Guided Combinatorial Algorithms for Submodular Maximization
May 08, 2024Yixin Chen, Ankur Nath, Chunli Peng, Alan Kuhnle
For constrained, not necessarily monotone submodular maximization, guiding the measured continuous greedy algorithm with a local search algorithm currently obtains the state-of-the-art approximation factor of 0.401 \citep{buchbinder2023constrained}. These algorithms rely upon the multilinear extension and the Lovasz extension of a submodular set function. However, the state-of-the-art approximation factor of combinatorial algorithms has remained $1/e \approx 0.367$ \citep{buchbinder2014submodular}. In this work, we develop combinatorial analogues of the guided measured continuous greedy algorithm and obtain approximation ratio of $0.385$ in $\oh{ kn }$ queries to the submodular set function for size constraint, and $0.305$ for a general matroid constraint. Further, we derandomize these algorithms, maintaining the same ratio and asymptotic time complexity. Finally, we develop a deterministic, nearly linear time algorithm with ratio $0.377$.
SPGNN: Recognizing Salient Subgraph Patterns via Enhanced Graph Convolution and Pooling
Apr 29, 2024Zehao Dong, Muhan Zhang, Yixin Chen
Graph neural networks (GNNs) have revolutionized the field of machine learning on non-Euclidean data such as graphs and networks. GNNs effectively implement node representation learning through neighborhood aggregation and achieve impressive results in many graph-related tasks. However, most neighborhood aggregation approaches are summation-based, which can be problematic as they may not be sufficiently expressive to encode informative graph structures. Furthermore, though the graph pooling module is also of vital importance for graph learning, especially for the task of graph classification, research on graph down-sampling mechanisms is rather limited. To address the above challenges, we propose a concatenation-based graph convolution mechanism that injectively updates node representations to maximize the discriminative power in distinguishing non-isomorphic subgraphs. In addition, we design a novel graph pooling module, called WL-SortPool, to learn important subgraph patterns in a deep-learning manner. WL-SortPool layer-wise sorts node representations (i.e. continuous WL colors) to separately learn the relative importance of subtrees with different depths for the purpose of classification, thus better characterizing the complex graph topology and rich information encoded in the graph. We propose a novel Subgraph Pattern GNN (SPGNN) architecture that incorporates these enhancements. We test the proposed SPGNN architecture on many graph classification benchmarks. Experimental results show that our method can achieve highly competitive results with state-of-the-art graph kernels and other GNN approaches.
PhyRecon: Physically Plausible Neural Scene Reconstruction
Apr 28, 2024Junfeng Ni, Yixin Chen, Bohan Jing, Nan Jiang, Bin Wang, Bo Dai, Yixin Zhu, Song-Chun Zhu, Siyuan Huang
While neural implicit representations have gained popularity in multi-view 3D reconstruction, previous work struggles to yield physically plausible results, thereby limiting their applications in physics-demanding domains like embodied AI and robotics. The lack of plausibility originates from both the absence of physics modeling in the existing pipeline and their inability to recover intricate geometrical structures. In this paper, we introduce PhyRecon, which stands as the first approach to harness both differentiable rendering and differentiable physics simulation to learn implicit surface representations. Our framework proposes a novel differentiable particle-based physical simulator seamlessly integrated with the neural implicit representation. At its core is an efficient transformation between SDF-based implicit representation and explicit surface points by our proposed algorithm, Surface Points Marching Cubes (SP-MC), enabling differentiable learning with both rendering and physical losses. Moreover, we model both rendering and physical uncertainty to identify and compensate for the inconsistent and inaccurate monocular geometric priors. The physical uncertainty additionally enables a physics-guided pixel sampling to enhance the learning of slender structures. By amalgamating these techniques, our model facilitates efficient joint modeling with appearance, geometry, and physics. Extensive experiments demonstrate that PhyRecon significantly outperforms all state-of-the-art methods in terms of reconstruction quality. Our reconstruction results also yield superior physical stability, verified by Isaac Gym, with at least a 40% improvement across all datasets, opening broader avenues for future physics-based applications.
Mini-Gemini: Mining the Potential of Multi-modality Vision Language Models
Mar 27, 2024Yanwei Li, Yuechen Zhang, Chengyao Wang, Zhisheng Zhong, Yixin Chen, Ruihang Chu, Shaoteng Liu, Jiaya Jia
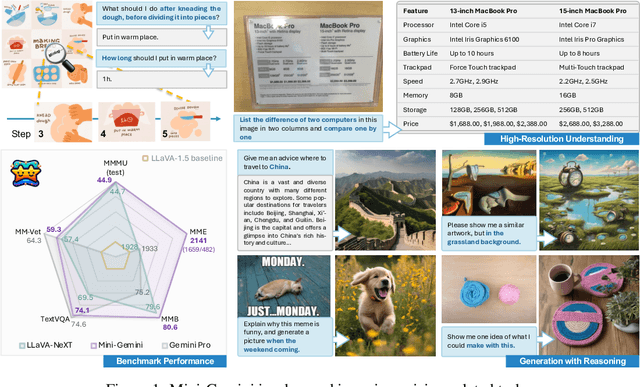
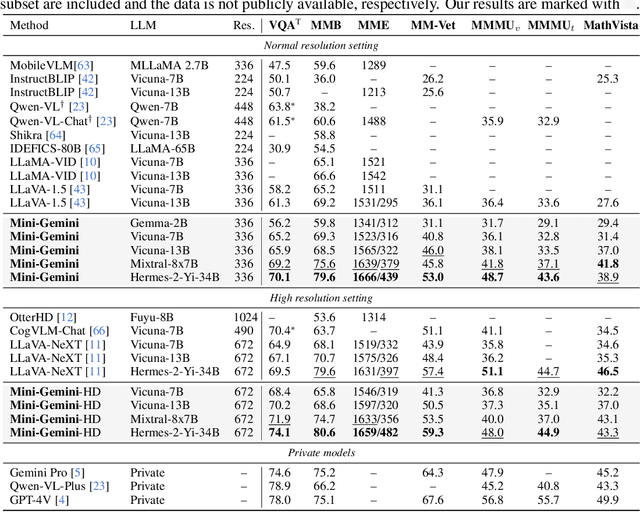
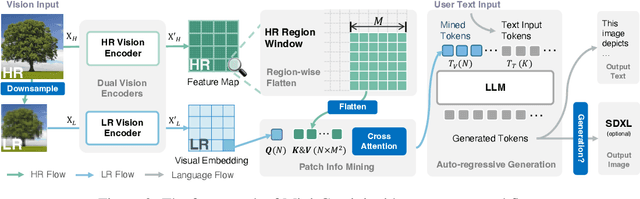
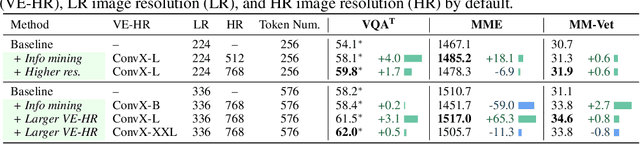
In this work, we introduce Mini-Gemini, a simple and effective framework enhancing multi-modality Vision Language Models (VLMs). Despite the advancements in VLMs facilitating basic visual dialog and reasoning, a performance gap persists compared to advanced models like GPT-4 and Gemini. We try to narrow the gap by mining the potential of VLMs for better performance and any-to-any workflow from three aspects, i.e., high-resolution visual tokens, high-quality data, and VLM-guided generation. To enhance visual tokens, we propose to utilize an additional visual encoder for high-resolution refinement without increasing the visual token count. We further construct a high-quality dataset that promotes precise image comprehension and reasoning-based generation, expanding the operational scope of current VLMs. In general, Mini-Gemini further mines the potential of VLMs and empowers current frameworks with image understanding, reasoning, and generation simultaneously. Mini-Gemini supports a series of dense and MoE Large Language Models (LLMs) from 2B to 34B. It is demonstrated to achieve leading performance in several zero-shot benchmarks and even surpasses the developed private models. Code and models are available at https://github.com/dvlab-research/MiniGemini.
Move as You Say, Interact as You Can: Language-guided Human Motion Generation with Scene Affordance
Mar 26, 2024Zan Wang, Yixin Chen, Baoxiong Jia, Puhao Li, Jinlu Zhang, Jingze Zhang, Tengyu Liu, Yixin Zhu, Wei Liang, Siyuan Huang

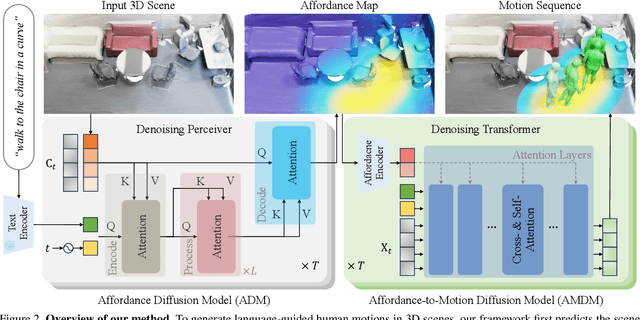


Despite significant advancements in text-to-motion synthesis, generating language-guided human motion within 3D environments poses substantial challenges. These challenges stem primarily from (i) the absence of powerful generative models capable of jointly modeling natural language, 3D scenes, and human motion, and (ii) the generative models' intensive data requirements contrasted with the scarcity of comprehensive, high-quality, language-scene-motion datasets. To tackle these issues, we introduce a novel two-stage framework that employs scene affordance as an intermediate representation, effectively linking 3D scene grounding and conditional motion generation. Our framework comprises an Affordance Diffusion Model (ADM) for predicting explicit affordance map and an Affordance-to-Motion Diffusion Model (AMDM) for generating plausible human motions. By leveraging scene affordance maps, our method overcomes the difficulty in generating human motion under multimodal condition signals, especially when training with limited data lacking extensive language-scene-motion pairs. Our extensive experiments demonstrate that our approach consistently outperforms all baselines on established benchmarks, including HumanML3D and HUMANISE. Additionally, we validate our model's exceptional generalization capabilities on a specially curated evaluation set featuring previously unseen descriptions and scenes.
Scaling Up Dynamic Human-Scene Interaction Modeling
Mar 13, 2024Nan Jiang, Zhiyuan Zhang, Hongjie Li, Xiaoxuan Ma, Zan Wang, Yixin Chen, Tengyu Liu, Yixin Zhu, Siyuan Huang
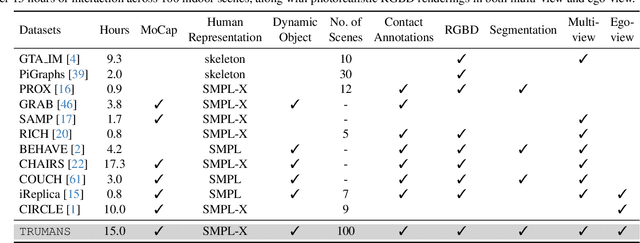
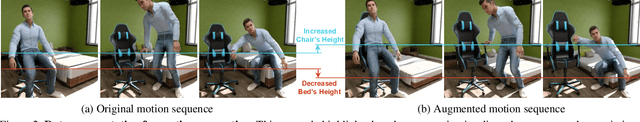
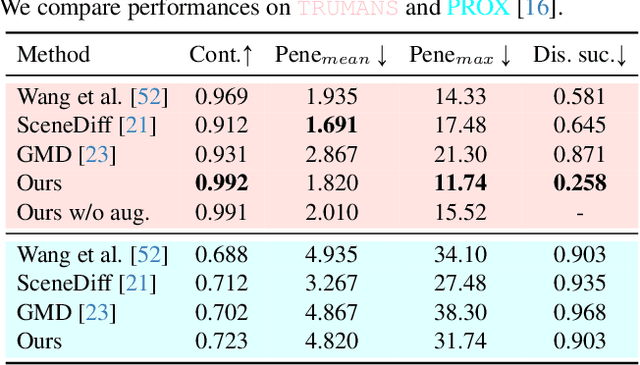
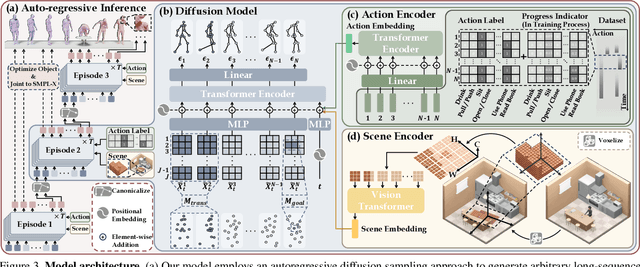
Confronting the challenges of data scarcity and advanced motion synthesis in human-scene interaction modeling, we introduce the TRUMANS dataset alongside a novel HSI motion synthesis method. TRUMANS stands as the most comprehensive motion-captured HSI dataset currently available, encompassing over 15 hours of human interactions across 100 indoor scenes. It intricately captures whole-body human motions and part-level object dynamics, focusing on the realism of contact. This dataset is further scaled up by transforming physical environments into exact virtual models and applying extensive augmentations to appearance and motion for both humans and objects while maintaining interaction fidelity. Utilizing TRUMANS, we devise a diffusion-based autoregressive model that efficiently generates HSI sequences of any length, taking into account both scene context and intended actions. In experiments, our approach shows remarkable zero-shot generalizability on a range of 3D scene datasets (e.g., PROX, Replica, ScanNet, ScanNet++), producing motions that closely mimic original motion-captured sequences, as confirmed by quantitative experiments and human studies.
Large-Language-Model Empowered Dose Volume Histogram Prediction for Intensity Modulated Radiotherapy
Feb 11, 2024Zehao Dong, Yixin Chen, Hiram Gay, Yao Hao, Geoffrey D. Hugo, Pamela Samson, Tianyu Zhao
Treatment planning is currently a patient specific, time-consuming, and resource demanding task in radiotherapy. Dose-volume histogram (DVH) prediction plays a critical role in automating this process. The geometric relationship between DVHs in radiotherapy plans and organs-at-risk (OAR) and planning target volume (PTV) has been well established. This study explores the potential of deep learning models for predicting DVHs using images and subsequent human intervention facilitated by a large-language model (LLM) to enhance the planning quality. We propose a pipeline to convert unstructured images to a structured graph consisting of image-patch nodes and dose nodes. A novel Dose Graph Neural Network (DoseGNN) model is developed for predicting DVHs from the structured graph. The proposed DoseGNN is enhanced with the LLM to encode massive knowledge from prescriptions and interactive instructions from clinicians. In this study, we introduced an online human-AI collaboration (OHAC) system as a practical implementation of the concept proposed for the automation of intensity-modulated radiotherapy (IMRT) planning. In comparison to the widely-employed DL models used in radiotherapy, DoseGNN achieved mean square errors that were 80$\%$, 76$\%$ and 41.0$\%$ of those predicted by Swin U-Net Transformer, 3D U-Net CNN and vanilla MLP, respectively. Moreover, the LLM-empowered DoseGNN model facilitates seamless adjustment to treatment plans through interaction with clinicians using natural language.
Highly Accurate Disease Diagnosis and Highly Reproducible Biomarker Identification with PathFormer
Feb 11, 2024Zehao Dong, Qihang Zhao, Philip R. O. Payne, Michael A Province, Carlos Cruchaga, Muhan Zhang, Tianyu Zhao, Yixin Chen, Fuhai Li
Biomarker identification is critical for precise disease diagnosis and understanding disease pathogenesis in omics data analysis, like using fold change and regression analysis. Graph neural networks (GNNs) have been the dominant deep learning model for analyzing graph-structured data. However, we found two major limitations of existing GNNs in omics data analysis, i.e., limited-prediction (diagnosis) accuracy and limited-reproducible biomarker identification capacity across multiple datasets. The root of the challenges is the unique graph structure of biological signaling pathways, which consists of a large number of targets and intensive and complex signaling interactions among these targets. To resolve these two challenges, in this study, we presented a novel GNN model architecture, named PathFormer, which systematically integrate signaling network, priori knowledge and omics data to rank biomarkers and predict disease diagnosis. In the comparison results, PathFormer outperformed existing GNN models significantly in terms of highly accurate prediction capability ( 30% accuracy improvement in disease diagnosis compared with existing GNN models) and high reproducibility of biomarker ranking across different datasets. The improvement was confirmed using two independent Alzheimer's Disease (AD) and cancer transcriptomic datasets. The PathFormer model can be directly applied to other omics data analysis studies.
Large Language Model Meets Graph Neural Network in Knowledge Distillation
Feb 09, 2024Shengxiang Hu, Guobing Zou, Song Yang, Yanglan Gan, Bofeng Zhang, Yixin Chen
Despite recent community revelations about the advancements and potential applications of Large Language Models (LLMs) in understanding Text-Attributed Graph (TAG), the deployment of LLMs for production is hindered by its high computational and storage requirements, as well as long latencies during model inference. Simultaneously, although traditional Graph Neural Networks (GNNs) are light weight and adept at learning structural features of graphs, their ability to grasp the complex semantics in TAG is somewhat constrained for real applications. To address these limitations, we concentrate on the downstream task of node classification in TAG and propose a novel graph knowledge distillation framework, termed Linguistic Graph Knowledge Distillation (LinguGKD), using LLMs as teacher models and GNNs as student models for knowledge distillation. It involves TAG-oriented instruction tuning of LLM on designed tailored prompts, followed by propagating knowledge and aligning the hierarchically learned node features from the teacher LLM to the student GNN in latent space, employing a layer-adaptive contrastive learning strategy. Through extensive experiments on a variety of LLM and GNN models and multiple benchmark datasets, the proposed LinguGKD significantly boosts the student GNN's predictive accuracy and convergence rate, without the need of extra data or model parameters. Compared to teacher LLM, distilled GNN achieves superior inference speed equipped with much fewer computing and storage demands, when surpassing the teacher LLM's classification accuracy on some of benchmark datasets.
DoseGNN: Improving the Performance of Deep Learning Models in Adaptive Dose-Volume Histogram Prediction through Graph Neural Networks
Feb 02, 2024Zehao Dong, Yixin Chen, Tianyu Zhao
Dose-Volume Histogram (DVH) prediction is fundamental in radiation therapy that facilitate treatment planning, dose evaluation, plan comparison and etc. It helps to increase the ability to deliver precise and effective radiation treatments while managing potential toxicities to healthy tissues as needed to reduce the risk of complications. This paper extends recently disclosed research findings presented on AAPM (AAPM 65th Annual Meeting $\&$ Exhibition) and includes necessary technique details. The objective is to design efficient deep learning models for DVH prediction on general radiotherapy platform equipped with high performance CBCT system, where input CT images and target dose images to predict may have different origins, spacing and sizes. Deep learning models widely-adopted in DVH prediction task are evaluated on the novel radiotherapy platform, and graph neural networks (GNNs) are shown to be the ideal architecture to construct a plug-and-play framework to improve predictive performance of base deep learning models in the adaptive setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge