Investigating Generalization Behaviours of Generative Flow Networks
Feb 07, 2024Lazar Atanackovic, Emmanuel Bengio
Generative Flow Networks (GFlowNets, GFNs) are a generative framework for learning unnormalized probability mass functions over discrete spaces. Since their inception, GFlowNets have proven to be useful for learning generative models in applications where the majority of the discrete space is unvisited during training. This has inspired some to hypothesize that GFlowNets, when paired with deep neural networks (DNNs), have favourable generalization properties. In this work, we empirically verify some of the hypothesized mechanisms of generalization of GFlowNets. In particular, we find that the functions that GFlowNets learn to approximate have an implicit underlying structure which facilitate generalization. We also find that GFlowNets are sensitive to being trained offline and off-policy; however, the reward implicitly learned by GFlowNets is robust to changes in the training distribution.
QGFN: Controllable Greediness with Action Values
Feb 07, 2024Elaine Lau, Stephen Zhewen Lu, Ling Pan, Doina Precup, Emmanuel Bengio
Generative Flow Networks (GFlowNets; GFNs) are a family of reward/energy-based generative methods for combinatorial objects, capable of generating diverse and high-utility samples. However, biasing GFNs towards producing high-utility samples is non-trivial. In this work, we leverage connections between GFNs and reinforcement learning (RL) and propose to combine the GFN policy with an action-value estimate, $Q$, to create greedier sampling policies which can be controlled by a mixing parameter. We show that several variants of the proposed method, QGFN, are able to improve on the number of high-reward samples generated in a variety of tasks without sacrificing diversity.
Maximum entropy GFlowNets with soft Q-learning
Dec 21, 2023Sobhan Mohammadpour, Emmanuel Bengio, Emma Frejinger, Pierre-Luc Bacon
Generative Flow Networks (GFNs) have emerged as a powerful tool for sampling discrete objects from unnormalized distributions, offering a scalable alternative to Markov Chain Monte Carlo (MCMC) methods. While GFNs draw inspiration from maximum entropy reinforcement learning (RL), the connection between the two has largely been unclear and seemingly applicable only in specific cases. This paper addresses the connection by constructing an appropriate reward function, thereby establishing an exact relationship between GFNs and maximum entropy RL. This construction allows us to introduce maximum entropy GFNs, which, in contrast to GFNs with uniform backward policy, achieve the maximum entropy attainable by GFNs without constraints on the state space.
DGFN: Double Generative Flow Networks
Nov 06, 2023Elaine Lau, Nikhil Vemgal, Doina Precup, Emmanuel Bengio
Deep learning is emerging as an effective tool in drug discovery, with potential applications in both predictive and generative models. Generative Flow Networks (GFlowNets/GFNs) are a recently introduced method recognized for the ability to generate diverse candidates, in particular in small molecule generation tasks. In this work, we introduce double GFlowNets (DGFNs). Drawing inspiration from reinforcement learning and Double Deep Q-Learning, we introduce a target network used to sample trajectories, while updating the main network with these sampled trajectories. Empirical results confirm that DGFNs effectively enhance exploration in sparse reward domains and high-dimensional state spaces, both challenging aspects of de-novo design in drug discovery.
Local Search GFlowNets
Oct 04, 2023Minsu Kim, Taeyoung Yun, Emmanuel Bengio, Dinghuai Zhang, Yoshua Bengio, Sungsoo Ahn, Jinkyoo Park
Generative Flow Networks (GFlowNets) are amortized sampling methods that learn a distribution over discrete objects proportional to their rewards. GFlowNets exhibit a remarkable ability to generate diverse samples, yet occasionally struggle to consistently produce samples with high rewards due to over-exploration on wide sample space. This paper proposes to train GFlowNets with local search which focuses on exploiting high rewarded sample space to resolve this issue. Our main idea is to explore the local neighborhood via destruction and reconstruction guided by backward and forward policies, respectively. This allows biasing the samples toward high-reward solutions, which is not possible for a typical GFlowNet solution generation scheme which uses the forward policy to generate the solution from scratch. Extensive experiments demonstrate a remarkable performance improvement in several biochemical tasks. Source code is available: \url{https://github.com/dbsxodud-11/ls_gfn}.
Goal-conditioned GFlowNets for Controllable Multi-Objective Molecular Design
Jun 29, 2023Julien Roy, Pierre-Luc Bacon, Christopher Pal, Emmanuel Bengio
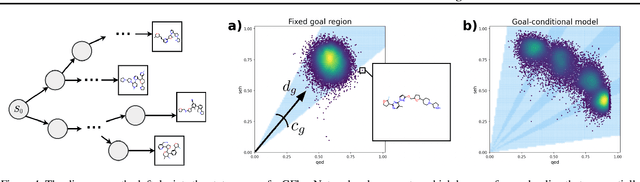
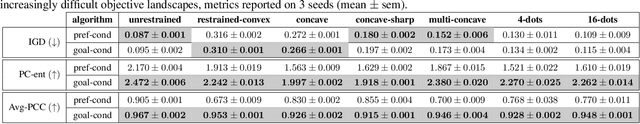
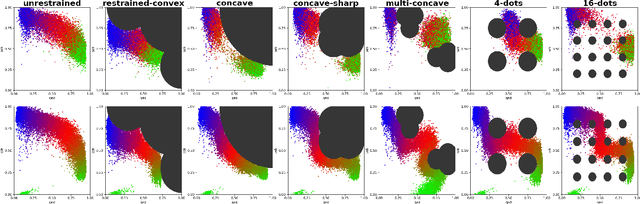
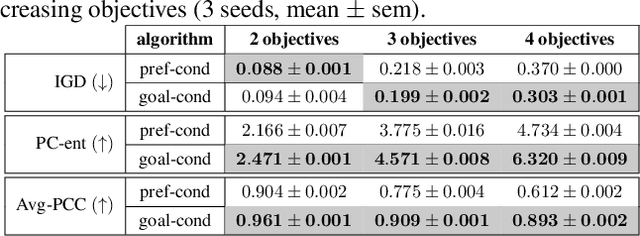
In recent years, in-silico molecular design has received much attention from the machine learning community. When designing a new compound for pharmaceutical applications, there are usually multiple properties of such molecules that need to be optimised: binding energy to the target, synthesizability, toxicity, EC50, and so on. While previous approaches have employed a scalarization scheme to turn the multi-objective problem into a preference-conditioned single objective, it has been established that this kind of reduction may produce solutions that tend to slide towards the extreme points of the objective space when presented with a problem that exhibits a concave Pareto front. In this work we experiment with an alternative formulation of goal-conditioned molecular generation to obtain a more controllable conditional model that can uniformly explore solutions along the entire Pareto front.
Towards Understanding and Improving GFlowNet Training
May 11, 2023Max W. Shen, Emmanuel Bengio, Ehsan Hajiramezanali, Andreas Loukas, Kyunghyun Cho, Tommaso Biancalani
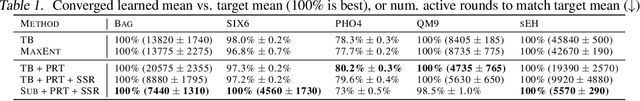

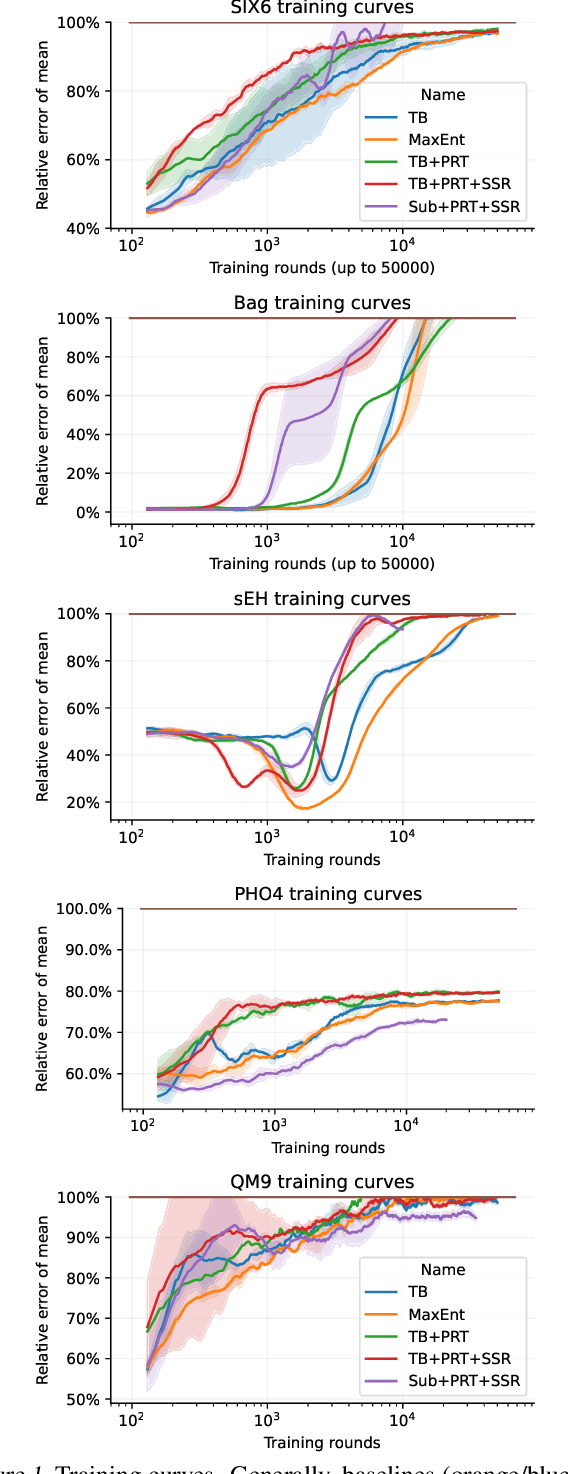
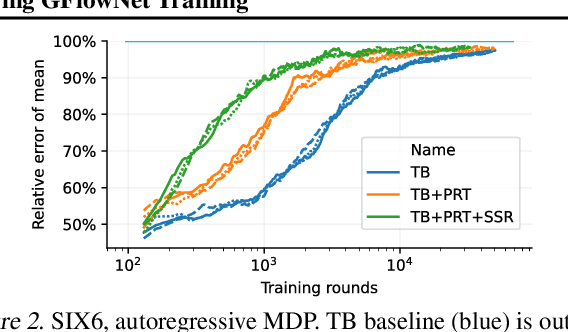
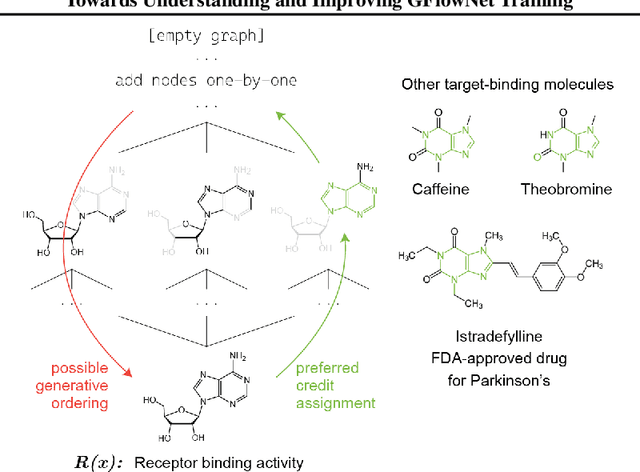
Generative flow networks (GFlowNets) are a family of algorithms that learn a generative policy to sample discrete objects $x$ with non-negative reward $R(x)$. Learning objectives guarantee the GFlowNet samples $x$ from the target distribution $p^*(x) \propto R(x)$ when loss is globally minimized over all states or trajectories, but it is unclear how well they perform with practical limits on training resources. We introduce an efficient evaluation strategy to compare the learned sampling distribution to the target reward distribution. As flows can be underdetermined given training data, we clarify the importance of learned flows to generalization and matching $p^*(x)$ in practice. We investigate how to learn better flows, and propose (i) prioritized replay training of high-reward $x$, (ii) relative edge flow policy parametrization, and (iii) a novel guided trajectory balance objective, and show how it can solve a substructure credit assignment problem. We substantially improve sample efficiency on biochemical design tasks.
Multi-Objective GFlowNets
Oct 23, 2022Moksh Jain, Sharath Chandra Raparthy, Alex Hernandez-Garcia, Jarrid Rector-Brooks, Yoshua Bengio, Santiago Miret, Emmanuel Bengio
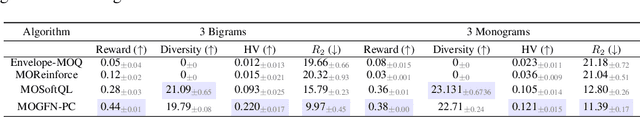
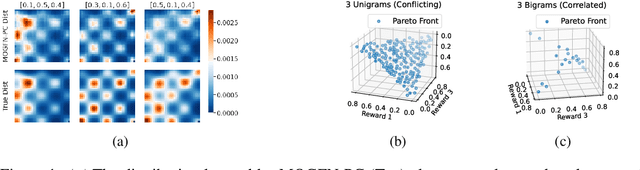


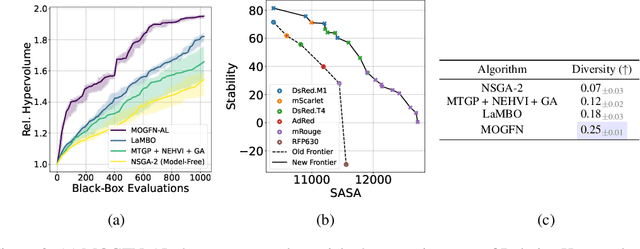
In many applications of machine learning, like drug discovery and material design, the goal is to generate candidates that simultaneously maximize a set of objectives. As these objectives are often conflicting, there is no single candidate that simultaneously maximizes all objectives, but rather a set of Pareto-optimal candidates where one objective cannot be improved without worsening another. Moreover, in practice, these objectives are often under-specified, making the diversity of candidates a key consideration. The existing multi-objective optimization methods focus predominantly on covering the Pareto front, failing to capture diversity in the space of candidates. Motivated by the success of GFlowNets for generation of diverse candidates in a single objective setting, in this paper we consider Multi-Objective GFlowNets (MOGFNs). MOGFNs consist of a novel Conditional GFlowNet which models a family of single-objective sub-problems derived by decomposing the multi-objective optimization problem. Our work is the first to empirically demonstrate conditional GFlowNets. Through a series of experiments on synthetic and benchmark tasks, we empirically demonstrate that MOGFNs outperform existing methods in terms of Hypervolume, R2-distance and candidate diversity. We also demonstrate the effectiveness of MOGFNs over existing methods in active learning settings. Finally, we supplement our empirical results with a careful analysis of each component of MOGFNs.
Learning GFlowNets from partial episodes for improved convergence and stability
Sep 30, 2022Kanika Madan, Jarrid Rector-Brooks, Maksym Korablyov, Emmanuel Bengio, Moksh Jain, Andrei Nica, Tom Bosc, Yoshua Bengio, Nikolay Malkin
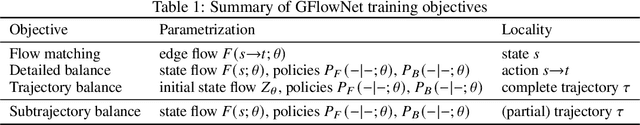

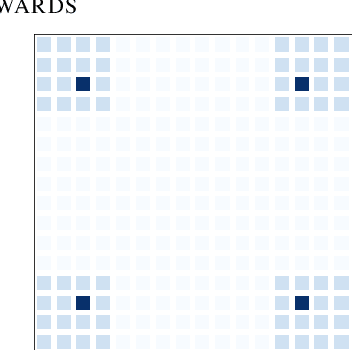

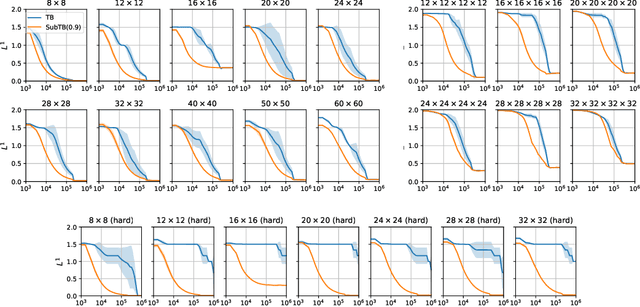
Generative flow networks (GFlowNets) are a family of algorithms for training a sequential sampler of discrete objects under an unnormalized target density and have been successfully used for various probabilistic modeling tasks. Existing training objectives for GFlowNets are either local to states or transitions, or propagate a reward signal over an entire sampling trajectory. We argue that these alternatives represent opposite ends of a gradient bias-variance tradeoff and propose a way to exploit this tradeoff to mitigate its harmful effects. Inspired by the TD($\lambda$) algorithm in reinforcement learning, we introduce subtrajectory balance or SubTB($\lambda$), a GFlowNet training objective that can learn from partial action subsequences of varying lengths. We show that SubTB($\lambda$) accelerates sampler convergence in previously studied and new environments and enables training GFlowNets in environments with longer action sequences and sparser reward landscapes than what was possible before. We also perform a comparative analysis of stochastic gradient dynamics, shedding light on the bias-variance tradeoff in GFlowNet training and the advantages of subtrajectory balance.
Biological Sequence Design with GFlowNets
Mar 02, 2022Moksh Jain, Emmanuel Bengio, Alex-Hernandez Garcia, Jarrid Rector-Brooks, Bonaventure F. P. Dossou, Chanakya Ekbote, Jie Fu, Tianyu Zhang, Micheal Kilgour, Dinghuai Zhang, Lena Simine, Payel Das, Yoshua Bengio
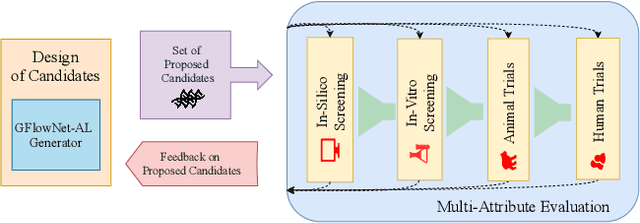
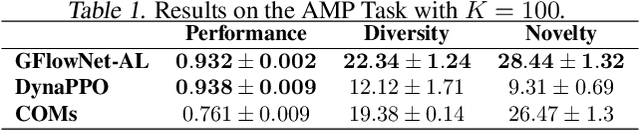
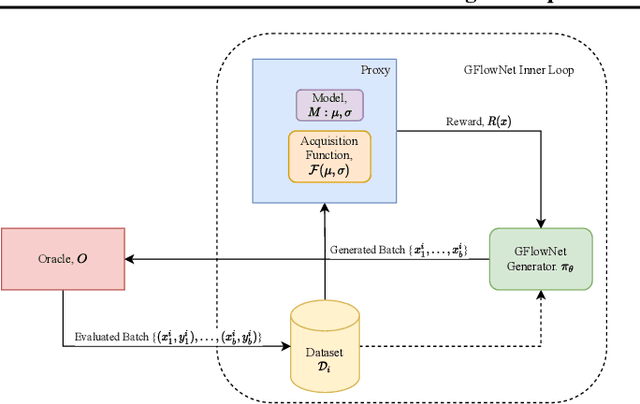
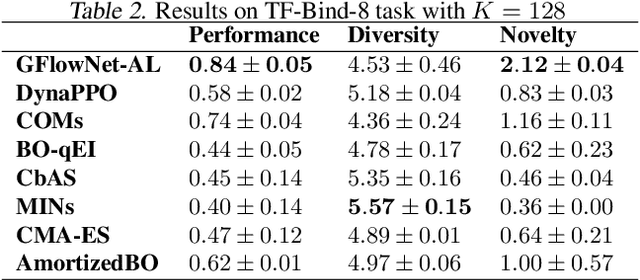
Design of de novo biological sequences with desired properties, like protein and DNA sequences, often involves an active loop with several rounds of molecule ideation and expensive wet-lab evaluations. These experiments can consist of multiple stages, with increasing levels of precision and cost of evaluation, where candidates are filtered. This makes the diversity of proposed candidates a key consideration in the ideation phase. In this work, we propose an active learning algorithm leveraging epistemic uncertainty estimation and the recently proposed GFlowNets as a generator of diverse candidate solutions, with the objective to obtain a diverse batch of useful (as defined by some utility function, for example, the predicted anti-microbial activity of a peptide) and informative candidates after each round. We also propose a scheme to incorporate existing labeled datasets of candidates, in addition to a reward function, to speed up learning in GFlowNets. We present empirical results on several biological sequence design tasks, and we find that our method generates more diverse and novel batches with high scoring candidates compared to existing approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge