DAWN: Domain-Adaptive Weakly Supervised Nuclei Segmentation via Cross-Task Interactions
Apr 24, 2024Ye Zhang, Yifeng Wang, Zijie Fang, Hao Bian, Linghan Cai, Ziyue Wang, Yongbing Zhang
Weakly supervised segmentation methods have gained significant attention due to their ability to reduce the reliance on costly pixel-level annotations during model training. However, the current weakly supervised nuclei segmentation approaches typically follow a two-stage pseudo-label generation and network training process. The performance of the nuclei segmentation heavily relies on the quality of the generated pseudo-labels, thereby limiting its effectiveness. This paper introduces a novel domain-adaptive weakly supervised nuclei segmentation framework using cross-task interaction strategies to overcome the challenge of pseudo-label generation. Specifically, we utilize weakly annotated data to train an auxiliary detection task, which assists the domain adaptation of the segmentation network. To enhance the efficiency of domain adaptation, we design a consistent feature constraint module integrating prior knowledge from the source domain. Furthermore, we develop pseudo-label optimization and interactive training methods to improve the domain transfer capability. To validate the effectiveness of our proposed method, we conduct extensive comparative and ablation experiments on six datasets. The results demonstrate the superiority of our approach over existing weakly supervised approaches. Remarkably, our method achieves comparable or even better performance than fully supervised methods. Our code will be released in https://github.com/zhangye-zoe/DAWN.
MamMIL: Multiple Instance Learning for Whole Slide Images with State Space Models
Mar 08, 2024Zijie Fang, Yifeng Wang, Zhi Wang, Jian Zhang, Xiangyang Ji, Yongbing Zhang
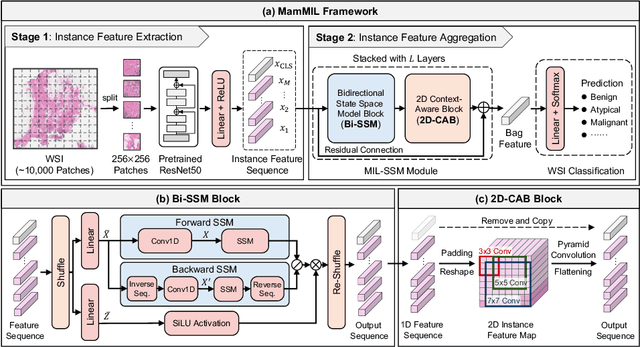
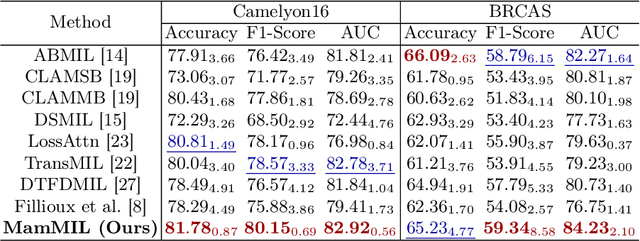
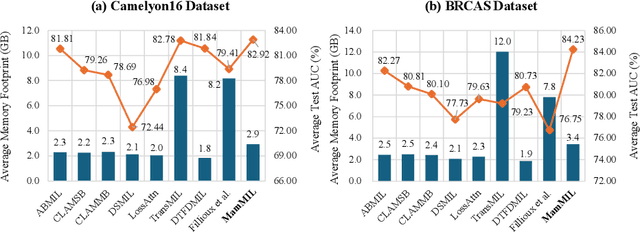
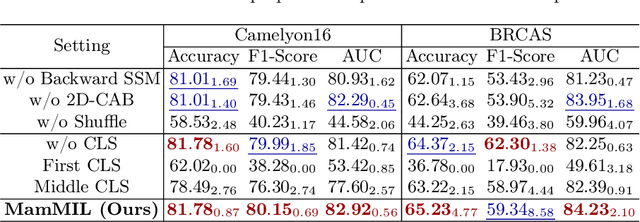
Recently, pathological diagnosis, the gold standard for cancer diagnosis, has achieved superior performance by combining the Transformer with the multiple instance learning (MIL) framework using whole slide images (WSIs). However, the giga-pixel nature of WSIs poses a great challenge for the quadratic-complexity self-attention mechanism in Transformer to be applied in MIL. Existing studies usually use linear attention to improve computing efficiency but inevitably bring performance bottlenecks. To tackle this challenge, we propose a MamMIL framework for WSI classification by cooperating the selective structured state space model (i.e., Mamba) with MIL for the first time, enabling the modeling of instance dependencies while maintaining linear complexity. Specifically, to solve the problem that Mamba can only conduct unidirectional one-dimensional (1D) sequence modeling, we innovatively introduce a bidirectional state space model and a 2D context-aware block to enable MamMIL to learn the bidirectional instance dependencies with 2D spatial relationships. Experiments on two datasets show that MamMIL can achieve advanced classification performance with smaller memory footprints than the state-of-the-art MIL frameworks based on the Transformer. The code will be open-sourced if accepted.
Boundary-aware Contrastive Learning for Semi-supervised Nuclei Instance Segmentation
Feb 07, 2024Ye Zhang, Ziyue Wang, Yifeng Wang, Hao Bian, Linghan Cai, Hengrui Li, Lingbo Zhang, Yongbing Zhang
Semi-supervised segmentation methods have demonstrated promising results in natural scenarios, providing a solution to reduce dependency on manual annotation. However, these methods face significant challenges when directly applied to pathological images due to the subtle color differences between nuclei and tissues, as well as the significant morphological variations among nuclei. Consequently, the generated pseudo-labels often contain much noise, especially at the nuclei boundaries. To address the above problem, this paper proposes a boundary-aware contrastive learning network to denoise the boundary noise in a semi-supervised nuclei segmentation task. The model has two key designs: a low-resolution denoising (LRD) module and a cross-RoI contrastive learning (CRC) module. The LRD improves the smoothness of the nuclei boundary by pseudo-labels denoising, and the CRC enhances the discrimination between foreground and background by boundary feature contrastive learning. We conduct extensive experiments to demonstrate the superiority of our proposed method over existing semi-supervised instance segmentation methods.
MedTransformer: Accurate AD Diagnosis for 3D MRI Images through 2D Vision Transformers
Jan 12, 2024Yifeng Wang, Ke Chen, Yihan Zhang, Haohan Wang
Automated diagnosis of AD in brain images is becoming a clinically important technique to support precision and efficient diagnosis and treatment planning. A few efforts have been made to automatically diagnose AD in magnetic resonance imaging (MRI) using three-dimensional CNNs. However, due to the complexity of 3D models, the performance is still unsatisfactory, both in terms of accuracy and efficiency. To overcome the complexities of 3D images and 3D models, in this study, we aim to attack this problem with 2D vision Transformers. We propose a 2D transformer-based medical image model with various transformer attention encoders to diagnose AD in 3D MRI images, by cutting the 3D images into multiple 2D slices.The model consists of four main components: shared encoders across three dimensions, dimension-specific encoders, attention across images from the same dimension, and attention across three dimensions. It is used to obtain attention relationships among multiple sequences from different dimensions (axial, coronal, and sagittal) and multiple slices. We also propose morphology augmentation, an erosion and dilation based method to increase the structural difference between AD and normal images. In this experiment, we use multiple datasets from ADNI, AIBL, MIRAID, OASIS to show the performance of our model. Our proposed MedTransformer demonstrates a strong ability in diagnosing AD. These results demonstrate the effectiveness of MedTransformer in learning from 3D data using a much smaller model and its capability to generalize among different medical tasks, which provides a possibility to help doctors diagnose AD in a simpler way.
Wavelet Dynamic Selection Network for Inertial Sensor Signal Enhancement
Dec 29, 2023Yifeng Wang, Yi Zhao
As attitude and motion sensing components, inertial sensors are widely used in various portable devices. But the severe errors of inertial sensors restrain their function, especially the trajectory recovery and semantic recognition. As a mainstream signal processing method, wavelet is hailed as the mathematical microscope of signal due to the plentiful and diverse wavelet basis functions. However, complicated noise types and application scenarios of inertial sensors make selecting wavelet basis perplexing. To this end, we propose a wavelet dynamic selection network (WDSNet), which intelligently selects the appropriate wavelet basis for variable inertial signals. In addition, existing deep learning architectures excel at extracting features from input data but neglect to learn the characteristics of target categories, which is essential to enhance the category awareness capability, thereby improving the selection of wavelet basis. Therefore, we propose a category representation mechanism (CRM), which enables the network to extract and represent category features without increasing trainable parameters. Furthermore, CRM transforms the common fully connected network into category representations, which provide closer supervision to the feature extractor than the far and trivial one-hot classification labels. We call this process of imposing interpretability on a network and using it to supervise the feature extractor the feature supervision mechanism, and its effectiveness is demonstrated experimentally and theoretically in this paper. The enhanced inertial signal can perform impracticable tasks with regard to the original signal, such as trajectory reconstruction. Both quantitative and visual results show that WDSNet outperforms the existing methods. Remarkably, WDSNet, as a weakly-supervised method, achieves the state-of-the-art performance of all the compared fully-supervised methods.
Deep Learning Predicts Biomarker Status and Discovers Related Histomorphology Characteristics for Low-Grade Glioma
Oct 11, 2023Zijie Fang, Yihan Liu, Yifeng Wang, Xiangyang Zhang, Yang Chen, Changjing Cai, Yiyang Lin, Ying Han, Zhi Wang, Shan Zeng, Hong Shen, Jun Tan, Yongbing Zhang

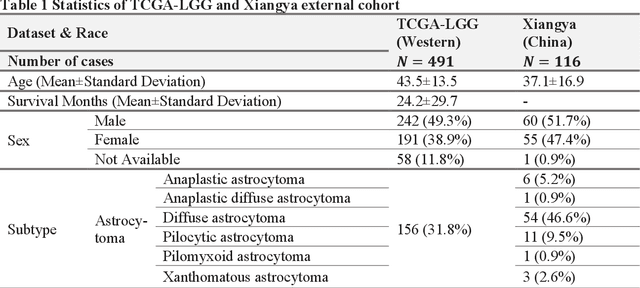

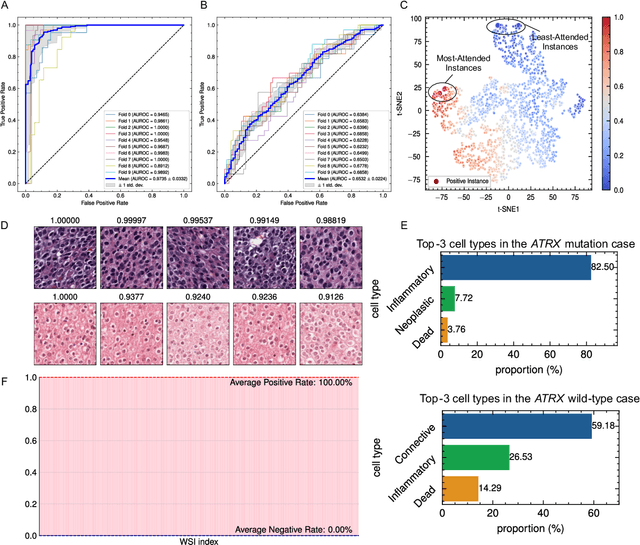
Biomarker detection is an indispensable part in the diagnosis and treatment of low-grade glioma (LGG). However, current LGG biomarker detection methods rely on expensive and complex molecular genetic testing, for which professionals are required to analyze the results, and intra-rater variability is often reported. To overcome these challenges, we propose an interpretable deep learning pipeline, a Multi-Biomarker Histomorphology Discoverer (Multi-Beholder) model based on the multiple instance learning (MIL) framework, to predict the status of five biomarkers in LGG using only hematoxylin and eosin-stained whole slide images and slide-level biomarker status labels. Specifically, by incorporating the one-class classification into the MIL framework, accurate instance pseudo-labeling is realized for instance-level supervision, which greatly complements the slide-level labels and improves the biomarker prediction performance. Multi-Beholder demonstrates superior prediction performance and generalizability for five LGG biomarkers (AUROC=0.6469-0.9735) in two cohorts (n=607) with diverse races and scanning protocols. Moreover, the excellent interpretability of Multi-Beholder allows for discovering the quantitative and qualitative correlations between biomarker status and histomorphology characteristics. Our pipeline not only provides a novel approach for biomarker prediction, enhancing the applicability of molecular treatments for LGG patients but also facilitates the discovery of new mechanisms in molecular functionality and LGG progression.
Rapid Image Labeling via Neuro-Symbolic Learning
Jun 18, 2023Yifeng Wang, Zhi Tu, Yiwen Xiang, Shiyuan Zhou, Xiyuan Chen, Bingxuan Li, Tianyi Zhang


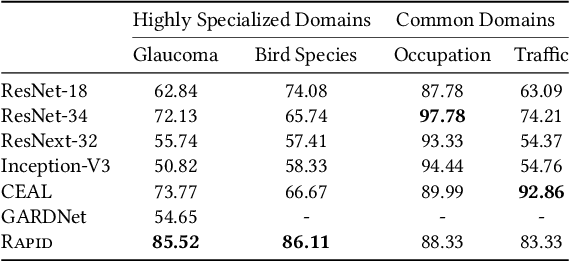
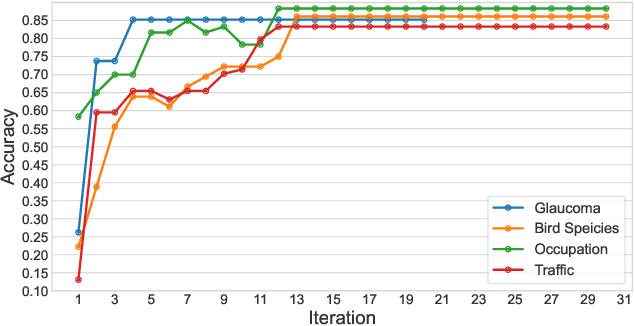
The success of Computer Vision (CV) relies heavily on manually annotated data. However, it is prohibitively expensive to annotate images in key domains such as healthcare, where data labeling requires significant domain expertise and cannot be easily delegated to crowd workers. To address this challenge, we propose a neuro-symbolic approach called Rapid, which infers image labeling rules from a small amount of labeled data provided by domain experts and automatically labels unannotated data using the rules. Specifically, Rapid combines pre-trained CV models and inductive logic learning to infer the logic-based labeling rules. Rapid achieves a labeling accuracy of 83.33% to 88.33% on four image labeling tasks with only 12 to 39 labeled samples. In particular, Rapid significantly outperforms finetuned CV models in two highly specialized tasks. These results demonstrate the effectiveness of Rapid in learning from small data and its capability to generalize among different tasks. Code and our dataset are publicly available at https://github.com/Neural-Symbolic-Image-Labeling/
Learning Agreement from Multi-source Annotations for Medical Image Segmentation
Apr 02, 2023Yifeng Wang, Luyang Luo, Mingxiang Wu, Qiong Wang, Hao Chen
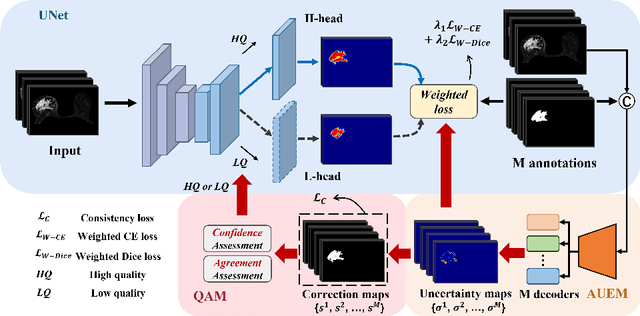

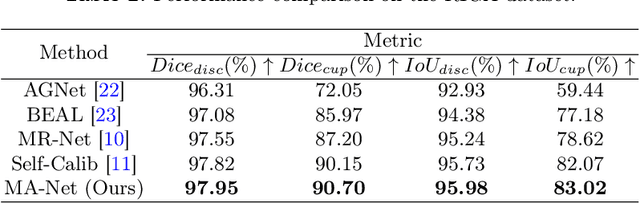
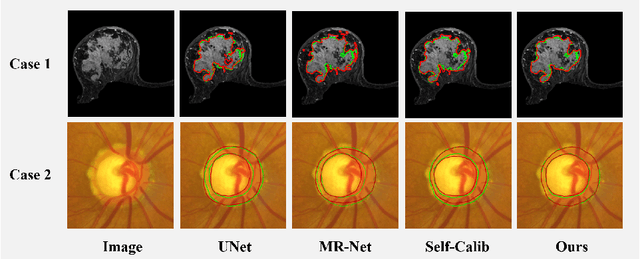
In medical image analysis, it is typical to merge multiple independent annotations as ground truth to mitigate the bias caused by individual annotation preference. However, arbitrating the final annotation is not always effective because new biases might be produced during the process, especially when there are significant variations among annotations. This paper proposes a novel Uncertainty-guided Multi-source Annotation Network (UMA-Net) to learn medical image segmentation directly from multiple annotations. UMA-Net consists of a UNet with two quality-specific predictors, an Annotation Uncertainty Estimation Module (AUEM) and a Quality Assessment Module (QAM). Specifically, AUEM estimates pixel-wise uncertainty maps of each annotation and encourages them to reach an agreement on reliable pixels/voxels. The uncertainty maps then guide the UNet to learn from the reliable pixels/voxels by weighting the segmentation loss. QAM grades the uncertainty maps into high-quality or low-quality groups based on assessment scores. The UNet is further implemented to contain a high-quality learning head (H-head) and a low-quality learning head (L-head). H-head purely learns with high-quality uncertainty maps to avoid error accumulation and keeps strong prediction ability, while L-head leverages the low-quality uncertainty maps to assist the backbone to learn maximum representation knowledge. UNet with H-head will be reserved during the inference stage, and the rest of the modules can be removed freely for computational efficiency. We conduct extensive experiments on an unsupervised 3D segmentation task and a supervised 2D segmentation task, respectively. The results show that our proposed UMA-Net outperforms state-of-the-art approaches, demonstrating its generality and effectiveness.
AugDiff: Diffusion based Feature Augmentation for Multiple Instance Learning in Whole Slide Image
Mar 11, 2023Zhuchen Shao, Liuxi Dai, Yifeng Wang, Haoqian Wang, Yongbing Zhang
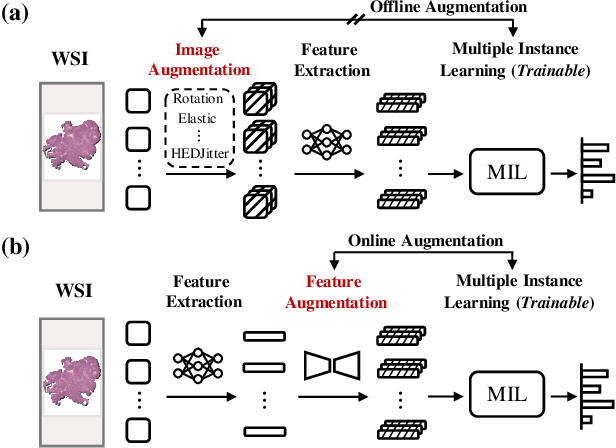

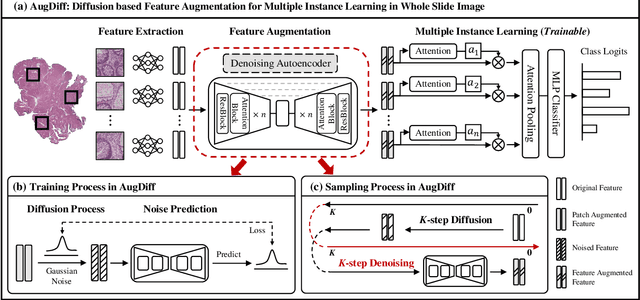
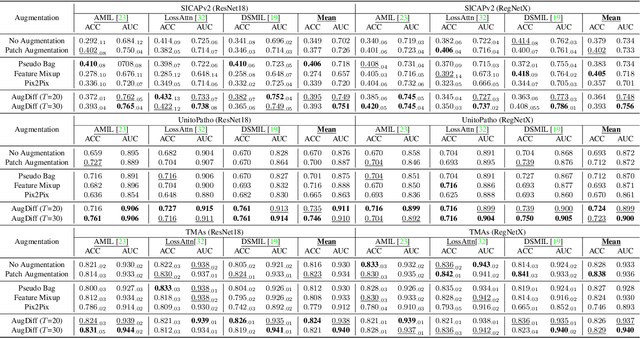
Multiple Instance Learning (MIL), a powerful strategy for weakly supervised learning, is able to perform various prediction tasks on gigapixel Whole Slide Images (WSIs). However, the tens of thousands of patches in WSIs usually incur a vast computational burden for image augmentation, limiting the MIL model's improvement in performance. Currently, the feature augmentation-based MIL framework is a promising solution, while existing methods such as Mixup often produce unrealistic features. To explore a more efficient and practical augmentation method, we introduce the Diffusion Model (DM) into MIL for the first time and propose a feature augmentation framework called AugDiff. Specifically, we employ the generation diversity of DM to improve the quality of feature augmentation and the step-by-step generation property to control the retention of semantic information. We conduct extensive experiments over three distinct cancer datasets, two different feature extractors, and three prevalent MIL algorithms to evaluate the performance of AugDiff. Ablation study and visualization further verify the effectiveness. Moreover, we highlight AugDiff's higher-quality augmented feature over image augmentation and its superiority over self-supervised learning. The generalization over external datasets indicates its broader applications.
Human Health Indicator Prediction from Gait Video
Dec 25, 2022Ziqing Li, Xuexin Yu, Xiaocong Lian, Yifeng Wang, Xiangyang Ji
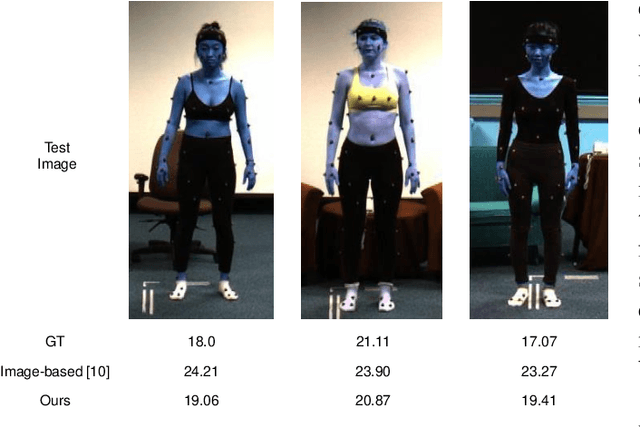
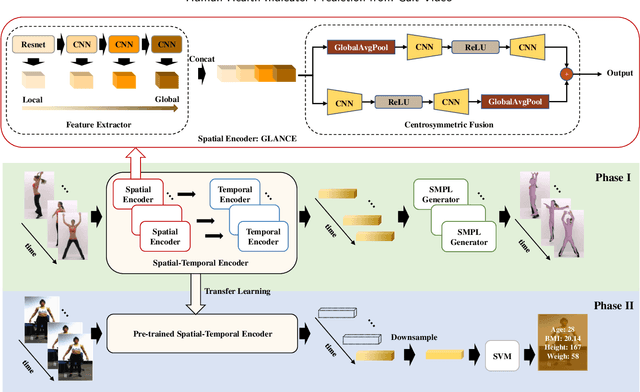
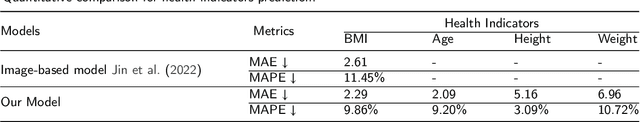

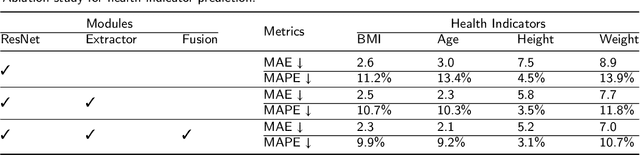
Body Mass Index (BMI), age, height and weight are important indicators of human health conditions, which can provide useful information for plenty of practical purposes, such as health care, monitoring and re-identification. Most existing methods of health indicator prediction mainly use front-view body or face images. These inputs are hard to be obtained in daily life and often lead to the lack of robustness for the models, considering their strict requirements on view and pose. In this paper, we propose to employ gait videos to predict health indicators, which are more prevalent in surveillance and home monitoring scenarios. However, the study of health indicator prediction from gait videos using deep learning was hindered due to the small amount of open-sourced data. To address this issue, we analyse the similarity and relationship between pose estimation and health indicator prediction tasks, and then propose a paradigm enabling deep learning for small health indicator datasets by pre-training on the pose estimation task. Furthermore, to better suit the health indicator prediction task, we bring forward Global-Local Aware aNd Centrosymmetric Encoder (GLANCE) module. It first extracts local and global features by progressive convolutions and then fuses multi-level features by a centrosymmetric double-path hourglass structure in two different ways. Experiments demonstrate that the proposed paradigm achieves state-of-the-art results for predicting health indicators on MoVi, and that the GLANCE module is also beneficial for pose estimation on 3DPW.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge