SegRap2023: A Benchmark of Organs-at-Risk and Gross Tumor Volume Segmentation for Radiotherapy Planning of Nasopharyngeal Carcinoma
Dec 15, 2023Xiangde Luo, Jia Fu, Yunxin Zhong, Shuolin Liu, Bing Han, Mehdi Astaraki, Simone Bendazzoli, Iuliana Toma-Dasu, Yiwen Ye, Ziyang Chen, Yong Xia, Yanzhou Su, Jin Ye, Junjun He, Zhaohu Xing, Hongqiu Wang, Lei Zhu, Kaixiang Yang, Xin Fang, Zhiwei Wang, Chan Woong Lee, Sang Joon Park, Jaehee Chun, Constantin Ulrich, Klaus H. Maier-Hein, Nchongmaje Ndipenoch, Alina Miron, Yongmin Li, Yimeng Zhang, Yu Chen, Lu Bai, Jinlong Huang, Chengyang An, Lisheng Wang, Kaiwen Huang, Yunqi Gu, Tao Zhou, Mu Zhou, Shichuan Zhang, Wenjun Liao, Guotai Wang, Shaoting Zhang
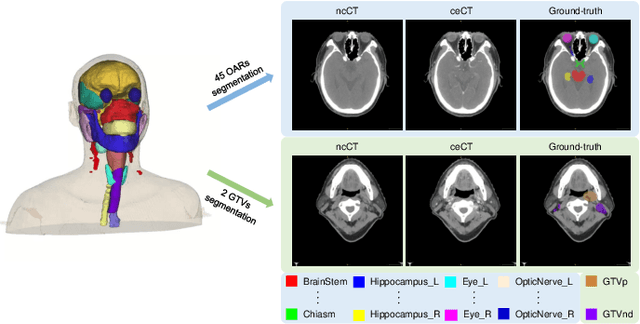
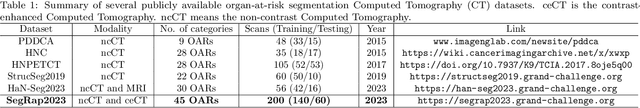
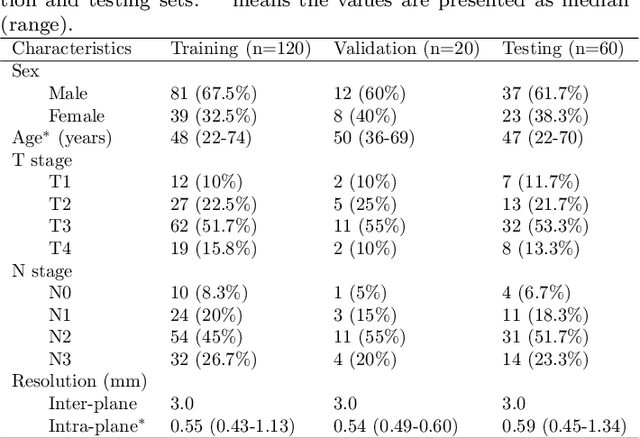
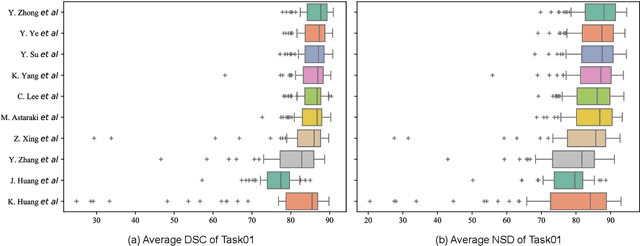
Radiation therapy is a primary and effective NasoPharyngeal Carcinoma (NPC) treatment strategy. The precise delineation of Gross Tumor Volumes (GTVs) and Organs-At-Risk (OARs) is crucial in radiation treatment, directly impacting patient prognosis. Previously, the delineation of GTVs and OARs was performed by experienced radiation oncologists. Recently, deep learning has achieved promising results in many medical image segmentation tasks. However, for NPC OARs and GTVs segmentation, few public datasets are available for model development and evaluation. To alleviate this problem, the SegRap2023 challenge was organized in conjunction with MICCAI2023 and presented a large-scale benchmark for OAR and GTV segmentation with 400 Computed Tomography (CT) scans from 200 NPC patients, each with a pair of pre-aligned non-contrast and contrast-enhanced CT scans. The challenge's goal was to segment 45 OARs and 2 GTVs from the paired CT scans. In this paper, we detail the challenge and analyze the solutions of all participants. The average Dice similarity coefficient scores for all submissions ranged from 76.68\% to 86.70\%, and 70.42\% to 73.44\% for OARs and GTVs, respectively. We conclude that the segmentation of large-size OARs is well-addressed, and more efforts are needed for GTVs and small-size or thin-structure OARs. The benchmark will remain publicly available here: https://segrap2023.grand-challenge.org
Each Test Image Deserves A Specific Prompt: Continual Test-Time Adaptation for 2D Medical Image Segmentation
Nov 30, 2023Ziyang Chen, Yiwen Ye, Mengkang Lu, Yongsheng Pan, Yong Xia
Distribution shift widely exists in medical images acquired from different medical centres and poses a significant obstacle to deploying the pre-trained semantic segmentation model in real-world applications. Test-time adaptation has proven its effectiveness in tackling the cross-domain distribution shift during inference. However, most existing methods achieve adaptation by updating the pre-trained models, rendering them susceptible to error accumulation and catastrophic forgetting when encountering a series of distribution shifts (i.e., under the continual test-time adaptation setup). To overcome these challenges caused by updating the models, in this paper, we freeze the pre-trained model and propose the Visual Prompt-based Test-Time Adaptation (VPTTA) method to train a specific prompt for each test image to align the statistics in the batch normalization layers. Specifically, we present the low-frequency prompt, which is lightweight with only a few parameters and can be effectively trained in a single iteration. To enhance prompt initialization, we equip VPTTA with a memory bank to benefit the current prompt from previous ones. Additionally, we design a warm-up mechanism, which mixes source and target statistics to construct warm-up statistics, thereby facilitating the training process. Extensive experiments demonstrate the superiority of our VPTTA over other state-of-the-art methods on two medical image segmentation benchmark tasks. The code and weights of pre-trained source models are available at https://github.com/Chen-Ziyang/VPTTA.
Continual Self-supervised Learning: Towards Universal Multi-modal Medical Data Representation Learning
Nov 30, 2023Yiwen Ye, Yutong Xie, Jianpeng Zhang, Ziyang Chen, Qi Wu, Yong Xia
Self-supervised learning is an efficient pre-training method for medical image analysis. However, current research is mostly confined to specific-modality data pre-training, consuming considerable time and resources without achieving universality across different modalities. A straightforward solution is combining all modality data for joint self-supervised pre-training, which poses practical challenges. Firstly, our experiments reveal conflicts in representation learning as the number of modalities increases. Secondly, multi-modal data collected in advance cannot cover all real-world scenarios. In this paper, we reconsider versatile self-supervised learning from the perspective of continual learning and propose MedCoSS, a continuous self-supervised learning approach for multi-modal medical data. Unlike joint self-supervised learning, MedCoSS assigns different modality data to different training stages, forming a multi-stage pre-training process. To balance modal conflicts and prevent catastrophic forgetting, we propose a rehearsal-based continual learning method. We introduce the k-means sampling strategy to retain data from previous modalities and rehearse it when learning new modalities. Instead of executing the pretext task on buffer data, a feature distillation strategy and an intra-modal mixup strategy are applied to these data for knowledge retention. We conduct continuous self-supervised pre-training on a large-scale multi-modal unlabeled dataset, including clinical reports, X-rays, CT scans, MRI scans, and pathological images. Experimental results demonstrate MedCoSS's exceptional generalization ability across nine downstream datasets and its significant scalability in integrating new modality data. Code and pre-trained weight are available at https://github.com/yeerwen/MedCoSS.
Treasure in Distribution: A Domain Randomization based Multi-Source Domain Generalization for 2D Medical Image Segmentation
May 31, 2023Ziyang Chen, Yongsheng Pan, Yiwen Ye, Hengfei Cui, Yong Xia




Although recent years have witnessed the great success of convolutional neural networks (CNNs) in medical image segmentation, the domain shift issue caused by the highly variable image quality of medical images hinders the deployment of CNNs in real-world clinical applications. Domain generalization (DG) methods aim to address this issue by training a robust model on the source domain, which has a strong generalization ability. Previously, many DG methods based on feature-space domain randomization have been proposed, which, however, suffer from the limited and unordered search space of feature styles. In this paper, we propose a multi-source DG method called Treasure in Distribution (TriD), which constructs an unprecedented search space to obtain the model with strong robustness by randomly sampling from a uniform distribution. To learn the domain-invariant representations explicitly, we further devise a style-mixing strategy in our TriD, which mixes the feature styles by randomly mixing the augmented and original statistics along the channel wise and can be extended to other DG methods. Extensive experiments on two medical segmentation tasks with different modalities demonstrate that our TriD achieves superior generalization performance on unseen target-domain data. Code is available at https://github.com/Chen-Ziyang/TriD.
UniSeg: A Prompt-driven Universal Segmentation Model as well as A Strong Representation Learner
Apr 07, 2023Yiwen Ye, Yutong Xie, Jianpeng Zhang, Ziyang Chen, Yong Xia
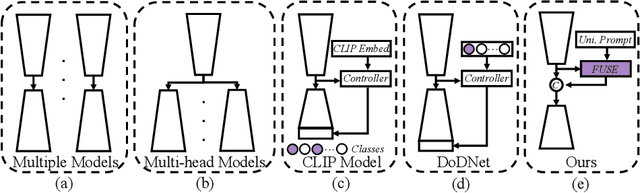
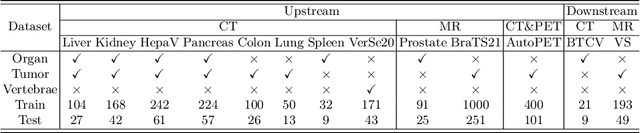
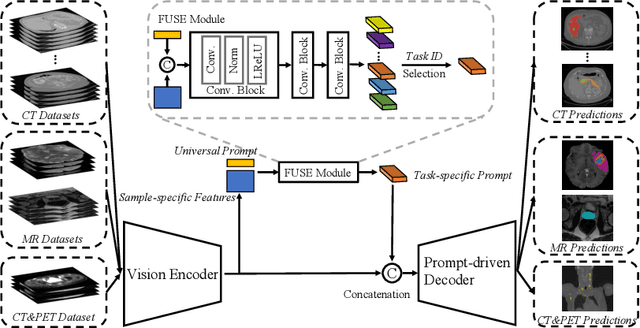
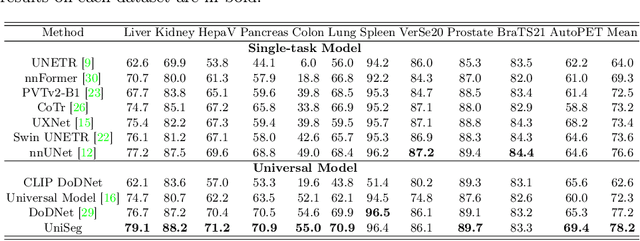
The universal model emerges as a promising trend for medical image segmentation, paving up the way to build medical imaging large model (MILM). One popular strategy to build universal models is to encode each task as a one-hot vector and generate dynamic convolutional layers at the end of the decoder to extract the interested target. Although successful, it ignores the correlations among tasks and meanwhile is too late to make the model 'aware' of the ongoing task. To address both issues, we propose a prompt-driven Universal Segmentation model (UniSeg) for multi-task medical image segmentation using diverse modalities and domains. We first devise a learnable universal prompt to describe the correlations among all tasks and then convert this prompt and image features into a task-specific prompt, which is fed to the decoder as a part of its input. Thus, we make the model 'aware' of the ongoing task early and boost the task-specific training of the whole decoder. Our results indicate that the proposed UniSeg outperforms other universal models and single-task models on 11 upstream tasks. Moreover, UniSeg also beats other pre-trained models on two downstream datasets, providing the community with a high-quality pre-trained model for 3D medical image segmentation. Code and model are available at https://github.com/yeerwen/UniSeg.
Boundary-Aware Network for Kidney Parsing
Aug 29, 2022Shishuai Hu, Yiwen Ye, Zehui Liao, Yong Xia




Kidney structures segmentation is a crucial yet challenging task in the computer-aided diagnosis of surgery-based renal cancer. Although numerous deep learning models have achieved remarkable success in many medical image segmentation tasks, accurate segmentation of kidney structures on computed tomography angiography (CTA) images remains challenging, due to the variable sizes of kidney tumors and the ambiguous boundaries between kidney structures and their surroundings. In this paper, we propose a boundary-aware network (BA-Net) to segment kidneys, kidney tumors, arteries, and veins on CTA scans. This model contains a shared encoder, a boundary decoder, and a segmentation decoder. The multi-scale deep supervision strategy is adopted on both decoders, which can alleviate the issues caused by variable tumor sizes. The boundary probability maps produced by the boundary decoder at each scale are used as attention to enhance the segmentation feature maps. We evaluated the BA-Net on the Kidney PArsing (KiPA) Challenge dataset and achieved an average Dice score of 89.65$\%$ for kidney structure segmentation on CTA scans using 4-fold cross-validation. The results demonstrate the effectiveness of the BA-Net.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge