Simple, Efficient and Scalable Structure-aware Adapter Boosts Protein Language Models
Apr 23, 2024Yang Tan, Mingchen Li, Bingxin Zhou, Bozitao Zhong, Lirong Zheng, Pan Tan, Ziyi Zhou, Huiqun Yu, Guisheng Fan, Liang Hong
Fine-tuning Pre-trained protein language models (PLMs) has emerged as a prominent strategy for enhancing downstream prediction tasks, often outperforming traditional supervised learning approaches. As a widely applied powerful technique in natural language processing, employing Parameter-Efficient Fine-Tuning techniques could potentially enhance the performance of PLMs. However, the direct transfer to life science tasks is non-trivial due to the different training strategies and data forms. To address this gap, we introduce SES-Adapter, a simple, efficient, and scalable adapter method for enhancing the representation learning of PLMs. SES-Adapter incorporates PLM embeddings with structural sequence embeddings to create structure-aware representations. We show that the proposed method is compatible with different PLM architectures and across diverse tasks. Extensive evaluations are conducted on 2 types of folding structures with notable quality differences, 9 state-of-the-art baselines, and 9 benchmark datasets across distinct downstream tasks. Results show that compared to vanilla PLMs, SES-Adapter improves downstream task performance by a maximum of 11% and an average of 3%, with significantly accelerated training speed by a maximum of 1034% and an average of 362%, the convergence rate is also improved by approximately 2 times. Moreover, positive optimization is observed even with low-quality predicted structures. The source code for SES-Adapter is available at https://github.com/tyang816/SES-Adapter.
PETA: Evaluating the Impact of Protein Transfer Learning with Sub-word Tokenization on Downstream Applications
Oct 26, 2023Yang Tan, Mingchen Li, Pan Tan, Ziyi Zhou, Huiqun Yu, Guisheng Fan, Liang Hong
Large protein language models are adept at capturing the underlying evolutionary information in primary structures, offering significant practical value for protein engineering. Compared to natural language models, protein amino acid sequences have a smaller data volume and a limited combinatorial space. Choosing an appropriate vocabulary size to optimize the pre-trained model is a pivotal issue. Moreover, despite the wealth of benchmarks and studies in the natural language community, there remains a lack of a comprehensive benchmark for systematically evaluating protein language model quality. Given these challenges, PETA trained language models with 14 different vocabulary sizes under three tokenization methods. It conducted thousands of tests on 33 diverse downstream datasets to assess the models' transfer learning capabilities, incorporating two classification heads and three random seeds to mitigate potential biases. Extensive experiments indicate that vocabulary sizes between 50 and 200 optimize the model, whereas sizes exceeding 800 detrimentally affect the model's representational performance. Our code, model weights and datasets are available at https://github.com/ginnm/ProteinPretraining.
MedChatZH: a Better Medical Adviser Learns from Better Instructions
Sep 03, 2023Yang Tan, Mingchen Li, Zijie Huang, Huiqun Yu, Guisheng Fan
Generative large language models (LLMs) have shown great success in various applications, including question-answering (QA) and dialogue systems. However, in specialized domains like traditional Chinese medical QA, these models may perform unsatisfactorily without fine-tuning on domain-specific datasets. To address this, we introduce MedChatZH, a dialogue model designed specifically for traditional Chinese medical QA. Our model is pre-trained on Chinese traditional medical books and fine-tuned with a carefully curated medical instruction dataset. It outperforms several solid baselines on a real-world medical dialogue dataset. We release our model, code, and dataset on https://github.com/tyang816/MedChatZH to facilitate further research in the domain of traditional Chinese medicine and LLMs.
Multi-level Protein Representation Learning for Blind Mutational Effect Prediction
Jun 08, 2023Yang Tan, Bingxin Zhou, Yuanhong Jiang, Yu Guang Wang, Liang Hong
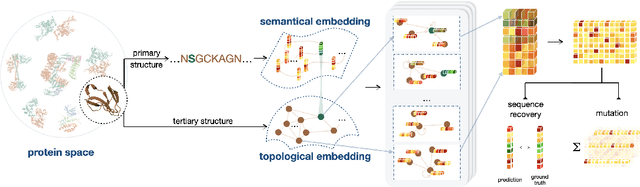

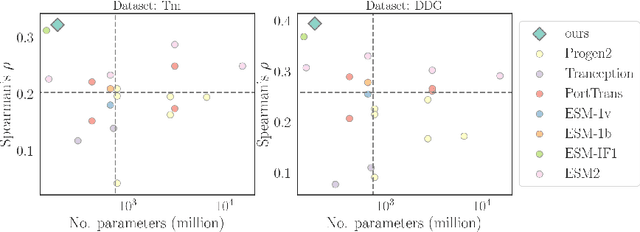
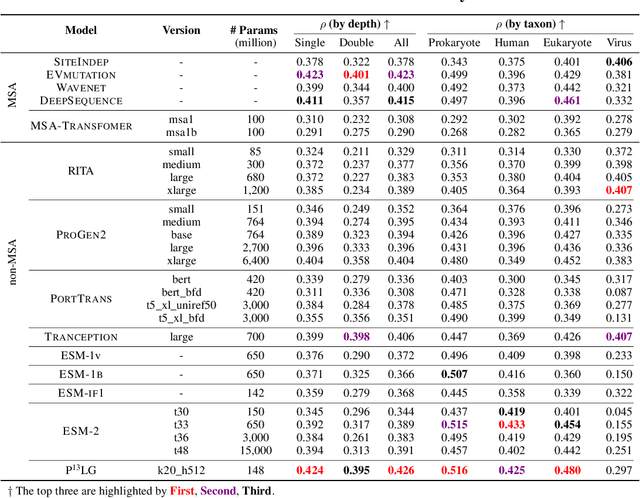
Directed evolution plays an indispensable role in protein engineering that revises existing protein sequences to attain new or enhanced functions. Accurately predicting the effects of protein variants necessitates an in-depth understanding of protein structure and function. Although large self-supervised language models have demonstrated remarkable performance in zero-shot inference using only protein sequences, these models inherently do not interpret the spatial characteristics of protein structures, which are crucial for comprehending protein folding stability and internal molecular interactions. This paper introduces a novel pre-training framework that cascades sequential and geometric analyzers for protein primary and tertiary structures. It guides mutational directions toward desired traits by simulating natural selection on wild-type proteins and evaluates the effects of variants based on their fitness to perform the function. We assess the proposed approach using a public database and two new databases for a variety of variant effect prediction tasks, which encompass a diverse set of proteins and assays from different taxa. The prediction results achieve state-of-the-art performance over other zero-shot learning methods for both single-site mutations and deep mutations.
A Comprehensive Survey on Heart Sound Analysis in the Deep Learning Era
Jan 23, 2023Zhao Ren, Yi Chang, Thanh Tam Nguyen, Yang Tan, Kun Qian, Björn W. Schuller


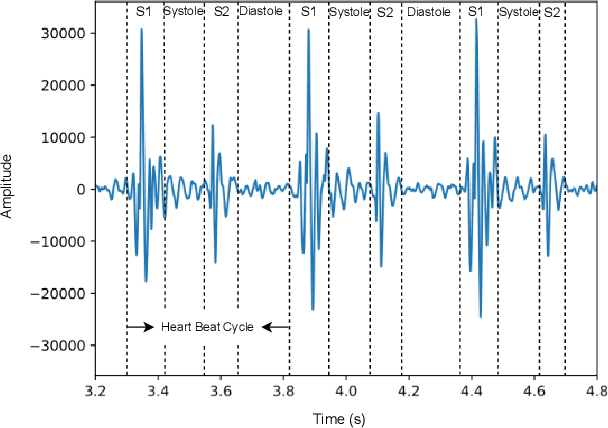
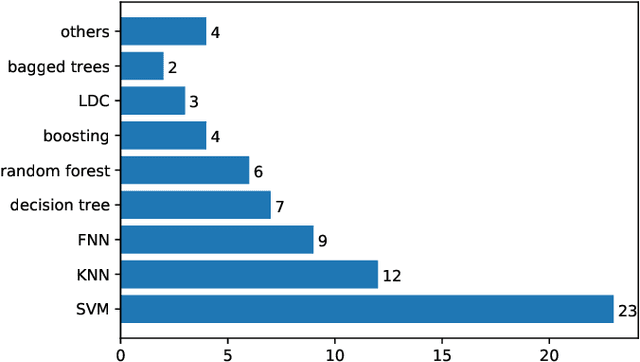
Heart sound auscultation has been demonstrated to be beneficial in clinical usage for early screening of cardiovascular diseases. Due to the high requirement of well-trained professionals for auscultation, automatic auscultation benefiting from signal processing and machine learning can help auxiliary diagnosis and reduce the burdens of training professional clinicians. Nevertheless, classic machine learning is limited to performance improvement in the era of big data. Deep learning has achieved better performance than classic machine learning in many research fields, as it employs more complex model architectures with stronger capability of extracting effective representations. Deep learning has been successfully applied to heart sound analysis in the past years. As most review works about heart sound analysis were given before 2017, the present survey is the first to work on a comprehensive overview to summarise papers on heart sound analysis with deep learning in the past six years 2017--2022. We introduce both classic machine learning and deep learning for comparison, and further offer insights about the advances and future research directions in deep learning for heart sound analysis.
Finding the Most Transferable Tasks for Brain Image Segmentation
Jan 03, 2023Yicong Li, Yang Tan, Jingyun Yang, Yang Li, Xiao-Ping Zhang
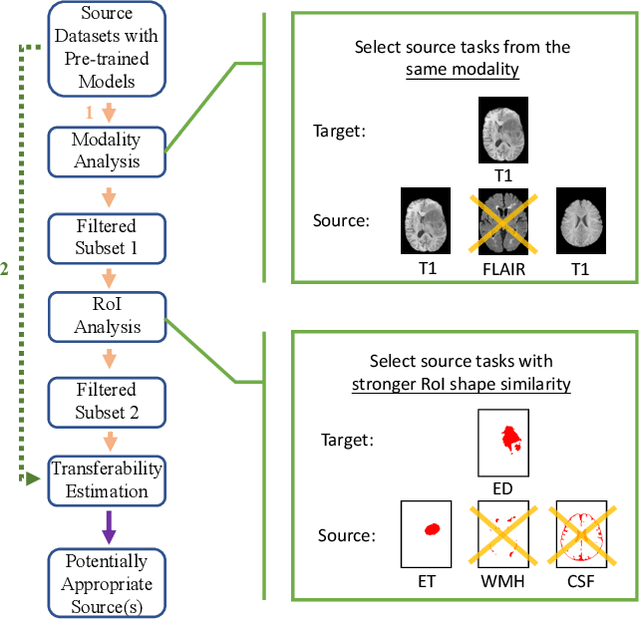
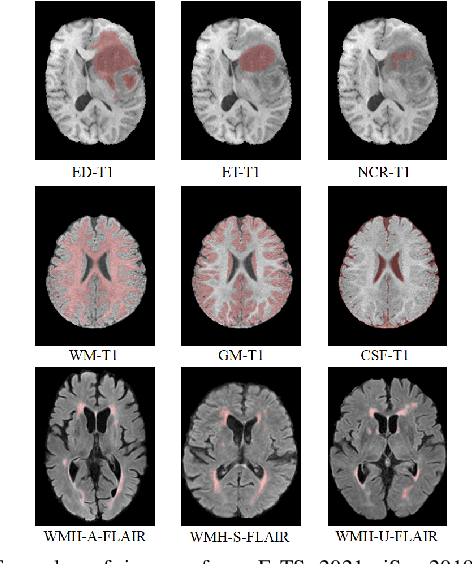


Although many studies have successfully applied transfer learning to medical image segmentation, very few of them have investigated the selection strategy when multiple source tasks are available for transfer. In this paper, we propose a prior knowledge guided and transferability based framework to select the best source tasks among a collection of brain image segmentation tasks, to improve the transfer learning performance on the given target task. The framework consists of modality analysis, RoI (region of interest) analysis, and transferability estimation, such that the source task selection can be refined step by step. Specifically, we adapt the state-of-the-art analytical transferability estimation metrics to medical image segmentation tasks and further show that their performance can be significantly boosted by filtering candidate source tasks based on modality and RoI characteristics. Our experiments on brain matter, brain tumor, and white matter hyperintensities segmentation datasets reveal that transferring from different tasks under the same modality is often more successful than transferring from the same task under different modalities. Furthermore, within the same modality, transferring from the source task that has stronger RoI shape similarity with the target task can significantly improve the final transfer performance. And such similarity can be captured using the Structural Similarity index in the label space.
Transferability-Guided Cross-Domain Cross-Task Transfer Learning
Jul 12, 2022Yang Tan, Yang Li, Shao-Lun Huang, Xiao-Ping Zhang
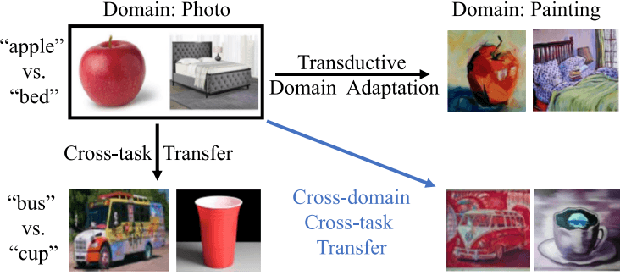
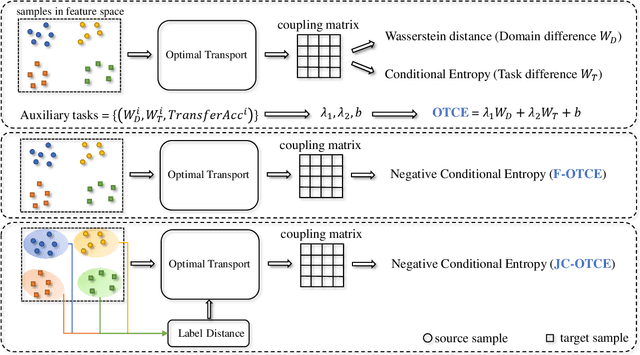
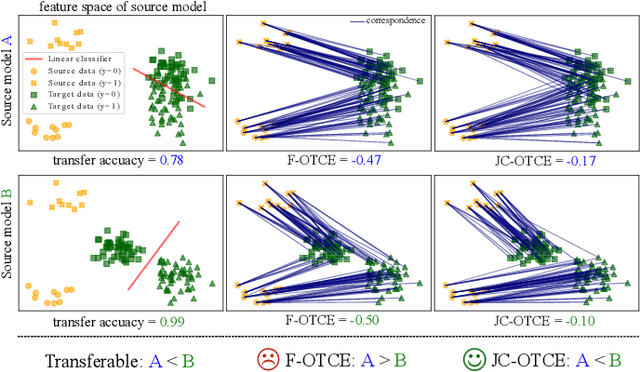
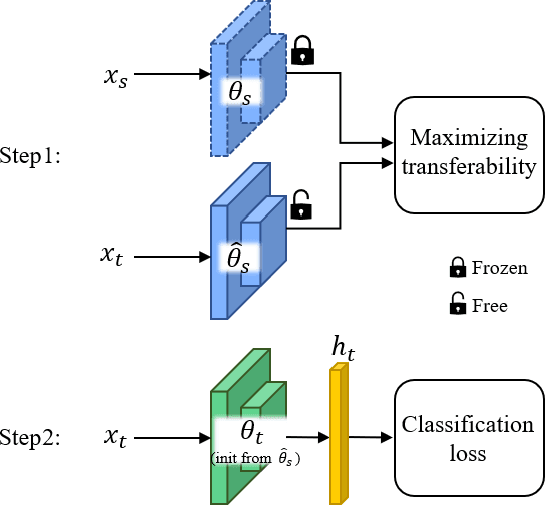
We propose two novel transferability metrics F-OTCE (Fast Optimal Transport based Conditional Entropy) and JC-OTCE (Joint Correspondence OTCE) to evaluate how much the source model (task) can benefit the learning of the target task and to learn more transferable representations for cross-domain cross-task transfer learning. Unlike the existing metric that requires evaluating the empirical transferability on auxiliary tasks, our metrics are auxiliary-free such that they can be computed much more efficiently. Specifically, F-OTCE estimates transferability by first solving an Optimal Transport (OT) problem between source and target distributions, and then uses the optimal coupling to compute the Negative Conditional Entropy between source and target labels. It can also serve as a loss function to maximize the transferability of the source model before finetuning on the target task. Meanwhile, JC-OTCE improves the transferability robustness of F-OTCE by including label distances in the OT problem, though it may incur additional computation cost. Extensive experiments demonstrate that F-OTCE and JC-OTCE outperform state-of-the-art auxiliary-free metrics by 18.85% and 28.88%, respectively in correlation coefficient with the ground-truth transfer accuracy. By eliminating the training cost of auxiliary tasks, the two metrics reduces the total computation time of the previous method from 43 minutes to 9.32s and 10.78s, respectively, for a pair of tasks. When used as a loss function, F-OTCE shows consistent improvements on the transfer accuracy of the source model in few-shot classification experiments, with up to 4.41% accuracy gain.
Transferability Estimation for Semantic Segmentation Task
Oct 11, 2021Yang Tan, Yang Li, Shao-Lun Huang


Transferability estimation is a fundamental problem in transfer learning to predict how good the performance is when transferring a source model (or source task) to a target task. With the guidance of transferability score, we can efficiently select the highly transferable source models without performing the real transfer in practice. Recent analytical transferability metrics are mainly designed for image classification problem, and currently there is no specific investigation for the transferability estimation of semantic segmentation task, which is an essential problem in autonomous driving, medical image analysis, etc. Consequently, we further extend the recent analytical transferability metric OTCE (Optimal Transport based Conditional Entropy) score to the semantic segmentation task. The challenge in applying the OTCE score is the high dimensional segmentation output, which is difficult to find the optimal coupling between so many pixels under an acceptable computation cost. Thus we propose to randomly sample N pixels for computing OTCE score and take the expectation over K repetitions as the final transferability score. Experimental evaluation on Cityscapes, BDD100K and GTA5 datasets demonstrates that the OTCE score highly correlates with the transfer performance.
Practical Transferability Estimation for Image Classification Tasks
Jun 30, 2021Yang Tan, Yang Li, Shao-Lun Huang




Transferability estimation is an essential problem in transfer learning to predict how good the performance is when transferring a source model (or source task) to a target task. Recent analytical transferability metrics have been widely used for source model selection and multi-task learning. A major challenge is how to make transfereability estimation robust under the cross-domain cross-task settings. The recently proposed OTCE score solves this problem by considering both domain and task differences, with the help of transfer experiences on auxiliary tasks, which causes an efficiency overhead. In this work, we propose a practical transferability metric called JC-NCE score that dramatically improves the robustness of the task difference estimation in OTCE, thus removing the need for auxiliary tasks. Specifically, we build the joint correspondences between source and target data via solving an optimal transport problem with a ground cost considering both the sample distance and label distance, and then compute the transferability score as the negative conditional entropy of the matched labels. Extensive validations under the intra-dataset and inter-dataset transfer settings demonstrate that our JC-NCE score outperforms the auxiliary-task free version of OTCE for 7% and 12%, respectively, and is also more robust than other existing transferability metrics on average.
OTCE: A Transferability Metric for Cross-Domain Cross-Task Representations
Mar 25, 2021Yang Tan, Yang Li, Shao-Lun Huang




Transfer learning across heterogeneous data distributions (a.k.a. domains) and distinct tasks is a more general and challenging problem than conventional transfer learning, where either domains or tasks are assumed to be the same. While neural network based feature transfer is widely used in transfer learning applications, finding the optimal transfer strategy still requires time-consuming experiments and domain knowledge. We propose a transferability metric called Optimal Transport based Conditional Entropy (OTCE), to analytically predict the transfer performance for supervised classification tasks in such cross-domain and cross-task feature transfer settings. Our OTCE score characterizes transferability as a combination of domain difference and task difference, and explicitly evaluates them from data in a unified framework. Specifically, we use optimal transport to estimate domain difference and the optimal coupling between source and target distributions, which is then used to derive the conditional entropy of the target task (task difference). Experiments on the largest cross-domain dataset DomainNet and Office31 demonstrate that OTCE shows an average of 21% gain in the correlation with the ground truth transfer accuracy compared to state-of-the-art methods. We also investigate two applications of the OTCE score including source model selection and multi-source feature fusion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge