Capabilities of Gemini Models in Medicine
May 01, 2024Khaled Saab, Tao Tu, Wei-Hung Weng, Ryutaro Tanno, David Stutz, Ellery Wulczyn, Fan Zhang, Tim Strother, Chunjong Park, Elahe Vedadi, Juanma Zambrano Chaves, Szu-Yeu Hu, Mike Schaekermann, Aishwarya Kamath, Yong Cheng, David G. T. Barrett, Cathy Cheung, Basil Mustafa, Anil Palepu, Daniel McDuff, Le Hou, Tomer Golany, Luyang Liu, Jean-baptiste Alayrac, Neil Houlsby, Nenad Tomasev, Jan Freyberg, Charles Lau, Jonas Kemp, Jeremy Lai, Shekoofeh Azizi, Kimberly Kanada, SiWai Man, Kavita Kulkarni, Ruoxi Sun, Siamak Shakeri, Luheng He, Ben Caine, Albert Webson, Natasha Latysheva, Melvin Johnson, Philip Mansfield, Jian Lu, Ehud Rivlin, Jesper Anderson, Bradley Green, Renee Wong, Jonathan Krause, Jonathon Shlens, Ewa Dominowska, S. M. Ali Eslami, Katherine Chou, Claire Cui, Oriol Vinyals, Koray Kavukcuoglu, James Manyika, Jeff Dean, Demis Hassabis, Yossi Matias, Dale Webster, Joelle Barral, Greg Corrado, Christopher Semturs, S. Sara Mahdavi, Juraj Gottweis, Alan Karthikesalingam, Vivek Natarajan
Excellence in a wide variety of medical applications poses considerable challenges for AI, requiring advanced reasoning, access to up-to-date medical knowledge and understanding of complex multimodal data. Gemini models, with strong general capabilities in multimodal and long-context reasoning, offer exciting possibilities in medicine. Building on these core strengths of Gemini, we introduce Med-Gemini, a family of highly capable multimodal models that are specialized in medicine with the ability to seamlessly use web search, and that can be efficiently tailored to novel modalities using custom encoders. We evaluate Med-Gemini on 14 medical benchmarks, establishing new state-of-the-art (SoTA) performance on 10 of them, and surpass the GPT-4 model family on every benchmark where a direct comparison is viable, often by a wide margin. On the popular MedQA (USMLE) benchmark, our best-performing Med-Gemini model achieves SoTA performance of 91.1% accuracy, using a novel uncertainty-guided search strategy. On 7 multimodal benchmarks including NEJM Image Challenges and MMMU (health & medicine), Med-Gemini improves over GPT-4V by an average relative margin of 44.5%. We demonstrate the effectiveness of Med-Gemini's long-context capabilities through SoTA performance on a needle-in-a-haystack retrieval task from long de-identified health records and medical video question answering, surpassing prior bespoke methods using only in-context learning. Finally, Med-Gemini's performance suggests real-world utility by surpassing human experts on tasks such as medical text summarization, alongside demonstrations of promising potential for multimodal medical dialogue, medical research and education. Taken together, our results offer compelling evidence for Med-Gemini's potential, although further rigorous evaluation will be crucial before real-world deployment in this safety-critical domain.
Predicting Generalization of AI Colonoscopy Models to Unseen Data
Mar 22, 2024Joel Shor, Carson McNeil, Yotam Intrator, Joseph R Ledsam, Hiro-o Yamano, Daisuke Tsurumaru, Hiroki Kayama, Atsushi Hamabe, Koji Ando, Mitsuhiko Ota, Haruei Ogino, Hiroshi Nakase, Kaho Kobayashi, Masaaki Miyo, Eiji Oki, Ichiro Takemasa, Ehud Rivlin, Roman Goldenberg
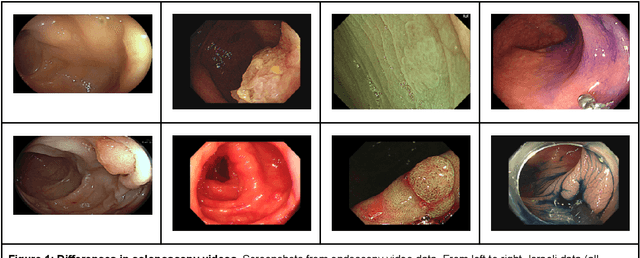
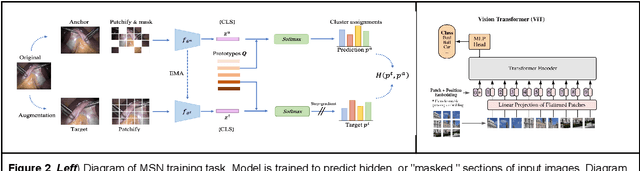
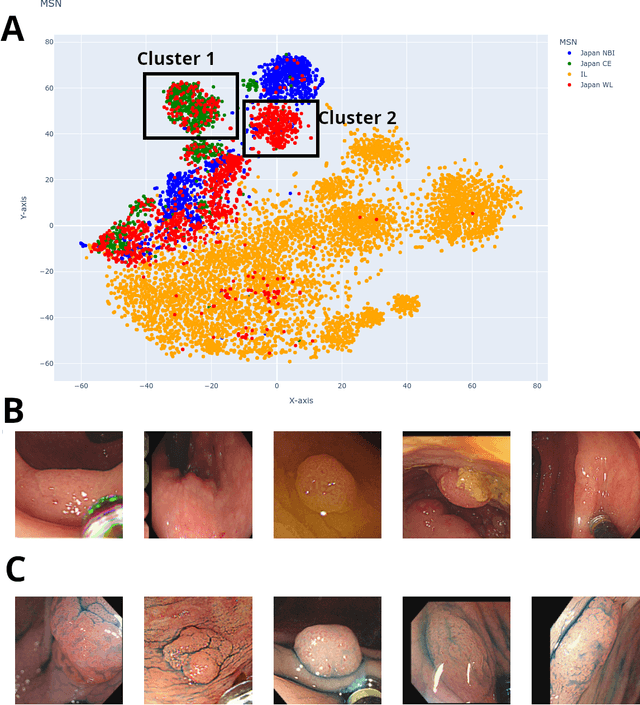
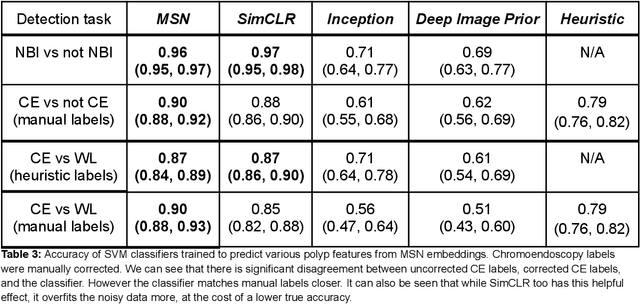
$\textbf{Background}$: Generalizability of AI colonoscopy algorithms is important for wider adoption in clinical practice. However, current techniques for evaluating performance on unseen data require expensive and time-intensive labels. $\textbf{Methods}$: We use a "Masked Siamese Network" (MSN) to identify novel phenomena in unseen data and predict polyp detector performance. MSN is trained to predict masked out regions of polyp images, without any labels. We test MSN's ability to be trained on data only from Israel and detect unseen techniques, narrow-band imaging (NBI) and chromendoscoy (CE), on colonoscopes from Japan (354 videos, 128 hours). We also test MSN's ability to predict performance of Computer Aided Detection (CADe) of polyps on colonoscopies from both countries, even though MSN is not trained on data from Japan. $\textbf{Results}$: MSN correctly identifies NBI and CE as less similar to Israel whitelight than Japan whitelight (bootstrapped z-test, |z| > 496, p < 10^-8 for both) using the label-free Frechet distance. MSN detects NBI with 99% accuracy, predicts CE better than our heuristic (90% vs 79% accuracy) despite being trained only on whitelight, and is the only method that is robust to noisy labels. MSN predicts CADe polyp detector performance on in-domain Israel and out-of-domain Japan colonoscopies (r=0.79, 0.37 respectively). With few examples of Japan detector performance to train on, MSN prediction of Japan performance improves (r=0.56). $\textbf{Conclusion}$: Our technique can identify distribution shifts in clinical data and can predict CADe detector performance on unseen data, without labels. Our self-supervised approach can aid in detecting when data in practice is different from training, such as between hospitals or data has meaningfully shifted from training. MSN has potential for application to medical image domains beyond colonoscopy.
Breaking the Language Barrier: Can Direct Inference Outperform Pre-Translation in Multilingual LLM Applications?
Mar 04, 2024Yotam Intrator, Matan Halfon, Roman Goldenberg, Reut Tsarfaty, Matan Eyal, Ehud Rivlin, Yossi Matias, Natalia Aizenberg

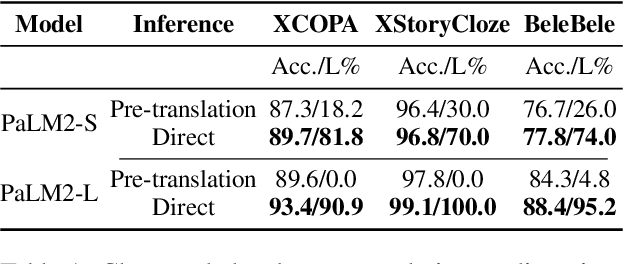
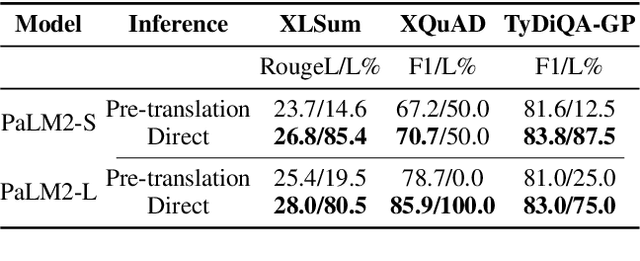
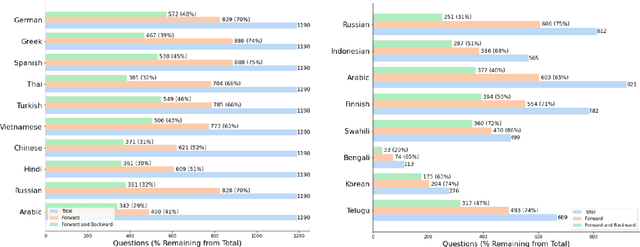
Large language models hold significant promise in multilingual applications. However, inherent biases stemming from predominantly English-centric pre-training have led to the widespread practice of pre-translation, i.e., translating non-English inputs to English before inference, leading to complexity and information loss. This study re-evaluates the need for pre-translation in the context of PaLM2 models (Anil et al., 2023), which have been established as highly performant in multilingual tasks. We offer a comprehensive investigation across 108 languages and 6 diverse benchmarks, including open-end generative tasks, which were excluded from previous similar studies. Our findings challenge the pre-translation paradigm established in prior research, highlighting the advantages of direct inference in PaLM2. Specifically, PaLM2-L consistently outperforms pre-translation in 94 out of 108 languages. These findings pave the way for more efficient and effective multilingual applications, alleviating the limitations associated with pre-translation and unlocking linguistic authenticity.
On the Semantic Latent Space of Diffusion-Based Text-to-Speech Models
Feb 19, 2024Miri Varshavsky Hassid, Roy Hirsch, Regev Cohen, Tomer Golany, Daniel Freedman, Ehud Rivlin
The incorporation of Denoising Diffusion Models (DDMs) in the Text-to-Speech (TTS) domain is rising, providing great value in synthesizing high quality speech. Although they exhibit impressive audio quality, the extent of their semantic capabilities is unknown, and controlling their synthesized speech's vocal properties remains a challenge. Inspired by recent advances in image synthesis, we explore the latent space of frozen TTS models, which is composed of the latent bottleneck activations of the DDM's denoiser. We identify that this space contains rich semantic information, and outline several novel methods for finding semantic directions within it, both supervised and unsupervised. We then demonstrate how these enable off-the-shelf audio editing, without any further training, architectural changes or data requirements. We present evidence of the semantic and acoustic qualities of the edited audio, and provide supplemental samples: https://latent-analysis-grad-tts.github.io/speech-samples/.
The unreasonable effectiveness of AI CADe polyp detectors to generalize to new countries
Dec 17, 2023Joel Shor, Hiro-o Yamano, Daisuke Tsurumaru, Yotami Intrator, Hiroki Kayama, Joe Ledsam, Atsushi Hamabe, Koji Ando, Mitsuhiko Ota, Haruei Ogino, Hiroshi Nakase, Kaho Kobayashi, Eiji Oki, Roman Goldenberg, Ehud Rivlin, Ichiro Takemasa
$\textbf{Background and aims}$: Artificial Intelligence (AI) Computer-Aided Detection (CADe) is commonly used for polyp detection, but data seen in clinical settings can differ from model training. Few studies evaluate how well CADe detectors perform on colonoscopies from countries not seen during training, and none are able to evaluate performance without collecting expensive and time-intensive labels. $\textbf{Methods}$: We trained a CADe polyp detector on Israeli colonoscopy videos (5004 videos, 1106 hours) and evaluated on Japanese videos (354 videos, 128 hours) by measuring the True Positive Rate (TPR) versus false alarms per minute (FAPM). We introduce a colonoscopy dissimilarity measure called "MAsked mediCal Embedding Distance" (MACE) to quantify differences between colonoscopies, without labels. We evaluated CADe on all Japan videos and on those with the highest MACE. $\textbf{Results}$: MACE correctly quantifies that narrow-band imaging (NBI) and chromoendoscopy (CE) frames are less similar to Israel data than Japan whitelight (bootstrapped z-test, |z| > 690, p < $10^{-8}$ for both). Despite differences in the data, CADe performance on Japan colonoscopies was non-inferior to Israel ones without additional training (TPR at 0.5 FAPM: 0.957 and 0.972 for Israel and Japan; TPR at 1.0 FAPM: 0.972 and 0.989 for Israel and Japan; superiority test t > 45.2, p < $10^{-8}$). Despite not being trained on NBI or CE, TPR on those subsets were non-inferior to Japan overall (non-inferiority test t > 47.3, p < $10^{-8}$, $\delta$ = 1.5% for both). $\textbf{Conclusion}$: Differences that prevent CADe detectors from performing well in non-medical settings do not degrade the performance of our AI CADe polyp detector when applied to data from a new country. MACE can help medical AI models internationalize by identifying the most "dissimilar" data on which to evaluate models.
Weakly-Supervised Surgical Phase Recognition
Oct 26, 2023Roy Hirsch, Regev Cohen, Mathilde Caron, Tomer Golany, Daniel Freedman, Ehud Rivlin
A key element of computer-assisted surgery systems is phase recognition of surgical videos. Existing phase recognition algorithms require frame-wise annotation of a large number of videos, which is time and money consuming. In this work we join concepts of graph segmentation with self-supervised learning to derive a random-walk solution for per-frame phase prediction. Furthermore, we utilize within our method two forms of weak supervision: sparse timestamps or few-shot learning. The proposed algorithm enjoys low complexity and can operate in lowdata regimes. We validate our method by running experiments with the public Cholec80 dataset of laparoscopic cholecystectomy videos, demonstrating promising performance in multiple setups.
Self-Supervised Learning for Endoscopic Video Analysis
Aug 23, 2023Roy Hirsch, Mathilde Caron, Regev Cohen, Amir Livne, Ron Shapiro, Tomer Golany, Roman Goldenberg, Daniel Freedman, Ehud Rivlin
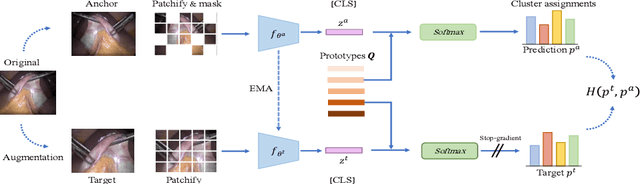



Self-supervised learning (SSL) has led to important breakthroughs in computer vision by allowing learning from large amounts of unlabeled data. As such, it might have a pivotal role to play in biomedicine where annotating data requires a highly specialized expertise. Yet, there are many healthcare domains for which SSL has not been extensively explored. One such domain is endoscopy, minimally invasive procedures which are commonly used to detect and treat infections, chronic inflammatory diseases or cancer. In this work, we study the use of a leading SSL framework, namely Masked Siamese Networks (MSNs), for endoscopic video analysis such as colonoscopy and laparoscopy. To fully exploit the power of SSL, we create sizable unlabeled endoscopic video datasets for training MSNs. These strong image representations serve as a foundation for secondary training with limited annotated datasets, resulting in state-of-the-art performance in endoscopic benchmarks like surgical phase recognition during laparoscopy and colonoscopic polyp characterization. Additionally, we achieve a 50% reduction in annotated data size without sacrificing performance. Thus, our work provides evidence that SSL can dramatically reduce the need of annotated data in endoscopy.
Self-Supervised Polyp Re-Identification in Colonoscopy
Jun 14, 2023Yotam Intrator, Natalie Aizenberg, Amir Livne, Ehud Rivlin, Roman Goldenberg
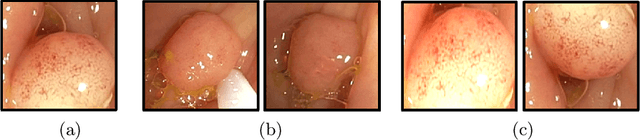
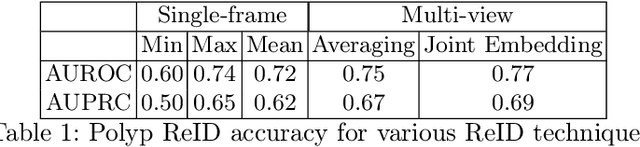
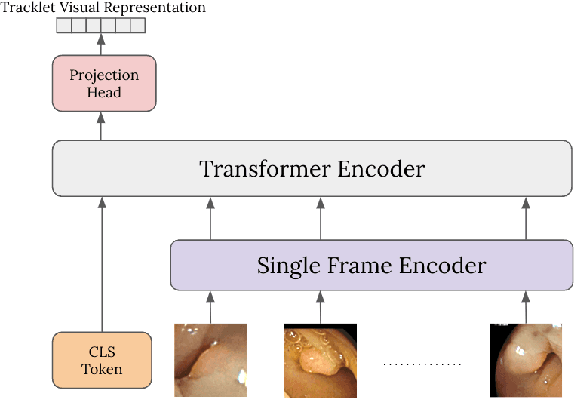

Computer-aided polyp detection (CADe) is becoming a standard, integral part of any modern colonoscopy system. A typical colonoscopy CADe detects a polyp in a single frame and does not track it through the video sequence. Yet, many downstream tasks including polyp characterization (CADx), quality metrics, automatic reporting, require aggregating polyp data from multiple frames. In this work we propose a robust long term polyp tracking method based on re-identification by visual appearance. Our solution uses an attention-based self-supervised ML model, specifically designed to leverage the temporal nature of video input. We quantitatively evaluate method's performance and demonstrate its value for the CADx task.
Semantic Parsing of Colonoscopy Videos with Multi-Label Temporal Networks
Jun 12, 2023Ori Kelner, Or Weinstein, Ehud Rivlin, Roman Goldenberg
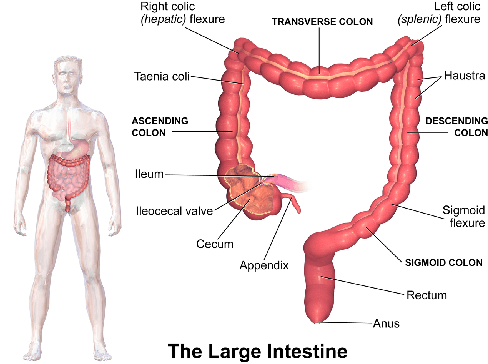
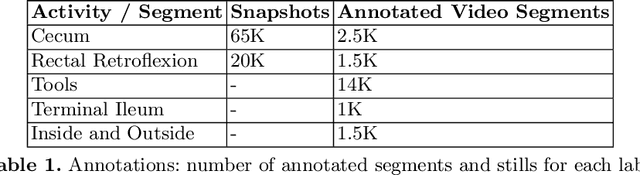
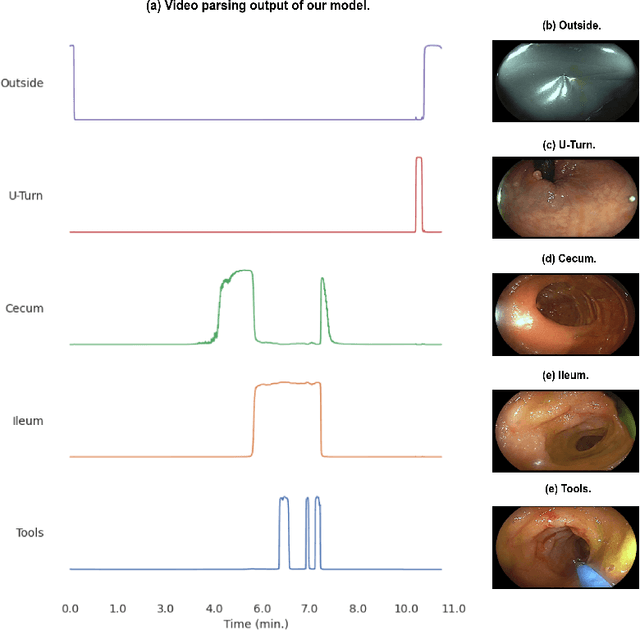

Following the successful debut of polyp detection and characterization, more advanced automation tools are being developed for colonoscopy. The new automation tasks, such as quality metrics or report generation, require understanding of the procedure flow that includes activities, events, anatomical landmarks, etc. In this work we present a method for automatic semantic parsing of colonoscopy videos. The method uses a novel DL multi-label temporal segmentation model trained in supervised and unsupervised regimes. We evaluate the accuracy of the method on a test set of over 300 annotated colonoscopy videos, and use ablation to explore the relative importance of various method's components.
Semi-supervised Quality Evaluation of Colonoscopy Procedures
May 17, 2023Idan Kligvasser, George Leifman, Roman Goldenberg, Ehud Rivlin, Michael Elad
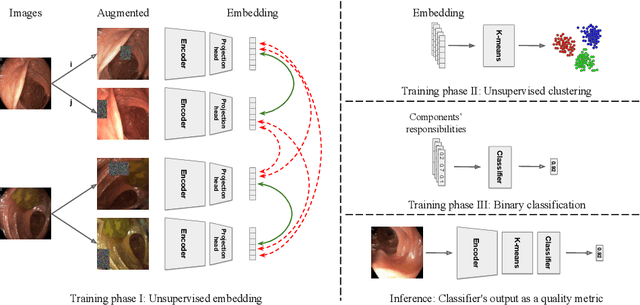


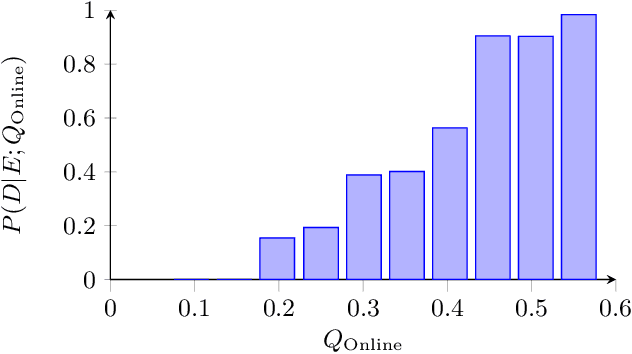
Colonoscopy is the standard of care technique for detecting and removing polyps for the prevention of colorectal cancer. Nevertheless, gastroenterologists (GI) routinely miss approximately 25% of polyps during colonoscopies. These misses are highly operator dependent, influenced by the physician skills, experience, vigilance, and fatigue. Standard quality metrics, such as Withdrawal Time or Cecal Intubation Rate, have been shown to be well correlated with Adenoma Detection Rate (ADR). However, those metrics are limited in their ability to assess the quality of a specific procedure, and they do not address quality aspects related to the style or technique of the examination. In this work we design novel online and offline quality metrics, based on visual appearance quality criteria learned by an ML model in an unsupervised way. Furthermore, we evaluate the likelihood of detecting an existing polyp as a function of quality and use it to demonstrate high correlation of the proposed metric to polyp detection sensitivity. The proposed online quality metric can be used to provide real time quality feedback to the performing GI. By integrating the local metric over the withdrawal phase, we build a global, offline quality metric, which is shown to be highly correlated to the standard Polyp Per Colonoscopy (PPC) quality metric.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge