JointViT: Modeling Oxygen Saturation Levels with Joint Supervision on Long-Tailed OCTA
Apr 18, 2024Zeyu Zhang, Xuyin Qi, Mingxi Chen, Guangxi Li, Ryan Pham, Ayub Qassim, Ella Berry, Zhibin Liao, Owen Siggs, Robert Mclaughlin, Jamie Craig, Minh-Son To



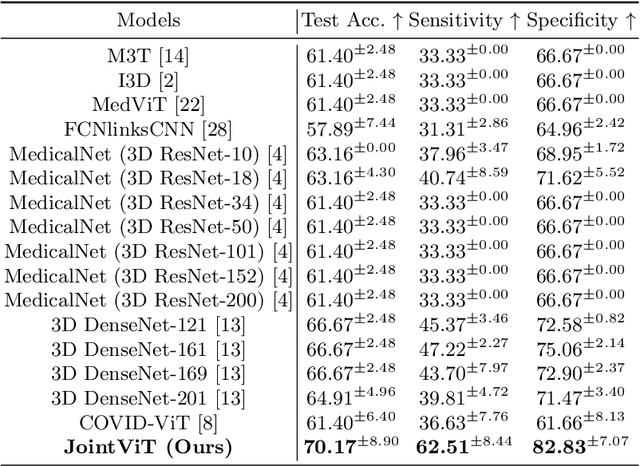
The oxygen saturation level in the blood (SaO2) is crucial for health, particularly in relation to sleep-related breathing disorders. However, continuous monitoring of SaO2 is time-consuming and highly variable depending on patients' conditions. Recently, optical coherence tomography angiography (OCTA) has shown promising development in rapidly and effectively screening eye-related lesions, offering the potential for diagnosing sleep-related disorders. To bridge this gap, our paper presents three key contributions. Firstly, we propose JointViT, a novel model based on the Vision Transformer architecture, incorporating a joint loss function for supervision. Secondly, we introduce a balancing augmentation technique during data preprocessing to improve the model's performance, particularly on the long-tail distribution within the OCTA dataset. Lastly, through comprehensive experiments on the OCTA dataset, our proposed method significantly outperforms other state-of-the-art methods, achieving improvements of up to 12.28% in overall accuracy. This advancement lays the groundwork for the future utilization of OCTA in diagnosing sleep-related disorders. See project website https://steve-zeyu-zhang.github.io/JointViT
CAPE: CAM as a Probabilistic Ensemble for Enhanced DNN Interpretation
Apr 04, 2024Townim Faisal Chowdhury, Kewen Liao, Vu Minh Hieu Phan, Minh-Son To, Yutong Xie, Kevin Hung, David Ross, Anton van den Hengel, Johan W. Verjans, Zhibin Liao
Deep Neural Networks (DNNs) are widely used for visual classification tasks, but their complex computation process and black-box nature hinder decision transparency and interpretability. Class activation maps (CAMs) and recent variants provide ways to visually explain the DNN decision-making process by displaying 'attention' heatmaps of the DNNs. Nevertheless, the CAM explanation only offers relative attention information, that is, on an attention heatmap, we can interpret which image region is more or less important than the others. However, these regions cannot be meaningfully compared across classes, and the contribution of each region to the model's class prediction is not revealed. To address these challenges that ultimately lead to better DNN Interpretation, in this paper, we propose CAPE, a novel reformulation of CAM that provides a unified and probabilistically meaningful assessment of the contributions of image regions. We quantitatively and qualitatively compare CAPE with state-of-the-art CAM methods on CUB and ImageNet benchmark datasets to demonstrate enhanced interpretability. We also test on a cytology imaging dataset depicting a challenging Chronic Myelomonocytic Leukemia (CMML) diagnosis problem. Code is available at: https://github.com/AIML-MED/CAPE.
Decomposing Disease Descriptions for Enhanced Pathology Detection: A Multi-Aspect Vision-Language Matching Framework
Mar 12, 2024Minh Hieu Phan, Yutong Xie, Yuankai Qi, Lingqiao Liu, Liyang Liu, Bowen Zhang, Zhibin Liao, Qi Wu, Minh-Son To, Johan W. Verjans
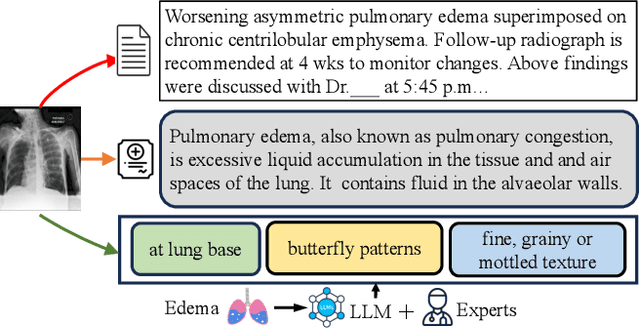
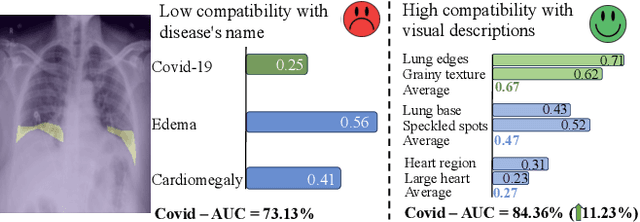
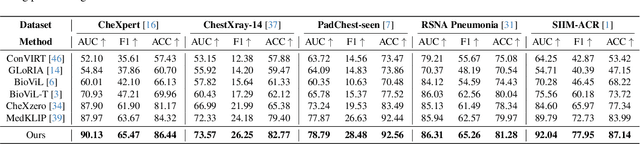

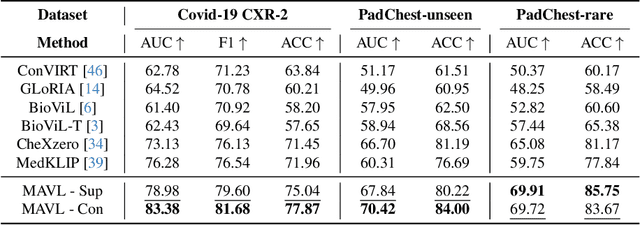
Medical vision language pre-training (VLP) has emerged as a frontier of research, enabling zero-shot pathological recognition by comparing the query image with the textual descriptions for each disease. Due to the complex semantics of biomedical texts, current methods struggle to align medical images with key pathological findings in unstructured reports. This leads to the misalignment with the target disease's textual representation. In this paper, we introduce a novel VLP framework designed to dissect disease descriptions into their fundamental aspects, leveraging prior knowledge about the visual manifestations of pathologies. This is achieved by consulting a large language model and medical experts. Integrating a Transformer module, our approach aligns an input image with the diverse elements of a disease, generating aspect-centric image representations. By consolidating the matches from each aspect, we improve the compatibility between an image and its associated disease. Additionally, capitalizing on the aspect-oriented representations, we present a dual-head Transformer tailored to process known and unknown diseases, optimizing the comprehensive detection efficacy. Conducting experiments on seven downstream datasets, ours outperforms recent methods by up to 8.07% and 11.23% in AUC scores for seen and novel categories, respectively. Our code is released at \href{https://github.com/HieuPhan33/MAVL}{https://github.com/HieuPhan33/MAVL}.
Structure-Preserving Synthesis: MaskGAN for Unpaired MR-CT Translation
Aug 01, 2023Minh Hieu Phan, Zhibin Liao, Johan W. Verjans, Minh-Son To
Medical image synthesis is a challenging task due to the scarcity of paired data. Several methods have applied CycleGAN to leverage unpaired data, but they often generate inaccurate mappings that shift the anatomy. This problem is further exacerbated when the images from the source and target modalities are heavily misaligned. Recently, current methods have aimed to address this issue by incorporating a supplementary segmentation network. Unfortunately, this strategy requires costly and time-consuming pixel-level annotations. To overcome this problem, this paper proposes MaskGAN, a novel and cost-effective framework that enforces structural consistency by utilizing automatically extracted coarse masks. Our approach employs a mask generator to outline anatomical structures and a content generator to synthesize CT contents that align with these structures. Extensive experiments demonstrate that MaskGAN outperforms state-of-the-art synthesis methods on a challenging pediatric dataset, where MR and CT scans are heavily misaligned due to rapid growth in children. Specifically, MaskGAN excels in preserving anatomical structures without the need for expert annotations. The code for this paper can be found at https://github.com/HieuPhan33/MaskGAN.
* Accepted to MICCAI 2023
CNN Attention Guidance for Improved Orthopedics Radiographic Fracture Classification
Mar 21, 2022Zhibin Liao, Kewen Liao, Haifeng Shen, Marouska F. van Boxel, Jasper Prijs, Ruurd L. Jaarsma, Job N. Doornberg, Anton van den Hengel, Johan W. Verjans




Convolutional neural networks (CNNs) have gained significant popularity in orthopedic imaging in recent years due to their ability to solve fracture classification problems. A common criticism of CNNs is their opaque learning and reasoning process, making it difficult to trust machine diagnosis and the subsequent adoption of such algorithms in clinical setting. This is especially true when the CNN is trained with limited amount of medical data, which is a common issue as curating sufficiently large amount of annotated medical imaging data is a long and costly process. While interest has been devoted to explaining CNN learnt knowledge by visualizing network attention, the utilization of the visualized attention to improve network learning has been rarely investigated. This paper explores the effectiveness of regularizing CNN network with human-provided attention guidance on where in the image the network should look for answering clues. On two orthopedics radiographic fracture classification datasets, through extensive experiments we demonstrate that explicit human-guided attention indeed can direct correct network attention and consequently significantly improve classification performance. The development code for the proposed attention guidance is publicly available on GitHub.
Pairwise Relation Learning for Semi-supervised Gland Segmentation
Aug 06, 2020Yutong Xie, Jianpeng Zhang, Zhibin Liao, Chunhua Shen, Johan Verjans, Yong Xia




Accurate and automated gland segmentation on histology tissue images is an essential but challenging task in the computer-aided diagnosis of adenocarcinoma. Despite their prevalence, deep learning models always require a myriad number of densely annotated training images, which are difficult to obtain due to extensive labor and associated expert costs related to histology image annotations. In this paper, we propose the pairwise relation-based semi-supervised (PRS^2) model for gland segmentation on histology images. This model consists of a segmentation network (S-Net) and a pairwise relation network (PR-Net). The S-Net is trained on labeled data for segmentation, and PR-Net is trained on both labeled and unlabeled data in an unsupervised way to enhance its image representation ability via exploiting the semantic consistency between each pair of images in the feature space. Since both networks share their encoders, the image representation ability learned by PR-Net can be transferred to S-Net to improve its segmentation performance. We also design the object-level Dice loss to address the issues caused by touching glands and combine it with other two loss functions for S-Net. We evaluated our model against five recent methods on the GlaS dataset and three recent methods on the CRAG dataset. Our results not only demonstrate the effectiveness of the proposed PR-Net and object-level Dice loss, but also indicate that our PRS^2 model achieves the state-of-the-art gland segmentation performance on both benchmarks.
On Modelling Label Uncertainty in Deep Neural Networks: Automatic Estimation of Intra-observer Variability in 2D Echocardiography Quality Assessment
Nov 02, 2019Zhibin Liao, Hany Girgis, Amir Abdi, Hooman Vaseli, Jorden Hetherington, Robert Rohling, Ken Gin, Teresa Tsang, Purang Abolmaesumi




Uncertainty of labels in clinical data resulting from intra-observer variability can have direct impact on the reliability of assessments made by deep neural networks. In this paper, we propose a method for modelling such uncertainty in the context of 2D echocardiography (echo), which is a routine procedure for detecting cardiovascular disease at point-of-care. Echo imaging quality and acquisition time is highly dependent on the operator's experience level. Recent developments have shown the possibility of automating echo image quality quantification by mapping an expert's assessment of quality to the echo image via deep learning techniques. Nevertheless, the observer variability in the expert's assessment can impact the quality quantification accuracy. Here, we aim to model the intra-observer variability in echo quality assessment as an aleatoric uncertainty modelling regression problem with the introduction of a novel method that handles the regression problem with categorical labels. A key feature of our design is that only a single forward pass is sufficient to estimate the level of uncertainty for the network output. Compared to the $0.11 \pm 0.09$ absolute error (in a scale from 0 to 1) archived by the conventional regression method, the proposed method brings the error down to $0.09 \pm 0.08$, where the improvement is statistically significant and equivalents to $5.7\%$ test accuracy improvement. The simplicity of the proposed approach means that it could be generalized to other applications of deep learning in medical imaging, where there is often uncertainty in clinical labels.
Approximate Fisher Information Matrix to Characterise the Training of Deep Neural Networks
Oct 16, 2018Zhibin Liao, Tom Drummond, Ian Reid, Gustavo Carneiro




In this paper, we introduce a novel methodology for characterising the performance of deep learning networks (ResNets and DenseNet) with respect to training convergence and generalisation as a function of mini-batch size and learning rate for image classification. This methodology is based on novel measurements derived from the eigenvalues of the approximate Fisher information matrix, which can be efficiently computed even for high capacity deep models. Our proposed measurements can help practitioners to monitor and control the training process (by actively tuning the mini-batch size and learning rate) to allow for good training convergence and generalisation. Furthermore, the proposed measurements also allow us to show that it is possible to optimise the training process with a new dynamic sampling training approach that continuously and automatically change the mini-batch size and learning rate during the training process. Finally, we show that the proposed dynamic sampling training approach has a faster training time and a competitive classification accuracy compared to the current state of the art.
Competitive Multi-scale Convolution
Nov 18, 2015Zhibin Liao, Gustavo Carneiro




In this paper, we introduce a new deep convolutional neural network (ConvNet) module that promotes competition among a set of multi-scale convolutional filters. This new module is inspired by the inception module, where we replace the original collaborative pooling stage (consisting of a concatenation of the multi-scale filter outputs) by a competitive pooling represented by a maxout activation unit. This extension has the following two objectives: 1) the selection of the maximum response among the multi-scale filters prevents filter co-adaptation and allows the formation of multiple sub-networks within the same model, which has been shown to facilitate the training of complex learning problems; and 2) the maxout unit reduces the dimensionality of the outputs from the multi-scale filters. We show that the use of our proposed module in typical deep ConvNets produces classification results that are either better than or comparable to the state of the art on the following benchmark datasets: MNIST, CIFAR-10, CIFAR-100 and SVHN.
On the Importance of Normalisation Layers in Deep Learning with Piecewise Linear Activation Units
Nov 01, 2015Zhibin Liao, Gustavo Carneiro




Deep feedforward neural networks with piecewise linear activations are currently producing the state-of-the-art results in several public datasets. The combination of deep learning models and piecewise linear activation functions allows for the estimation of exponentially complex functions with the use of a large number of subnetworks specialized in the classification of similar input examples. During the training process, these subnetworks avoid overfitting with an implicit regularization scheme based on the fact that they must share their parameters with other subnetworks. Using this framework, we have made an empirical observation that can improve even more the performance of such models. We notice that these models assume a balanced initial distribution of data points with respect to the domain of the piecewise linear activation function. If that assumption is violated, then the piecewise linear activation units can degenerate into purely linear activation units, which can result in a significant reduction of their capacity to learn complex functions. Furthermore, as the number of model layers increases, this unbalanced initial distribution makes the model ill-conditioned. Therefore, we propose the introduction of batch normalisation units into deep feedforward neural networks with piecewise linear activations, which drives a more balanced use of these activation units, where each region of the activation function is trained with a relatively large proportion of training samples. Also, this batch normalisation promotes the pre-conditioning of very deep learning models. We show that by introducing maxout and batch normalisation units to the network in network model results in a model that produces classification results that are better than or comparable to the current state of the art in CIFAR-10, CIFAR-100, MNIST, and SVHN datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge