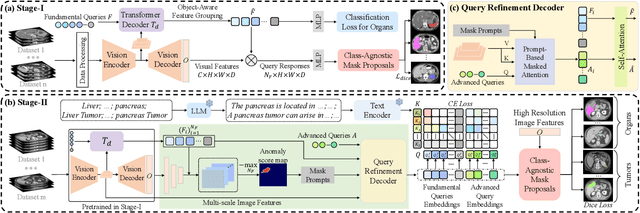
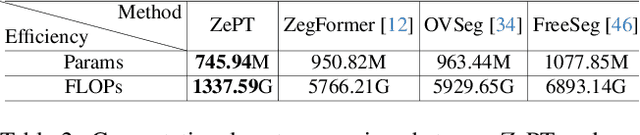
The long-tailed distribution problem in medical image analysis reflects a high prevalence of common conditions and a low prevalence of rare ones, which poses a significant challenge in developing a unified model capable of identifying rare or novel tumor categories not encountered during training. In this paper, we propose a new zero-shot pan-tumor segmentation framework (ZePT) based on query-disentangling and self-prompting to segment unseen tumor categories beyond the training set. ZePT disentangles the object queries into two subsets and trains them in two stages. Initially, it learns a set of fundamental queries for organ segmentation through an object-aware feature grouping strategy, which gathers organ-level visual features. Subsequently, it refines the other set of advanced queries that focus on the auto-generated visual prompts for unseen tumor segmentation. Moreover, we introduce query-knowledge alignment at the feature level to enhance each query's discriminative representation and generalizability. Extensive experiments on various tumor segmentation tasks demonstrate the performance superiority of ZePT, which surpasses the previous counterparts and evidence the promising ability for zero-shot tumor segmentation in real-world settings. Codes will be made publicly available.
Anatomical Invariance Modeling and Semantic Alignment for Self-supervised Learning in 3D Medical Image Segmentation
Feb 27, 2023Yankai Jiang, Mingze Sun, Heng Guo, Ke Yan, Le Lu, Minfeng Xu

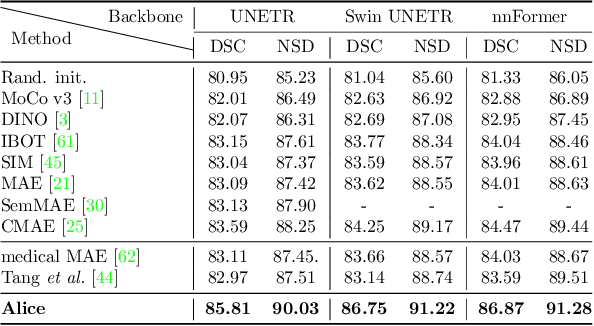
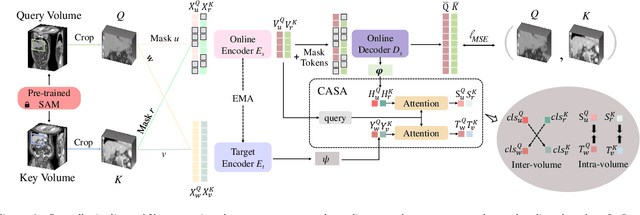
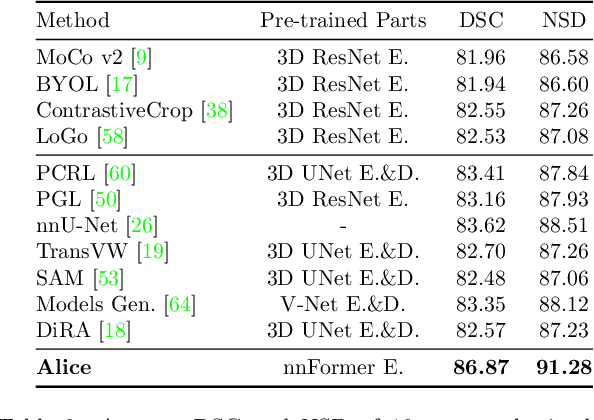
Self-supervised learning (SSL) has recently achieved promising performance for 3D medical image segmentation tasks. Most current methods follow existing SSL paradigm originally designed for photographic or natural images, which cannot explicitly and thoroughly exploit the intrinsic similar anatomical structures across varying medical images. This may in fact degrade the quality of learned deep representations by maximizing the similarity among features containing spatial misalignment information and different anatomical semantics. In this work, we propose a new self-supervised learning framework, namely Alice, that explicitly fulfills Anatomical invariance modeling and semantic alignment via elaborately combining discriminative and generative objectives. Alice introduces a new contrastive learning strategy which encourages the similarity between views that are diversely mined but with consistent high-level semantics, in order to learn invariant anatomical features. Moreover, we design a conditional anatomical feature alignment module to complement corrupted embeddings with globally matched semantics and inter-patch topology information, conditioned by the distribution of local image content, which permits to create better contrastive pairs. Our extensive quantitative experiments on two public 3D medical image segmentation benchmarks of FLARE 2022 and BTCV demonstrate and validate the performance superiority of Alice, surpassing the previous best SSL counterpart methods by 2.11% and 1.77% in Dice coefficients, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge