Capabilities of Gemini Models in Medicine
May 01, 2024Khaled Saab, Tao Tu, Wei-Hung Weng, Ryutaro Tanno, David Stutz, Ellery Wulczyn, Fan Zhang, Tim Strother, Chunjong Park, Elahe Vedadi, Juanma Zambrano Chaves, Szu-Yeu Hu, Mike Schaekermann, Aishwarya Kamath, Yong Cheng, David G. T. Barrett, Cathy Cheung, Basil Mustafa, Anil Palepu, Daniel McDuff, Le Hou, Tomer Golany, Luyang Liu, Jean-baptiste Alayrac, Neil Houlsby, Nenad Tomasev, Jan Freyberg, Charles Lau, Jonas Kemp, Jeremy Lai, Shekoofeh Azizi, Kimberly Kanada, SiWai Man, Kavita Kulkarni, Ruoxi Sun, Siamak Shakeri, Luheng He, Ben Caine, Albert Webson, Natasha Latysheva, Melvin Johnson, Philip Mansfield, Jian Lu, Ehud Rivlin, Jesper Anderson, Bradley Green, Renee Wong, Jonathan Krause, Jonathon Shlens, Ewa Dominowska, S. M. Ali Eslami, Katherine Chou, Claire Cui, Oriol Vinyals, Koray Kavukcuoglu, James Manyika, Jeff Dean, Demis Hassabis, Yossi Matias, Dale Webster, Joelle Barral, Greg Corrado, Christopher Semturs, S. Sara Mahdavi, Juraj Gottweis, Alan Karthikesalingam, Vivek Natarajan
Excellence in a wide variety of medical applications poses considerable challenges for AI, requiring advanced reasoning, access to up-to-date medical knowledge and understanding of complex multimodal data. Gemini models, with strong general capabilities in multimodal and long-context reasoning, offer exciting possibilities in medicine. Building on these core strengths of Gemini, we introduce Med-Gemini, a family of highly capable multimodal models that are specialized in medicine with the ability to seamlessly use web search, and that can be efficiently tailored to novel modalities using custom encoders. We evaluate Med-Gemini on 14 medical benchmarks, establishing new state-of-the-art (SoTA) performance on 10 of them, and surpass the GPT-4 model family on every benchmark where a direct comparison is viable, often by a wide margin. On the popular MedQA (USMLE) benchmark, our best-performing Med-Gemini model achieves SoTA performance of 91.1% accuracy, using a novel uncertainty-guided search strategy. On 7 multimodal benchmarks including NEJM Image Challenges and MMMU (health & medicine), Med-Gemini improves over GPT-4V by an average relative margin of 44.5%. We demonstrate the effectiveness of Med-Gemini's long-context capabilities through SoTA performance on a needle-in-a-haystack retrieval task from long de-identified health records and medical video question answering, surpassing prior bespoke methods using only in-context learning. Finally, Med-Gemini's performance suggests real-world utility by surpassing human experts on tasks such as medical text summarization, alongside demonstrations of promising potential for multimodal medical dialogue, medical research and education. Taken together, our results offer compelling evidence for Med-Gemini's potential, although further rigorous evaluation will be crucial before real-world deployment in this safety-critical domain.
Adaptive Collaboration Strategy for LLMs in Medical Decision Making
Apr 22, 2024Yubin Kim, Chanwoo Park, Hyewon Jeong, Yik Siu Chan, Xuhai Xu, Daniel McDuff, Cynthia Breazeal, Hae Won Park
Foundation models have become invaluable in advancing the medical field. Despite their promise, the strategic deployment of LLMs for effective utility in complex medical tasks remains an open question. Our novel framework, Medical Decision-making Agents (MDAgents) aims to address this gap by automatically assigning the effective collaboration structure for LLMs. Assigned solo or group collaboration structure is tailored to the complexity of the medical task at hand, emulating real-world medical decision making processes. We evaluate our framework and baseline methods with state-of-the-art LLMs across a suite of challenging medical benchmarks: MedQA, MedMCQA, PubMedQA, DDXPlus, PMC-VQA, Path-VQA, and MedVidQA, achieving the best performance in 5 out of 7 benchmarks that require an understanding of multi-modal medical reasoning. Ablation studies reveal that MDAgents excels in adapting the number of collaborating agents to optimize efficiency and accuracy, showcasing its robustness in diverse scenarios. We also explore the dynamics of group consensus, offering insights into how collaborative agents could behave in complex clinical team dynamics. Our code can be found at https://github.com/mitmedialab/MDAgents.
How Suboptimal is Training rPPG Models with Videos and Targets from Different Body Sites?
Mar 15, 2024Björn Braun, Daniel McDuff, Christian Holz




Remote camera measurement of the blood volume pulse via photoplethysmography (rPPG) is a compelling technology for scalable, low-cost, and accessible assessment of cardiovascular information. Neural networks currently provide the state-of-the-art for this task and supervised training or fine-tuning is an important step in creating these models. However, most current models are trained on facial videos using contact PPG measurements from the fingertip as targets/ labels. One of the reasons for this is that few public datasets to date have incorporated contact PPG measurements from the face. Yet there is copious evidence that the PPG signals at different sites on the body have very different morphological features. Is training a facial video rPPG model using contact measurements from another site on the body suboptimal? Using a recently released unique dataset with synchronized contact PPG and video measurements from both the hand and face, we can provide precise and quantitative answers to this question. We obtain up to 40 % lower mean squared errors between the waveforms of the predicted and the ground truth PPG signals using state-of-the-art neural models when using PPG signals from the forehead compared to using PPG signals from the fingertip. We also show qualitatively that the neural models learn to predict the morphology of the ground truth PPG signal better when trained on the forehead PPG signals. However, while models trained from the forehead PPG produce a more faithful waveform, models trained from a finger PPG do still learn the dominant frequency (i.e., the heart rate) well.
On the Standardization of Behavioral Use Clauses and Their Adoption for Responsible Licensing of AI
Feb 07, 2024Daniel McDuff, Tim Korjakow, Scott Cambo, Jesse Josua Benjamin, Jenny Lee, Yacine Jernite, Carlos Muñoz Ferrandis, Aaron Gokaslan, Alek Tarkowski, Joseph Lindley, A. Feder Cooper, Danish Contractor
Growing concerns over negligent or malicious uses of AI have increased the appetite for tools that help manage the risks of the technology. In 2018, licenses with behaviorial-use clauses (commonly referred to as Responsible AI Licenses) were proposed to give developers a framework for releasing AI assets while specifying their users to mitigate negative applications. As of the end of 2023, on the order of 40,000 software and model repositories have adopted responsible AI licenses licenses. Notable models licensed with behavioral use clauses include BLOOM (language) and LLaMA2 (language), Stable Diffusion (image), and GRID (robotics). This paper explores why and how these licenses have been adopted, and why and how they have been adapted to fit particular use cases. We use a mixed-methods methodology of qualitative interviews, clustering of license clauses, and quantitative analysis of license adoption. Based on this evidence we take the position that responsible AI licenses need standardization to avoid confusing users or diluting their impact. At the same time, customization of behavioral restrictions is also appropriate in some contexts (e.g., medical domains). We advocate for ``standardized customization'' that can meet users' needs and can be supported via tooling.
Health-LLM: Large Language Models for Health Prediction via Wearable Sensor Data
Jan 12, 2024Yubin Kim, Xuhai Xu, Daniel McDuff, Cynthia Breazeal, Hae Won Park
Large language models (LLMs) are capable of many natural language tasks, yet they are far from perfect. In health applications, grounding and interpreting domain-specific and non-linguistic data is important. This paper investigates the capacity of LLMs to deliver multi-modal health predictions based on contextual information (e.g. user demographics, health knowledge) and physiological data (e.g. resting heart rate, sleep minutes). We present a comprehensive evaluation of eight state-of-the-art LLMs with diverse prompting and fine-tuning techniques on six public health datasets (PM-Data, LifeSnaps, GLOBEM, AW_FB, MIT-BIH & MIMIC-III). Our experiments cover thirteen consumer health prediction tasks in mental health, activity, metabolic, sleep, and cardiac assessment. Our fine-tuned model, Health-Alpaca exhibits comparable performance to larger models (GPT-3.5 and GPT-4), achieving the best performance in 5 out of 13 tasks. Ablation studies highlight the effectiveness of context enhancement strategies, and generalization capability of the fine-tuned models across training datasets and the size of training samples. Notably, we observe that our context enhancement can yield up to 23.8% improvement in performance. While constructing contextually rich prompts (combining user context, health knowledge and temporal information) exhibits synergistic improvement, the inclusion of health knowledge context in prompts significantly enhances overall performance.
Towards Accurate Differential Diagnosis with Large Language Models
Nov 30, 2023Daniel McDuff, Mike Schaekermann, Tao Tu, Anil Palepu, Amy Wang, Jake Garrison, Karan Singhal, Yash Sharma, Shekoofeh Azizi, Kavita Kulkarni, Le Hou, Yong Cheng, Yun Liu, S Sara Mahdavi, Sushant Prakash, Anupam Pathak, Christopher Semturs, Shwetak Patel, Dale R Webster, Ewa Dominowska, Juraj Gottweis, Joelle Barral, Katherine Chou, Greg S Corrado, Yossi Matias, Jake Sunshine, Alan Karthikesalingam, Vivek Natarajan
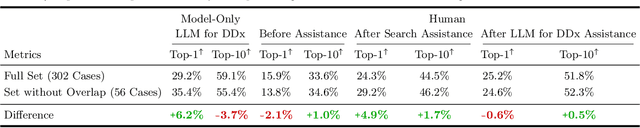
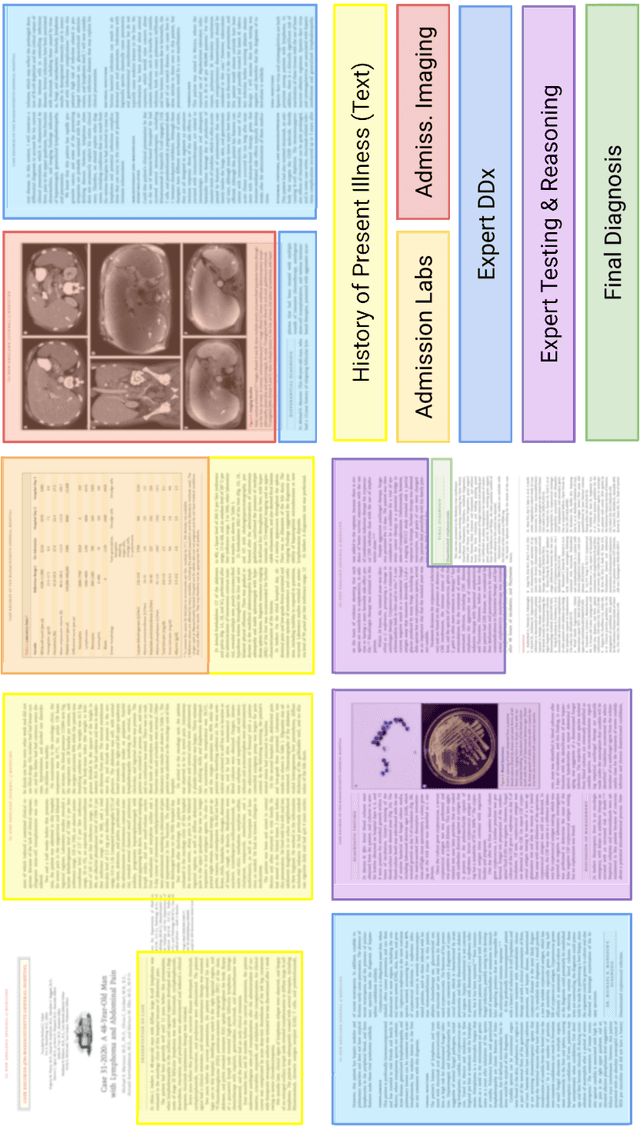
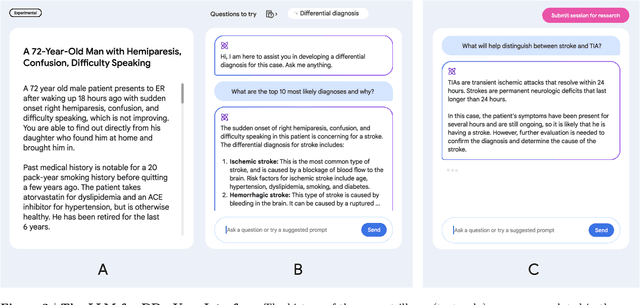
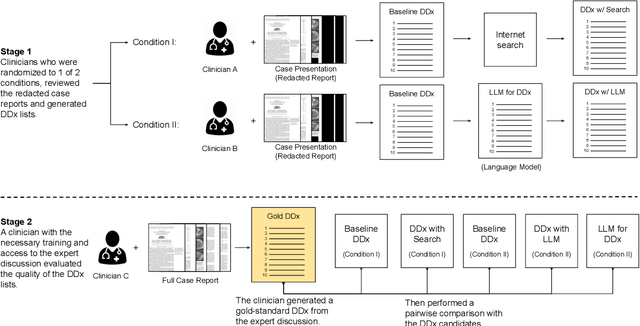
An accurate differential diagnosis (DDx) is a cornerstone of medical care, often reached through an iterative process of interpretation that combines clinical history, physical examination, investigations and procedures. Interactive interfaces powered by Large Language Models (LLMs) present new opportunities to both assist and automate aspects of this process. In this study, we introduce an LLM optimized for diagnostic reasoning, and evaluate its ability to generate a DDx alone or as an aid to clinicians. 20 clinicians evaluated 302 challenging, real-world medical cases sourced from the New England Journal of Medicine (NEJM) case reports. Each case report was read by two clinicians, who were randomized to one of two assistive conditions: either assistance from search engines and standard medical resources, or LLM assistance in addition to these tools. All clinicians provided a baseline, unassisted DDx prior to using the respective assistive tools. Our LLM for DDx exhibited standalone performance that exceeded that of unassisted clinicians (top-10 accuracy 59.1% vs 33.6%, [p = 0.04]). Comparing the two assisted study arms, the DDx quality score was higher for clinicians assisted by our LLM (top-10 accuracy 51.7%) compared to clinicians without its assistance (36.1%) (McNemar's Test: 45.7, p < 0.01) and clinicians with search (44.4%) (4.75, p = 0.03). Further, clinicians assisted by our LLM arrived at more comprehensive differential lists than those without its assistance. Our study suggests that our LLM for DDx has potential to improve clinicians' diagnostic reasoning and accuracy in challenging cases, meriting further real-world evaluation for its ability to empower physicians and widen patients' access to specialist-level expertise.
From Classification to Clinical Insights: Towards Analyzing and Reasoning About Mobile and Behavioral Health Data With Large Language Models
Nov 25, 2023Zachary Englhardt, Chengqian Ma, Margaret E. Morris, Xuhai "Orson" Xu, Chun-Cheng Chang, Lianhui Qin, Daniel McDuff, Xin Liu, Shwetak Patel, Vikram Iyer
Passively collected behavioral health data from ubiquitous sensors holds significant promise to provide mental health professionals insights from patient's daily lives; however, developing analysis tools to use this data in clinical practice requires addressing challenges of generalization across devices and weak or ambiguous correlations between the measured signals and an individual's mental health. To address these challenges, we take a novel approach that leverages large language models (LLMs) to synthesize clinically useful insights from multi-sensor data. We develop chain of thought prompting methods that use LLMs to generate reasoning about how trends in data such as step count and sleep relate to conditions like depression and anxiety. We first demonstrate binary depression classification with LLMs achieving accuracies of 61.1% which exceed the state of the art. While it is not robust for clinical use, this leads us to our key finding: even more impactful and valued than classification is a new human-AI collaboration approach in which clinician experts interactively query these tools and combine their domain expertise and context about the patient with AI generated reasoning to support clinical decision-making. We find models like GPT-4 correctly reference numerical data 75% of the time, and clinician participants express strong interest in using this approach to interpret self-tracking data.
Video-based sympathetic arousal assessment via peripheral blood flow estimation
Nov 12, 2023Bjoern Braun, Daniel McDuff, Tadas Baltrusaitis, Christian Holz
Electrodermal activity (EDA) is considered a standard marker of sympathetic activity. However, traditional EDA measurement requires electrodes in steady contact with the skin. Can sympathetic arousal be measured using only an optical sensor, such as an RGB camera? This paper presents a novel approach to infer sympathetic arousal by measuring the peripheral blood flow on the face or hand optically. We contribute a self-recorded dataset of 21 participants, comprising synchronized videos of participants' faces and palms and gold-standard EDA and photoplethysmography (PPG) signals. Our results show that we can measure peripheral sympathetic responses that closely correlate with the ground truth EDA. We obtain median correlations of 0.57 to 0.63 between our inferred signals and the ground truth EDA using only videos of the participants' palms or foreheads or PPG signals from the foreheads or fingers. We also show that sympathetic arousal is best inferred from the forehead, finger, or palm.
The Capability of Large Language Models to Measure Psychiatric Functioning
Aug 03, 2023Isaac R. Galatzer-Levy, Daniel McDuff, Vivek Natarajan, Alan Karthikesalingam, Matteo Malgaroli
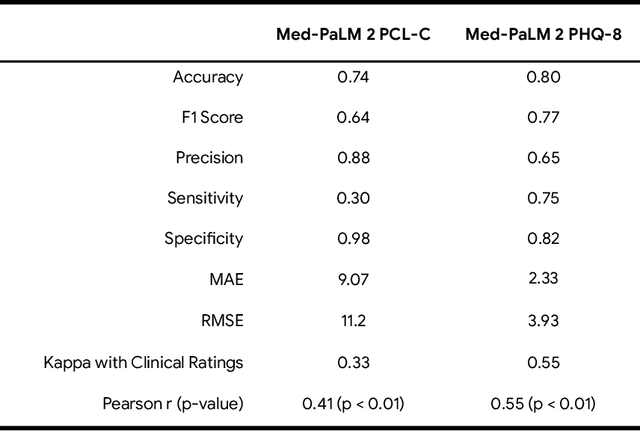
The current work investigates the capability of Large language models (LLMs) that are explicitly trained on large corpuses of medical knowledge (Med-PaLM 2) to predict psychiatric functioning from patient interviews and clinical descriptions without being trained to do so. To assess this, n = 145 depression and n =115 PTSD assessments and n = 46 clinical case studies across high prevalence/high comorbidity disorders (Depressive, Anxiety, Psychotic, trauma and stress, Addictive disorders) were analyzed using prompts to extract estimated clinical scores and diagnoses. Results demonstrate that Med-PaLM 2 is capable of assessing psychiatric functioning across a range of psychiatric conditions with the strongest performance being the prediction of depression scores based on standardized assessments (Accuracy range= 0.80 - 0.84) which were statistically indistinguishable from human clinical raters t(1,144) = 1.20; p = 0.23. Results show the potential for general clinical language models to flexibly predict psychiatric risk based on free descriptions of functioning from both patients and clinicians.
Large Language Models are Few-Shot Health Learners
May 24, 2023Xin Liu, Daniel McDuff, Geza Kovacs, Isaac Galatzer-Levy, Jacob Sunshine, Jiening Zhan, Ming-Zher Poh, Shun Liao, Paolo Di Achille, Shwetak Patel

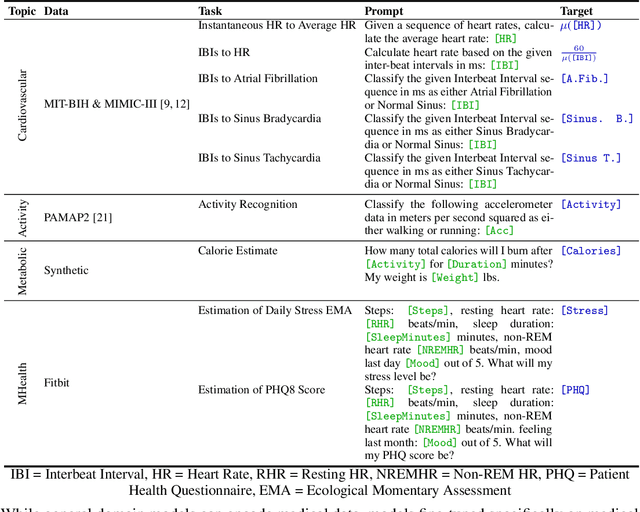
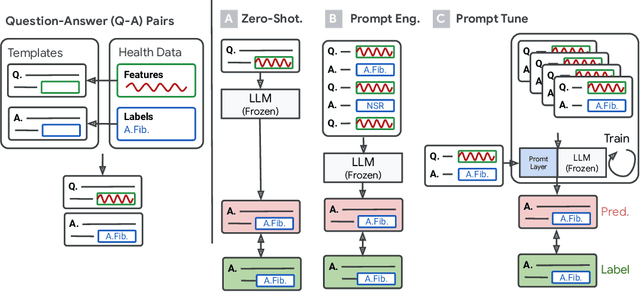
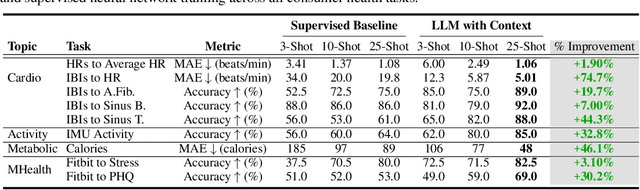
Large language models (LLMs) can capture rich representations of concepts that are useful for real-world tasks. However, language alone is limited. While existing LLMs excel at text-based inferences, health applications require that models be grounded in numerical data (e.g., vital signs, laboratory values in clinical domains; steps, movement in the wellness domain) that is not easily or readily expressed as text in existing training corpus. We demonstrate that with only few-shot tuning, a large language model is capable of grounding various physiological and behavioral time-series data and making meaningful inferences on numerous health tasks for both clinical and wellness contexts. Using data from wearable and medical sensor recordings, we evaluate these capabilities on the tasks of cardiac signal analysis, physical activity recognition, metabolic calculation (e.g., calories burned), and estimation of stress reports and mental health screeners.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge