RaffeSDG: Random Frequency Filtering enabled Single-source Domain Generalization for Medical Image Segmentation
May 02, 2024Heng Li, Haojin Li, Jianyu Chen, Zhongxi Qiu, Huazhu Fu, Lidai Wang, Yan Hu, Jiang Liu
Deep learning models often encounter challenges in making accurate inferences when there are domain shifts between the source and target data. This issue is particularly pronounced in clinical settings due to the scarcity of annotated data resulting from the professional and private nature of medical data. Despite the existence of decent solutions, many of them are hindered in clinical settings due to limitations in data collection and computational complexity. To tackle domain shifts in data-scarce medical scenarios, we propose a Random frequency filtering enabled Single-source Domain Generalization algorithm (RaffeSDG), which promises robust out-of-domain inference with segmentation models trained on a single-source domain. A filter-based data augmentation strategy is first proposed to promote domain variability within a single-source domain by introducing variations in frequency space and blending homologous samples. Then Gaussian filter-based structural saliency is also leveraged to learn robust representations across augmented samples, further facilitating the training of generalizable segmentation models. To validate the effectiveness of RaffeSDG, we conducted extensive experiments involving out-of-domain inference on segmentation tasks for three human tissues imaged by four diverse modalities. Through thorough investigations and comparisons, compelling evidence was observed in these experiments, demonstrating the potential and generalizability of RaffeSDG. The code is available at https://github.com/liamheng/Non-IID_Medical_Image_Segmentation.
An Aggregation-Free Federated Learning for Tackling Data Heterogeneity
Apr 29, 2024Yuan Wang, Huazhu Fu, Renuga Kanagavelu, Qingsong Wei, Yong Liu, Rick Siow Mong Goh
The performance of Federated Learning (FL) hinges on the effectiveness of utilizing knowledge from distributed datasets. Traditional FL methods adopt an aggregate-then-adapt framework, where clients update local models based on a global model aggregated by the server from the previous training round. This process can cause client drift, especially with significant cross-client data heterogeneity, impacting model performance and convergence of the FL algorithm. To address these challenges, we introduce FedAF, a novel aggregation-free FL algorithm. In this framework, clients collaboratively learn condensed data by leveraging peer knowledge, the server subsequently trains the global model using the condensed data and soft labels received from the clients. FedAF inherently avoids the issue of client drift, enhances the quality of condensed data amid notable data heterogeneity, and improves the global model performance. Extensive numerical studies on several popular benchmark datasets show FedAF surpasses various state-of-the-art FL algorithms in handling label-skew and feature-skew data heterogeneity, leading to superior global model accuracy and faster convergence.
Multimodal Fusion on Low-quality Data: A Comprehensive Survey
Apr 27, 2024Qingyang Zhang, Yake Wei, Zongbo Han, Huazhu Fu, Xi Peng, Cheng Deng, Qinghua Hu, Cai Xu, Jie Wen, Di Hu, Changqing Zhang
Multimodal fusion focuses on integrating information from multiple modalities with the goal of more accurate prediction, which has achieved remarkable progress in a wide range of scenarios, including autonomous driving and medical diagnosis. However, the reliability of multimodal fusion remains largely unexplored especially under low-quality data settings. This paper surveys the common challenges and recent advances of multimodal fusion in the wild and presents them in a comprehensive taxonomy. From a data-centric view, we identify four main challenges that are faced by multimodal fusion on low-quality data, namely (1) noisy multimodal data that are contaminated with heterogeneous noises, (2) incomplete multimodal data that some modalities are missing, (3) imbalanced multimodal data that the qualities or properties of different modalities are significantly different and (4) quality-varying multimodal data that the quality of each modality dynamically changes with respect to different samples. This new taxonomy will enable researchers to understand the state of the field and identify several potential directions. We also provide discussion for the open problems in this field together with interesting future research directions.
MedRG: Medical Report Grounding with Multi-modal Large Language Model
Apr 10, 2024Ke Zou, Yang Bai, Zhihao Chen, Yang Zhou, Yidi Chen, Kai Ren, Meng Wang, Xuedong Yuan, Xiaojing Shen, Huazhu Fu
Medical Report Grounding is pivotal in identifying the most relevant regions in medical images based on a given phrase query, a critical aspect in medical image analysis and radiological diagnosis. However, prevailing visual grounding approaches necessitate the manual extraction of key phrases from medical reports, imposing substantial burdens on both system efficiency and physicians. In this paper, we introduce a novel framework, Medical Report Grounding (MedRG), an end-to-end solution for utilizing a multi-modal Large Language Model to predict key phrase by incorporating a unique token, BOX, into the vocabulary to serve as an embedding for unlocking detection capabilities. Subsequently, the vision encoder-decoder jointly decodes the hidden embedding and the input medical image, generating the corresponding grounding box. The experimental results validate the effectiveness of MedRG, surpassing the performance of the existing state-of-the-art medical phrase grounding methods. This study represents a pioneering exploration of the medical report grounding task, marking the first-ever endeavor in this domain.
Instrument-tissue Interaction Detection Framework for Surgical Video Understanding
Mar 30, 2024Wenjun Lin, Yan Hu, Huazhu Fu, Mingming Yang, Chin-Boon Chng, Ryo Kawasaki, Cheekong Chui, Jiang Liu
Instrument-tissue interaction detection task, which helps understand surgical activities, is vital for constructing computer-assisted surgery systems but with many challenges. Firstly, most models represent instrument-tissue interaction in a coarse-grained way which only focuses on classification and lacks the ability to automatically detect instruments and tissues. Secondly, existing works do not fully consider relations between intra- and inter-frame of instruments and tissues. In the paper, we propose to represent instrument-tissue interaction as <instrument class, instrument bounding box, tissue class, tissue bounding box, action class> quintuple and present an Instrument-Tissue Interaction Detection Network (ITIDNet) to detect the quintuple for surgery videos understanding. Specifically, we propose a Snippet Consecutive Feature (SCF) Layer to enhance features by modeling relationships of proposals in the current frame using global context information in the video snippet. We also propose a Spatial Corresponding Attention (SCA) Layer to incorporate features of proposals between adjacent frames through spatial encoding. To reason relationships between instruments and tissues, a Temporal Graph (TG) Layer is proposed with intra-frame connections to exploit relationships between instruments and tissues in the same frame and inter-frame connections to model the temporal information for the same instance. For evaluation, we build a cataract surgery video (PhacoQ) dataset and a cholecystectomy surgery video (CholecQ) dataset. Experimental results demonstrate the promising performance of our model, which outperforms other state-of-the-art models on both datasets.
AIC-UNet: Anatomy-informed Cascaded UNet for Robust Multi-Organ Segmentation
Mar 27, 2024Young Seok Jeon, Hongfei Yang, Huazhu Fu, Mengling Feng
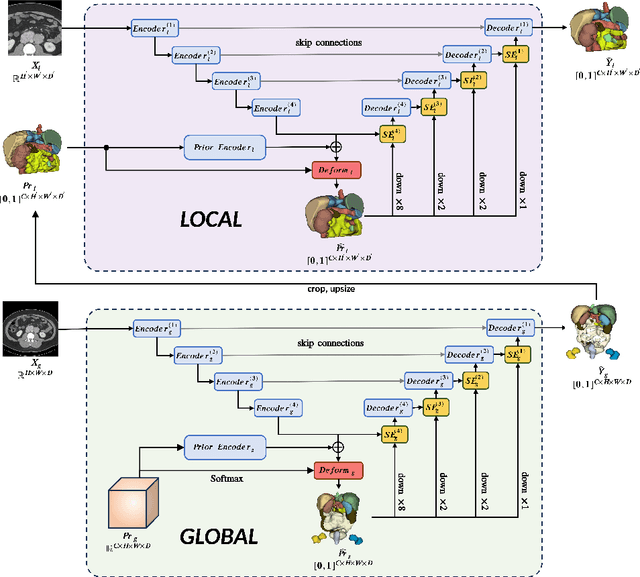
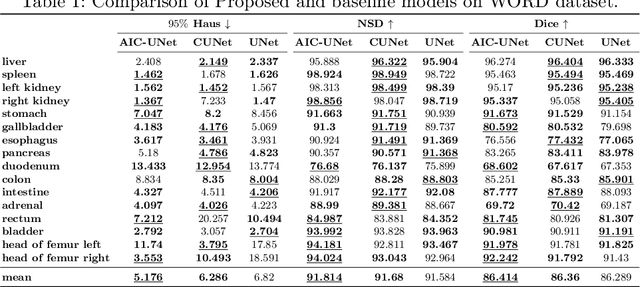

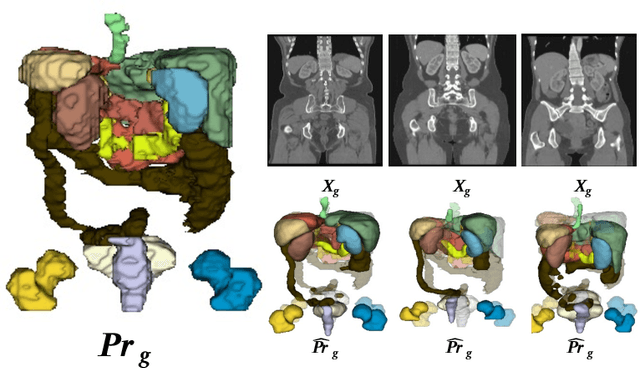
Imposing key anatomical features, such as the number of organs, their shapes, sizes, and relative positions, is crucial for building a robust multi-organ segmentation model. Current attempts to incorporate anatomical features include broadening effective receptive fields (ERF) size with resource- and data-intensive modules such as self-attention or introducing organ-specific topology regularizers, which may not scale to multi-organ segmentation problems where inter-organ relation also plays a huge role. We introduce a new approach to impose anatomical constraints on any existing encoder-decoder segmentation model by conditioning model prediction with learnable anatomy prior. More specifically, given an abdominal scan, a part of the encoder spatially warps a learnable prior to align with the given input scan using thin plate spline (TPS) grid interpolation. The warped prior is then integrated during the decoding phase to guide the model for more anatomy-informed predictions. Code is available at \hyperlink{https://anonymous.4open.science/r/AIC-UNet-7048}{https://anonymous.4open.science/r/AIC-UNet-7048}.
Training-free image style alignment for self-adapting domain shift on handheld ultrasound devices
Feb 17, 2024Hongye Zeng, Ke Zou, Zhihao Chen, Yuchong Gao, Hongbo Chen, Haibin Zhang, Kang Zhou, Meng Wang, Rick Siow Mong Goh, Yong Liu, Chang Jiang, Rui Zheng, Huazhu Fu
Handheld ultrasound devices face usage limitations due to user inexperience and cannot benefit from supervised deep learning without extensive expert annotations. Moreover, the models trained on standard ultrasound device data are constrained by training data distribution and perform poorly when directly applied to handheld device data. In this study, we propose the Training-free Image Style Alignment (TISA) framework to align the style of handheld device data to those of standard devices. The proposed TISA can directly infer handheld device images without extra training and is suited for clinical applications. We show that TISA performs better and more stably in medical detection and segmentation tasks for handheld device data. We further validate TISA as the clinical model for automatic measurements of spinal curvature and carotid intima-media thickness. The automatic measurements agree well with manual measurements made by human experts and the measurement errors remain within clinically acceptable ranges. We demonstrate the potential for TISA to facilitate automatic diagnosis on handheld ultrasound devices and expedite their eventual widespread use.
Dual-scale Enhanced and Cross-generative Consistency Learning for Semi-supervised Polyp Segmentation
Dec 26, 2023Yunqi Gu, Tao Zhou, Yizhe Zhang, Yi Zhou, Kelei He, Chen Gong, Huazhu Fu
Automatic polyp segmentation plays a crucial role in the early diagnosis and treatment of colorectal cancer (CRC). However, existing methods heavily rely on fully supervised training, which requires a large amount of labeled data with time-consuming pixel-wise annotations. Moreover, accurately segmenting polyps poses challenges due to variations in shape, size, and location. To address these issues, we propose a novel Dual-scale Enhanced and Cross-generative consistency learning framework for semi-supervised polyp Segmentation (DEC-Seg) from colonoscopy images. First, we propose a Cross-level Feature Aggregation (CFA) module that integrates cross-level adjacent layers to enhance the feature representation ability across different resolutions. To address scale variation, we present a scale-enhanced consistency constraint, which ensures consistency in the segmentation maps generated from the same input image at different scales. This constraint helps handle variations in polyp sizes and improves the robustness of the model. Additionally, we design a scale-aware perturbation consistency scheme to enhance the robustness of the mean teacher model. Furthermore, we propose a cross-generative consistency scheme, in which the original and perturbed images can be reconstructed using cross-segmentation maps. This consistency constraint allows us to mine effective feature representations and boost the segmentation performance. To produce more accurate segmentation maps, we propose a Dual-scale Complementary Fusion (DCF) module that integrates features from two scale-specific decoders operating at different scales. Extensive experimental results on five benchmark datasets demonstrate the effectiveness of our DEC-Seg against other state-of-the-art semi-supervised segmentation approaches. The implementation code will be released at https://github.com/taozh2017/DECSeg.
VSR-Net: Vessel-like Structure Rehabilitation Network with Graph Clustering
Dec 20, 2023Haili Ye, Xiaoqing Zhang, Yan Hu, Huazhu Fu, Jiang Liu
The morphologies of vessel-like structures, such as blood vessels and nerve fibres, play significant roles in disease diagnosis, e.g., Parkinson's disease. Deep network-based refinement segmentation methods have recently achieved promising vessel-like structure segmentation results. There are still two challenges: (1) existing methods have limitations in rehabilitating subsection ruptures in segmented vessel-like structures; (2) they are often overconfident in predicted segmentation results. To tackle these two challenges, this paper attempts to leverage the potential of spatial interconnection relationships among subsection ruptures from the structure rehabilitation perspective. Based on this, we propose a novel Vessel-like Structure Rehabilitation Network (VSR-Net) to rehabilitate subsection ruptures and improve the model calibration based on coarse vessel-like structure segmentation results. VSR-Net first constructs subsection rupture clusters with Curvilinear Clustering Module (CCM). Then, the well-designed Curvilinear Merging Module (CMM) is applied to rehabilitate the subsection ruptures to obtain the refined vessel-like structures. Extensive experiments on five 2D/3D medical image datasets show that VSR-Net significantly outperforms state-of-the-art (SOTA) refinement segmentation methods with lower calibration error. Additionally, we provide quantitative analysis to explain the morphological difference between the rehabilitation results of VSR-Net and ground truth (GT), which is smaller than SOTA methods and GT, demonstrating that our method better rehabilitates vessel-like structures by restoring subsection ruptures.
Enhancing and Adapting in the Clinic: Source-free Unsupervised Domain Adaptation for Medical Image Enhancement
Dec 03, 2023Heng Li, Ziqin Lin, Zhongxi Qiu, Zinan Li, Huazhu Fu, Yan Hu, Jiang Liu
Medical imaging provides many valuable clues involving anatomical structure and pathological characteristics. However, image degradation is a common issue in clinical practice, which can adversely impact the observation and diagnosis by physicians and algorithms. Although extensive enhancement models have been developed, these models require a well pre-training before deployment, while failing to take advantage of the potential value of inference data after deployment. In this paper, we raise an algorithm for source-free unsupervised domain adaptive medical image enhancement (SAME), which adapts and optimizes enhancement models using test data in the inference phase. A structure-preserving enhancement network is first constructed to learn a robust source model from synthesized training data. Then a teacher-student model is initialized with the source model and conducts source-free unsupervised domain adaptation (SFUDA) by knowledge distillation with the test data. Additionally, a pseudo-label picker is developed to boost the knowledge distillation of enhancement tasks. Experiments were implemented on ten datasets from three medical image modalities to validate the advantage of the proposed algorithm, and setting analysis and ablation studies were also carried out to interpret the effectiveness of SAME. The remarkable enhancement performance and benefits for downstream tasks demonstrate the potential and generalizability of SAME. The code is available at https://github.com/liamheng/Annotation-free-Medical-Image-Enhancement.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge