Deep learning with noisy labels in medical prediction problems: a scoping review
Mar 19, 2024Yishu Wei, Yu Deng, Cong Sun, Mingquan Lin, Hongmei Jiang, Yifan Peng
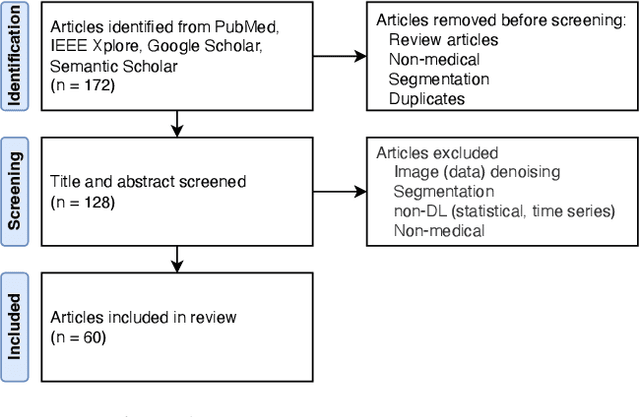
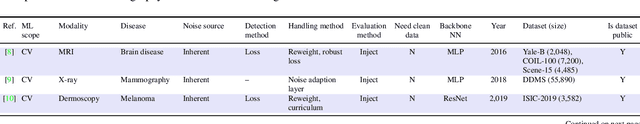
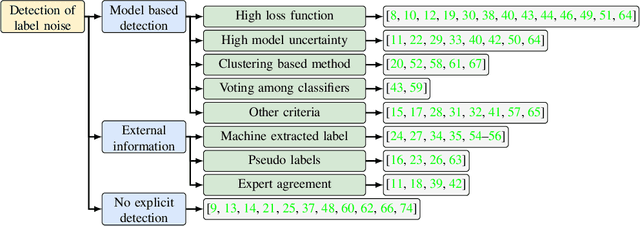
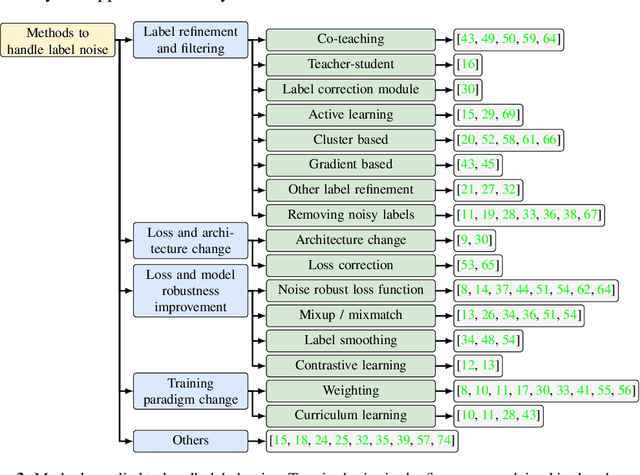
Objectives: Medical research faces substantial challenges from noisy labels attributed to factors like inter-expert variability and machine-extracted labels. Despite this, the adoption of label noise management remains limited, and label noise is largely ignored. To this end, there is a critical need to conduct a scoping review focusing on the problem space. This scoping review aims to comprehensively review label noise management in deep learning-based medical prediction problems, which includes label noise detection, label noise handling, and evaluation. Research involving label uncertainty is also included. Methods: Our scoping review follows the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. We searched 4 databases, including PubMed, IEEE Xplore, Google Scholar, and Semantic Scholar. Our search terms include "noisy label AND medical / healthcare / clinical", "un-certainty AND medical / healthcare / clinical", and "noise AND medical / healthcare / clinical". Results: A total of 60 papers met inclusion criteria between 2016 and 2023. A series of practical questions in medical research are investigated. These include the sources of label noise, the impact of label noise, the detection of label noise, label noise handling techniques, and their evaluation. Categorization of both label noise detection methods and handling techniques are provided. Discussion: From a methodological perspective, we observe that the medical community has been up to date with the broader deep-learning community, given that most techniques have been evaluated on medical data. We recommend considering label noise as a standard element in medical research, even if it is not dedicated to handling noisy labels. Initial experiments can start with easy-to-implement methods, such as noise-robust loss functions, weighting, and curriculum learning.
A survey of recent methods for addressing AI fairness and bias in biomedicine
Feb 13, 2024Yifan Yang, Mingquan Lin, Han Zhao, Yifan Peng, Furong Huang, Zhiyong Lu
Artificial intelligence (AI) systems have the potential to revolutionize clinical practices, including improving diagnostic accuracy and surgical decision-making, while also reducing costs and manpower. However, it is important to recognize that these systems may perpetuate social inequities or demonstrate biases, such as those based on race or gender. Such biases can occur before, during, or after the development of AI models, making it critical to understand and address potential biases to enable the accurate and reliable application of AI models in clinical settings. To mitigate bias concerns during model development, we surveyed recent publications on different debiasing methods in the fields of biomedical natural language processing (NLP) or computer vision (CV). Then we discussed the methods that have been applied in the biomedical domain to address bias. We performed our literature search on PubMed, ACM digital library, and IEEE Xplore of relevant articles published between January 2018 and December 2023 using multiple combinations of keywords. We then filtered the result of 10,041 articles automatically with loose constraints, and manually inspected the abstracts of the remaining 890 articles to identify the 55 articles included in this review. Additional articles in the references are also included in this review. We discuss each method and compare its strengths and weaknesses. Finally, we review other potential methods from the general domain that could be applied to biomedicine to address bias and improve fairness.The bias of AIs in biomedicine can originate from multiple sources. Existing debiasing methods that focus on algorithms can be categorized into distributional or algorithmic.
Improving Fairness of Automated Chest X-ray Diagnosis by Contrastive Learning
Jan 25, 2024Mingquan Lin, Tianhao Li, Zhaoyi Sun, Gregory Holste, Ying Ding, Fei Wang, George Shih, Yifan Peng
Purpose: Limited studies exploring concrete methods or approaches to tackle and enhance model fairness in the radiology domain. Our proposed AI model utilizes supervised contrastive learning to minimize bias in CXR diagnosis. Materials and Methods: In this retrospective study, we evaluated our proposed method on two datasets: the Medical Imaging and Data Resource Center (MIDRC) dataset with 77,887 CXR images from 27,796 patients collected as of April 20, 2023 for COVID-19 diagnosis, and the NIH Chest X-ray (NIH-CXR) dataset with 112,120 CXR images from 30,805 patients collected between 1992 and 2015. In the NIH-CXR dataset, thoracic abnormalities include atelectasis, cardiomegaly, effusion, infiltration, mass, nodule, pneumonia, pneumothorax, consolidation, edema, emphysema, fibrosis, pleural thickening, or hernia. Our proposed method utilizes supervised contrastive learning with carefully selected positive and negative samples to generate fair image embeddings, which are fine-tuned for subsequent tasks to reduce bias in chest X-ray (CXR) diagnosis. We evaluated the methods using the marginal AUC difference ($\delta$ mAUC). Results: The proposed model showed a significant decrease in bias across all subgroups when compared to the baseline models, as evidenced by a paired T-test (p<0.0001). The $\delta$ mAUC obtained by our method were 0.0116 (95\% CI, 0.0110-0.0123), 0.2102 (95% CI, 0.2087-0.2118), and 0.1000 (95\% CI, 0.0988-0.1011) for sex, race, and age on MIDRC, and 0.0090 (95\% CI, 0.0082-0.0097) for sex and 0.0512 (95% CI, 0.0512-0.0532) for age on NIH-CXR, respectively. Conclusion: Employing supervised contrastive learning can mitigate bias in CXR diagnosis, addressing concerns of fairness and reliability in deep learning-based diagnostic methods.
Towards long-tailed, multi-label disease classification from chest X-ray: Overview of the CXR-LT challenge
Oct 24, 2023Gregory Holste, Yiliang Zhou, Song Wang, Ajay Jaiswal, Mingquan Lin, Sherry Zhuge, Yuzhe Yang, Dongkyun Kim, Trong-Hieu Nguyen-Mau, Minh-Triet Tran, Jaehyup Jeong, Wongi Park, Jongbin Ryu, Feng Hong, Arsh Verma, Yosuke Yamagishi, Changhyun Kim, Hyeryeong Seo, Myungjoo Kang, Leo Anthony Celi, Zhiyong Lu, Ronald M. Summers, George Shih, Zhangyang Wang, Yifan Peng
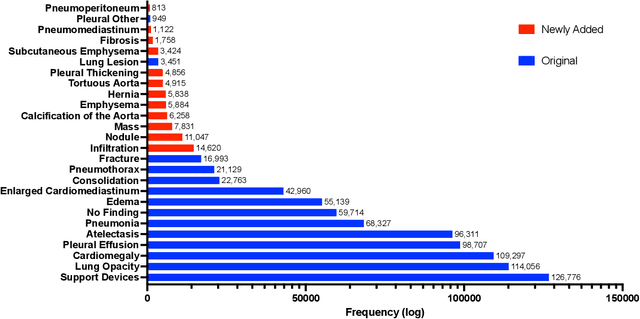
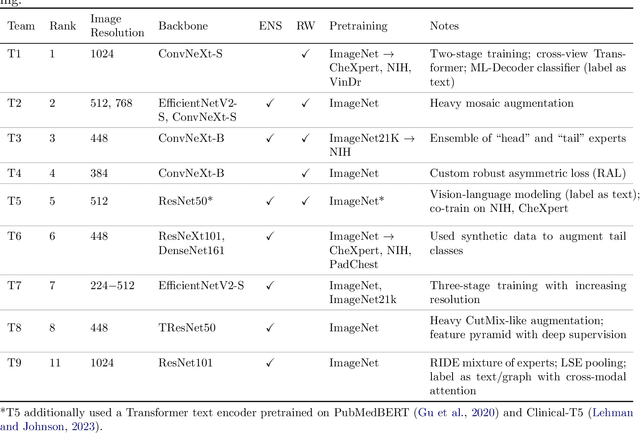
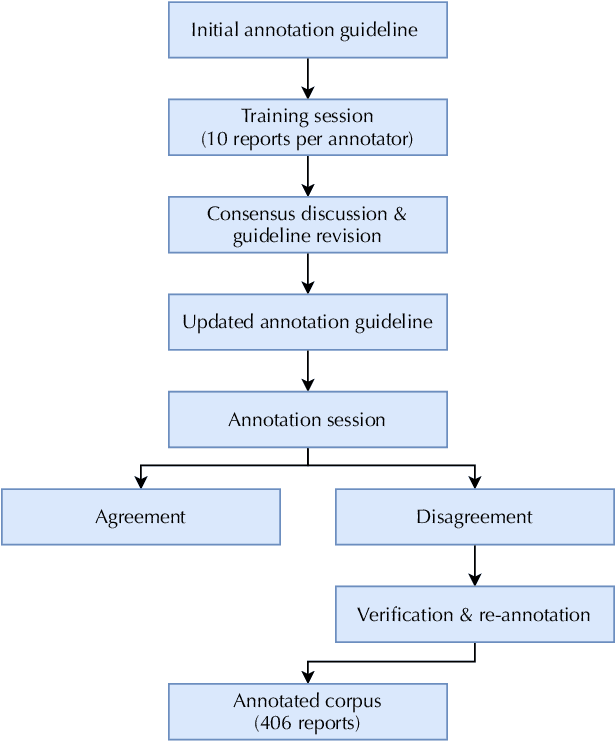
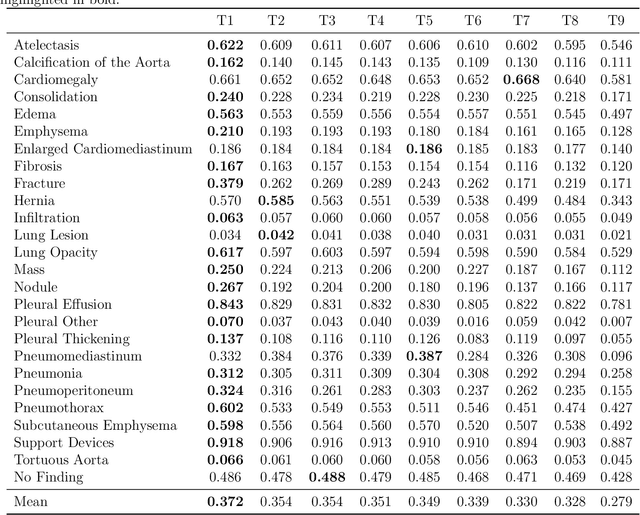
Many real-world image recognition problems, such as diagnostic medical imaging exams, are "long-tailed" $\unicode{x2013}$ there are a few common findings followed by many more relatively rare conditions. In chest radiography, diagnosis is both a long-tailed and multi-label problem, as patients often present with multiple findings simultaneously. While researchers have begun to study the problem of long-tailed learning in medical image recognition, few have studied the interaction of label imbalance and label co-occurrence posed by long-tailed, multi-label disease classification. To engage with the research community on this emerging topic, we conducted an open challenge, CXR-LT, on long-tailed, multi-label thorax disease classification from chest X-rays (CXRs). We publicly release a large-scale benchmark dataset of over 350,000 CXRs, each labeled with at least one of 26 clinical findings following a long-tailed distribution. We synthesize common themes of top-performing solutions, providing practical recommendations for long-tailed, multi-label medical image classification. Finally, we use these insights to propose a path forward involving vision-language foundation models for few- and zero-shot disease classification.
A scoping review on multimodal deep learning in biomedical images and texts
Jul 14, 2023Zhaoyi Sun, Mingquan Lin, Qingqing Zhu, Qianqian Xie, Fei Wang, Zhiyong Lu, Yifan Peng
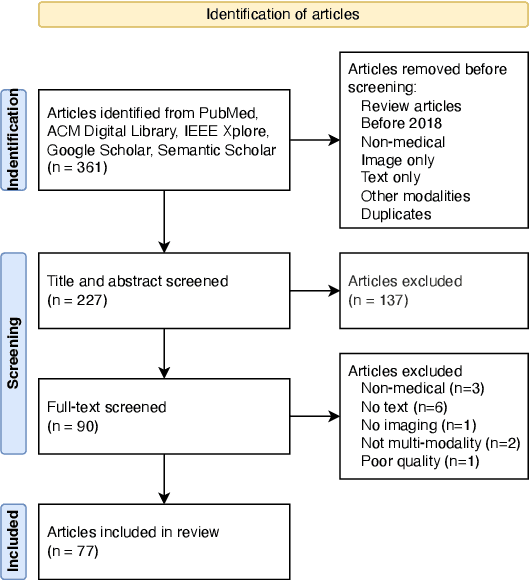
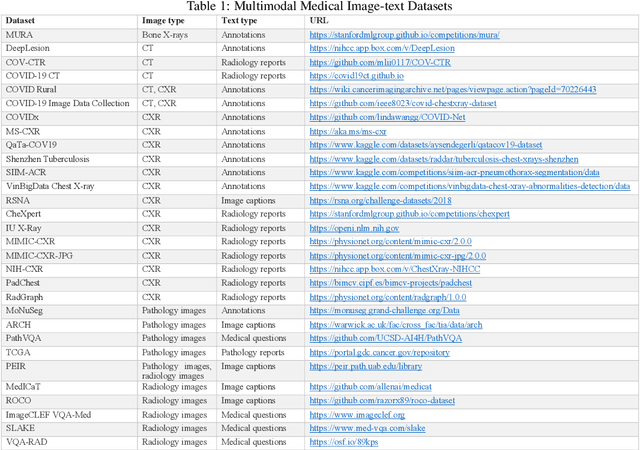
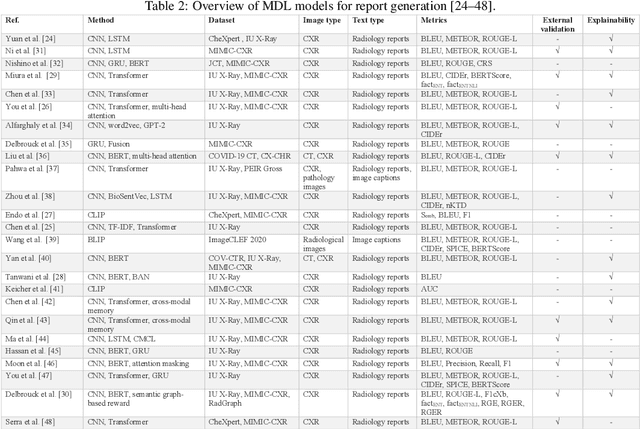
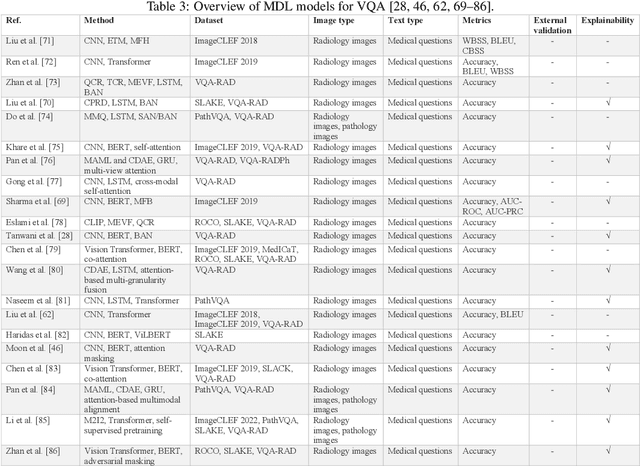
Computer-assisted diagnostic and prognostic systems of the future should be capable of simultaneously processing multimodal data. Multimodal deep learning (MDL), which involves the integration of multiple sources of data, such as images and text, has the potential to revolutionize the analysis and interpretation of biomedical data. However, it only caught researchers' attention recently. To this end, there is a critical need to conduct a systematic review on this topic, identify the limitations of current work, and explore future directions. In this scoping review, we aim to provide a comprehensive overview of the current state of the field and identify key concepts, types of studies, and research gaps with a focus on biomedical images and texts joint learning, mainly because these two were the most commonly available data types in MDL research. This study reviewed the current uses of multimodal deep learning on five tasks: (1) Report generation, (2) Visual question answering, (3) Cross-modal retrieval, (4) Computer-aided diagnosis, and (5) Semantic segmentation. Our results highlight the diverse applications and potential of MDL and suggest directions for future research in the field. We hope our review will facilitate the collaboration of natural language processing (NLP) and medical imaging communities and support the next generation of decision-making and computer-assisted diagnostic system development.
Evaluate underdiagnosis and overdiagnosis bias of deep learning model on primary open-angle glaucoma diagnosis in under-served patient populations
Jan 29, 2023Mingquan Lin, Yuyun Xiao, Bojian Hou, Tingyi Wanyan, Mohit Manoj Sharma, Zhangyang Wang, Fei Wang, Sarah Van Tassel, Yifan Peng
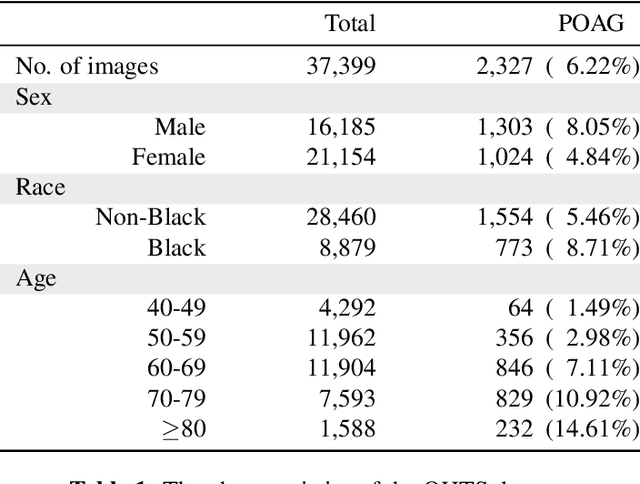
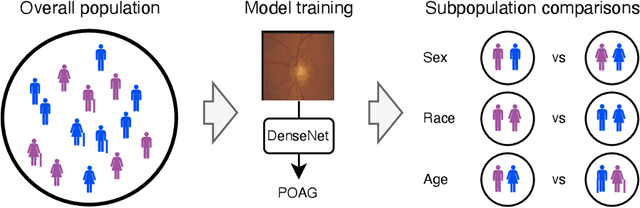
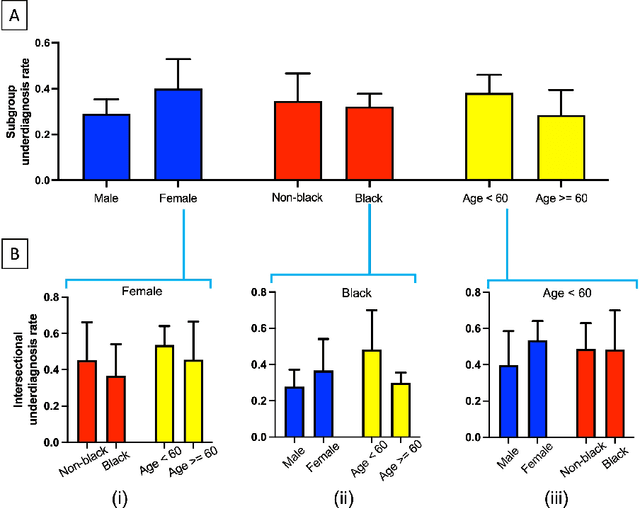
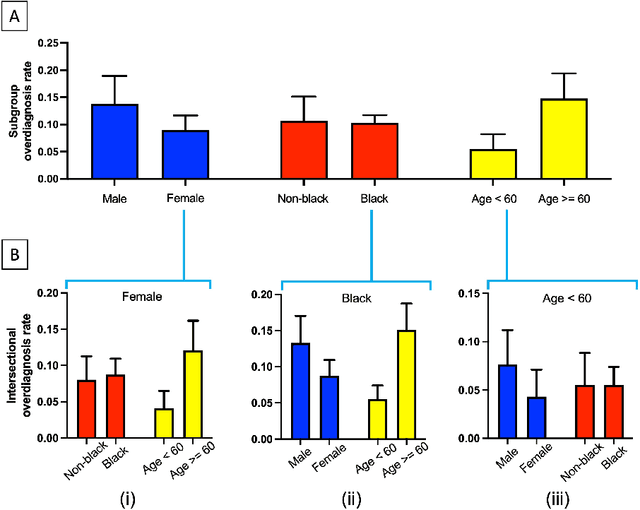
In the United States, primary open-angle glaucoma (POAG) is the leading cause of blindness, especially among African American and Hispanic individuals. Deep learning has been widely used to detect POAG using fundus images as its performance is comparable to or even surpasses diagnosis by clinicians. However, human bias in clinical diagnosis may be reflected and amplified in the widely-used deep learning models, thus impacting their performance. Biases may cause (1) underdiagnosis, increasing the risks of delayed or inadequate treatment, and (2) overdiagnosis, which may increase individuals' stress, fear, well-being, and unnecessary/costly treatment. In this study, we examined the underdiagnosis and overdiagnosis when applying deep learning in POAG detection based on the Ocular Hypertension Treatment Study (OHTS) from 22 centers across 16 states in the United States. Our results show that the widely-used deep learning model can underdiagnose or overdiagnose underserved populations. The most underdiagnosed group is female younger (< 60 yrs) group, and the most overdiagnosed group is Black older (>=60 yrs) group. Biased diagnosis through traditional deep learning methods may delay disease detection, treatment and create burdens among under-served populations, thereby, raising ethical concerns about using deep learning models in ophthalmology clinics.
* 9 pages, 2 figures, Accepted by AMIA 2023 Informatics Summit
Prior Knowledge Enhances Radiology Report Generation
Jan 11, 2022Song Wang, Liyan Tang, Mingquan Lin, George Shih, Ying Ding, Yifan Peng
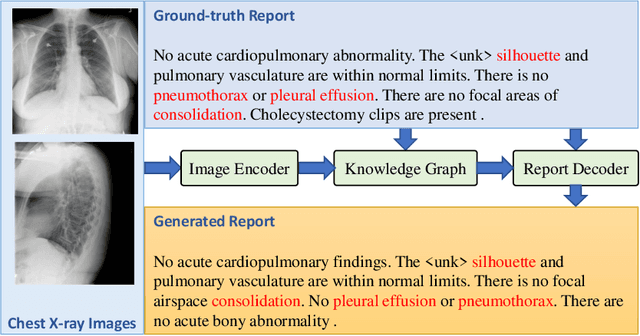

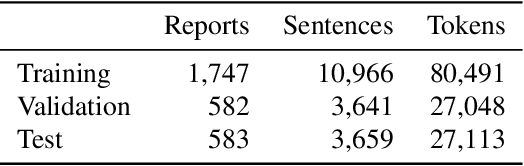
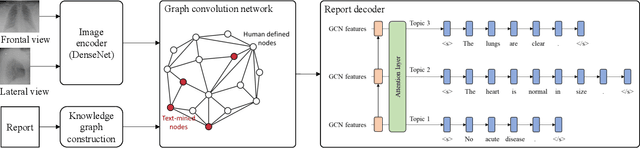
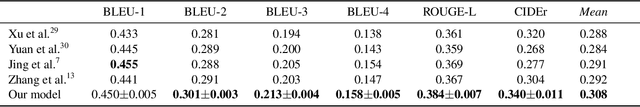
Radiology report generation aims to produce computer-aided diagnoses to alleviate the workload of radiologists and has drawn increasing attention recently. However, previous deep learning methods tend to neglect the mutual influences between medical findings, which can be the bottleneck that limits the quality of generated reports. In this work, we propose to mine and represent the associations among medical findings in an informative knowledge graph and incorporate this prior knowledge with radiology report generation to help improve the quality of generated reports. Experiment results demonstrate the superior performance of our proposed method on the IU X-ray dataset with a ROUGE-L of 0.384$\pm$0.007 and CIDEr of 0.340$\pm$0.011. Compared with previous works, our model achieves an average of 1.6% improvement (2.0% and 1.5% improvements in CIDEr and ROUGE-L, respectively). The experiments suggest that prior knowledge can bring performance gains to accurate radiology report generation. We will make the code publicly available at https://github.com/bionlplab/report_generation_amia2022.
Artificial Intelligence in Tumor Subregion Analysis Based on Medical Imaging: A Review
Mar 25, 2021Mingquan Lin, Jacob Wynne, Yang Lei, Tonghe Wang, Walter J. Curran, Tian Liu, Xiaofeng Yang
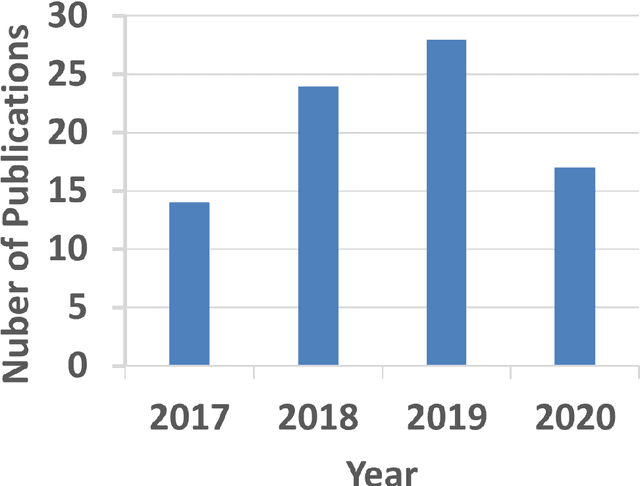
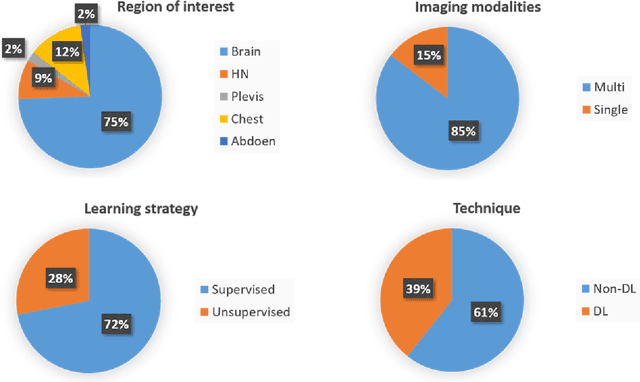

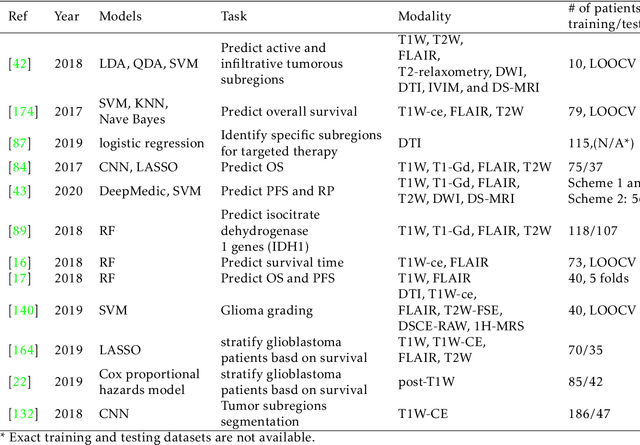
Medical imaging is widely used in cancer diagnosis and treatment, and artificial intelligence (AI) has achieved tremendous success in various tasks of medical image analysis. This paper reviews AI-based tumor subregion analysis in medical imaging. We summarize the latest AI-based methods for tumor subregion analysis and their applications. Specifically, we categorize the AI-based methods by training strategy: supervised and unsupervised. A detailed review of each category is presented, highlighting important contributions and achievements. Specific challenges and potential AI applications in tumor subregion analysis are discussed.
Using Radiomics as Prior Knowledge for Abnormality Classification and Localization in Chest X-rays
Nov 25, 2020Yan Han, Chongyan Chen, Liyan Tang, Mingquan Lin, Ajay Jaiswal, Ying Ding, Yifan Peng
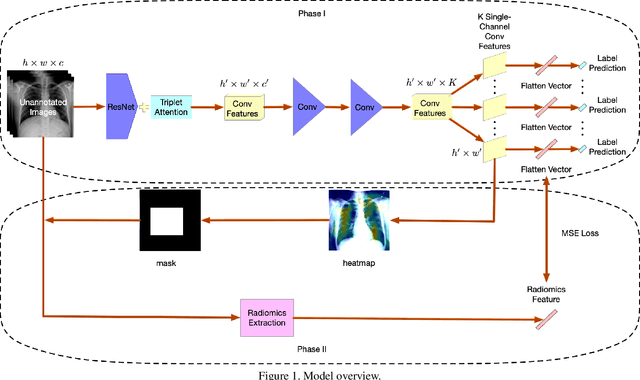
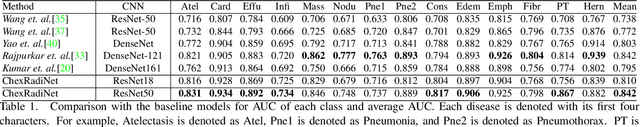
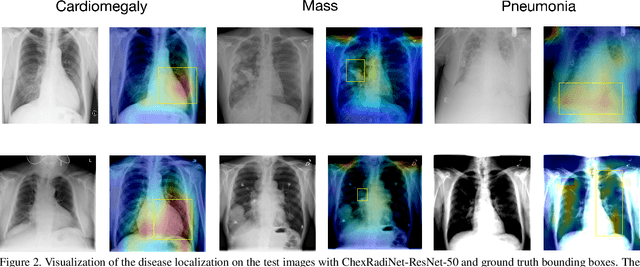
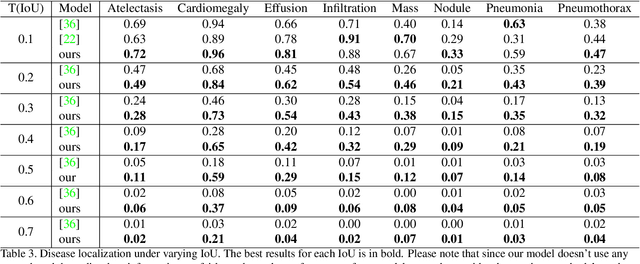
Chest X-rays become one of the most common medical diagnoses due to its noninvasiveness. The number of chest X-ray images has skyrocketed, but reading chest X-rays still have been manually performed by radiologists, which creates huge burnouts and delays. Traditionally, radiomics, as a subfield of radiology that can extract a large number of quantitative features from medical images, demonstrates its potential to facilitate medical imaging diagnosis before the deep learning era. In this paper, we develop an algorithm that can utilize the radiomics features to improve the abnormality classification performance. Our algorithm, ChexRadiNet, applies a light-weight but efficient triplet-attention mechanism for highlighting the meaningful image regions to improve localization accuracy. We first apply ChexRadiNet to classify the chest X-rays by using only image features. Then we use the generated heatmaps of chest X-rays to extract radiomics features. Finally, the extracted radiomics features could be used to guide our model to learn more robust accurate image features. After a number of iterations, our model could focus on more accurate image regions and extract more robust features. The empirical evaluation of our method supports our intuition and outperforms other state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge