OpenMEDLab: An Open-source Platform for Multi-modality Foundation Models in Medicine
Mar 04, 2024Xiaosong Wang, Xiaofan Zhang, Guotai Wang, Junjun He, Zhongyu Li, Wentao Zhu, Yi Guo, Qi Dou, Xiaoxiao Li, Dequan Wang, Liang Hong, Qicheng Lao, Tong Ruan, Yukun Zhou, Yixue Li, Jie Zhao, Kang Li, Xin Sun, Lifeng Zhu, Shaoting Zhang
The emerging trend of advancing generalist artificial intelligence, such as GPTv4 and Gemini, has reshaped the landscape of research (academia and industry) in machine learning and many other research areas. However, domain-specific applications of such foundation models (e.g., in medicine) remain untouched or often at their very early stages. It will require an individual set of transfer learning and model adaptation techniques by further expanding and injecting these models with domain knowledge and data. The development of such technologies could be largely accelerated if the bundle of data, algorithms, and pre-trained foundation models were gathered together and open-sourced in an organized manner. In this work, we present OpenMEDLab, an open-source platform for multi-modality foundation models. It encapsulates not only solutions of pioneering attempts in prompting and fine-tuning large language and vision models for frontline clinical and bioinformatic applications but also building domain-specific foundation models with large-scale multi-modal medical data. Importantly, it opens access to a group of pre-trained foundation models for various medical image modalities, clinical text, protein engineering, etc. Inspiring and competitive results are also demonstrated for each collected approach and model in a variety of benchmarks for downstream tasks. We welcome researchers in the field of medical artificial intelligence to continuously contribute cutting-edge methods and models to OpenMEDLab, which can be accessed via https://github.com/openmedlab.
Neural Network Diffusion
Feb 20, 2024Kai Wang, Zhaopan Xu, Yukun Zhou, Zelin Zang, Trevor Darrell, Zhuang Liu, Yang You
Diffusion models have achieved remarkable success in image and video generation. In this work, we demonstrate that diffusion models can also \textit{generate high-performing neural network parameters}. Our approach is simple, utilizing an autoencoder and a standard latent diffusion model. The autoencoder extracts latent representations of a subset of the trained network parameters. A diffusion model is then trained to synthesize these latent parameter representations from random noise. It then generates new representations that are passed through the autoencoder's decoder, whose outputs are ready to use as new subsets of network parameters. Across various architectures and datasets, our diffusion process consistently generates models of comparable or improved performance over trained networks, with minimal additional cost. Notably, we empirically find that the generated models perform differently with the trained networks. Our results encourage more exploration on the versatile use of diffusion models.
Expectation Maximization Pseudo Labelling for Segmentation with Limited Annotations
May 02, 2023Mou-Cheng Xu, Yukun Zhou, Chen Jin, Marius de Groot, Daniel C. Alexander, Neil P. Oxtoby, Yipeng Hu, Joseph Jacob
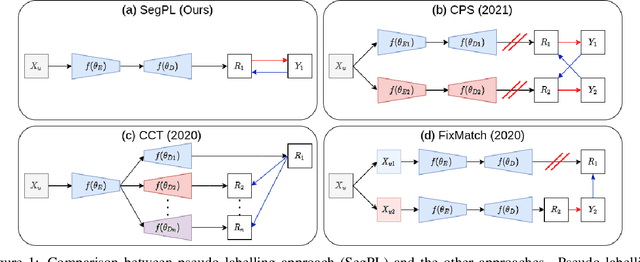
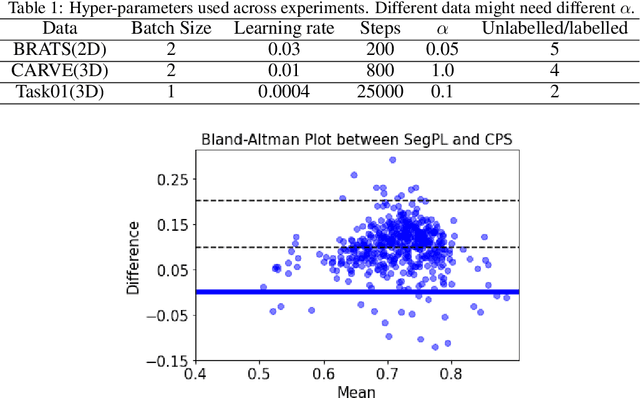
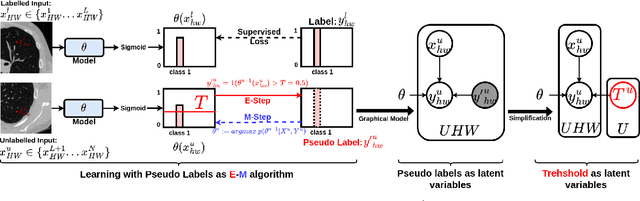
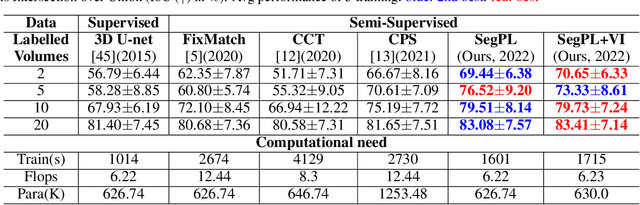
We study pseudo labelling and its generalisation for semi-supervised segmentation of medical images. Pseudo labelling has achieved great empirical successes in semi-supervised learning, by utilising raw inferences on unlabelled data as pseudo labels for self-training. In our paper, we build a connection between pseudo labelling and the Expectation Maximization algorithm which partially explains its empirical successes. We thereby realise that the original pseudo labelling is an empirical estimation of its underlying full formulation. Following this insight, we demonstrate the full generalisation of pseudo labels under Bayes' principle, called Bayesian Pseudo Labels. We then provide a variational approach to learn to approximate Bayesian Pseudo Labels, by learning a threshold to select good quality pseudo labels. In the rest of the paper, we demonstrate the applications of Pseudo Labelling and its generalisation Bayesian Psuedo Labelling in semi-supervised segmentation of medical images on: 1) 3D binary segmentation of lung vessels from CT volumes; 2) 2D multi class segmentation of brain tumours from MRI volumes; 3) 3D binary segmentation of brain tumours from MRI volumes. We also show that pseudo labels can enhance the robustness of the learnt representations.
An Experiment Design Paradigm using Joint Feature Selection and Task Optimization
Oct 13, 2022Stefano B. Blumberg, Hongxiang Lin, Yukun Zhou, Paddy Slator, Daniel C. Alexander
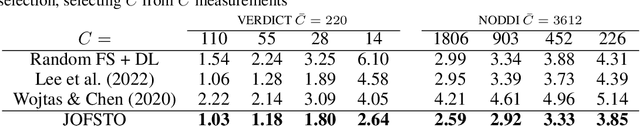
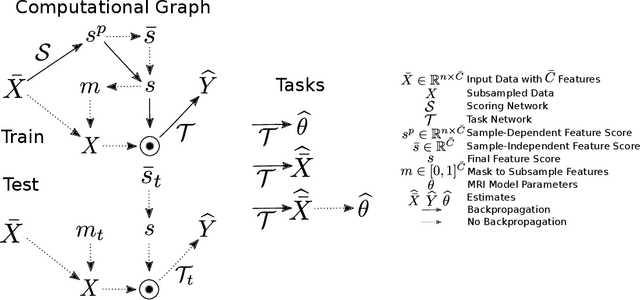
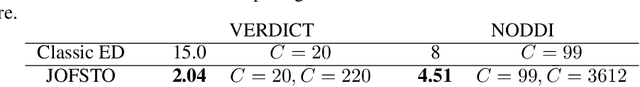

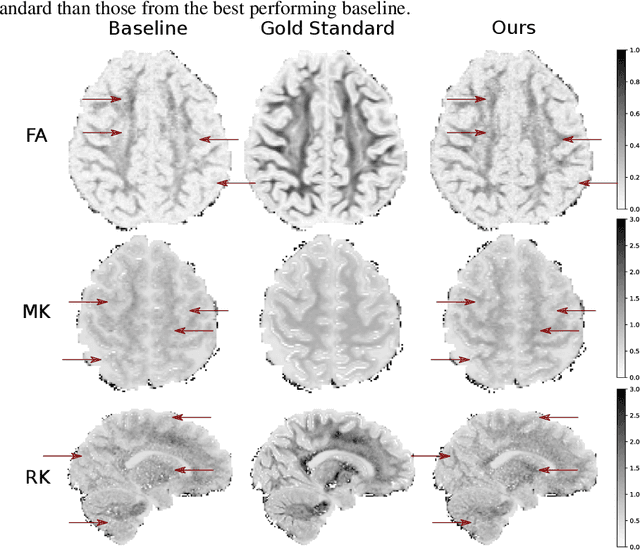
This paper presents a subsampling-task paradigm for data-driven task-specific experiment design (ED) and a novel method in populationwide supervised feature selection (FS). Optimal ED, the choice of sampling points under constraints of limited acquisition-time, arises in a wide variety of scientific and engineering contexts. However the continuous optimization used in classical approaches depend on a-priori parameter choices and challenging non-convex optimization landscapes. This paper proposes to replace this strategy with a subsampling-task paradigm, analogous to populationwide supervised FS. In particular, we introduce JOFSTO, which performs JOint Feature Selection and Task Optimization. JOFSTO jointly optimizes two coupled networks: one for feature scoring, which provides the ED, the other for execution of a downstream task or process. Unlike most FS problems, e.g. selecting protein expressions for classification, ED problems typically select from highly correlated globally informative candidates rather than seeking a small number of highly informative features among many uninformative features. JOFSTO's construction efficiently identifies potentially correlated, but effective subsets and returns a trained task network. We demonstrate the approach using parameter estimation and mapping problems in quantitative MRI, where economical ED is crucial for clinical application. Results from simulations and empirical data show the subsampling-task paradigm strongly outperforms classical ED, and within our paradigm, JOFSTO outperforms state-of-the-art supervised FS techniques. JOFSTO extends immediately to wider image-based ED problems and other scenarios where the design must be specified globally across large numbers of acquisitions. Code will be released.
Bayesian Pseudo Labels: Expectation Maximization for Robust and Efficient Semi-Supervised Segmentation
Aug 08, 2022Mou-Cheng Xu, Yukun Zhou, Chen Jin, Marius de Groot, Daniel C. Alexander, Neil P. Oxtoby, Yipeng Hu, Joseph Jacob
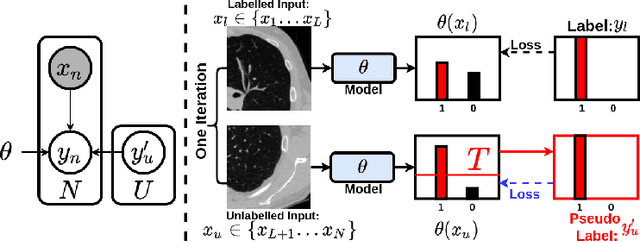
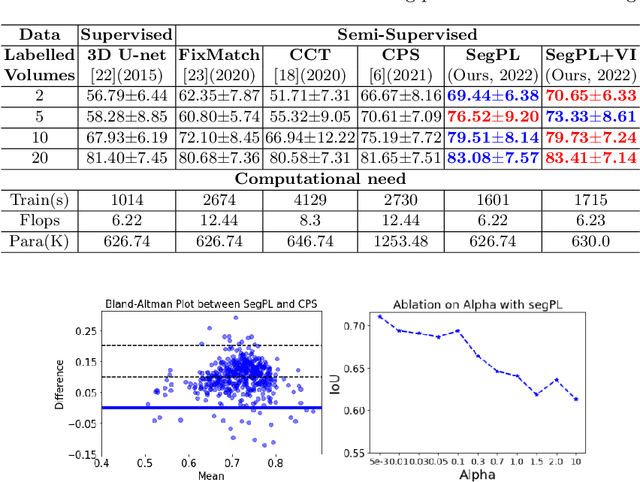
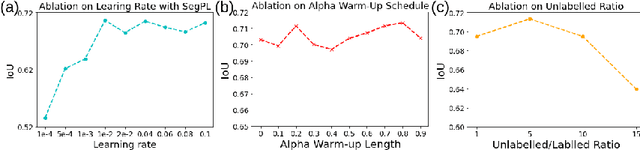
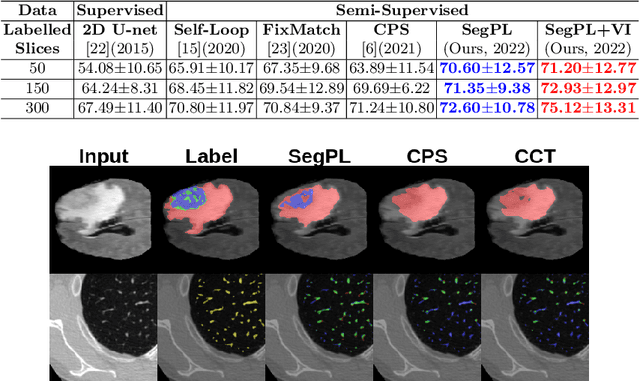
This paper concerns pseudo labelling in segmentation. Our contribution is fourfold. Firstly, we present a new formulation of pseudo-labelling as an Expectation-Maximization (EM) algorithm for clear statistical interpretation. Secondly, we propose a semi-supervised medical image segmentation method purely based on the original pseudo labelling, namely SegPL. We demonstrate SegPL is a competitive approach against state-of-the-art consistency regularisation based methods on semi-supervised segmentation on a 2D multi-class MRI brain tumour segmentation task and a 3D binary CT lung vessel segmentation task. The simplicity of SegPL allows less computational cost comparing to prior methods. Thirdly, we demonstrate that the effectiveness of SegPL may originate from its robustness against out-of-distribution noises and adversarial attacks. Lastly, under the EM framework, we introduce a probabilistic generalisation of SegPL via variational inference, which learns a dynamic threshold for pseudo labelling during the training. We show that SegPL with variational inference can perform uncertainty estimation on par with the gold-standard method Deep Ensemble.
Progressive Subsampling for Oversampled Data -- Application to Quantitative MRI
Apr 08, 2022Stefano B. Blumberg, Hongxiang Lin, Francesco Grussu, Yukun Zhou, Matteo Figini, Daniel C. Alexander
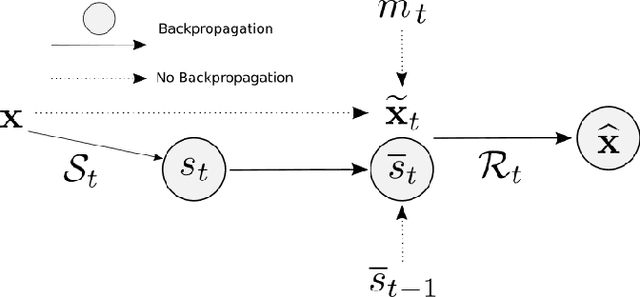
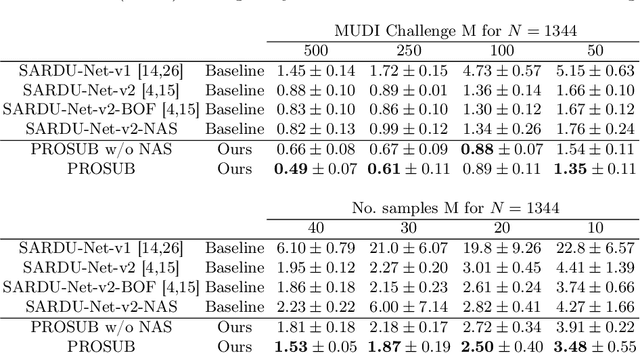
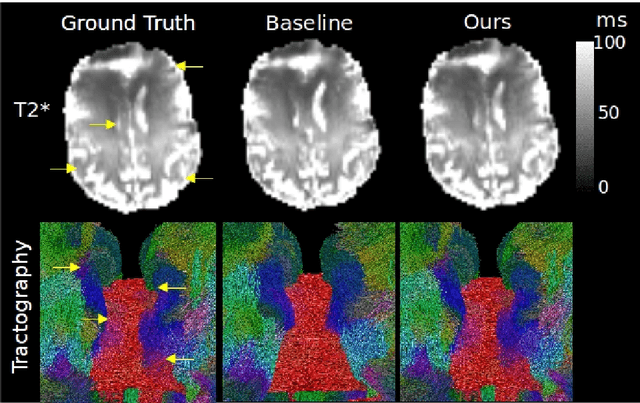
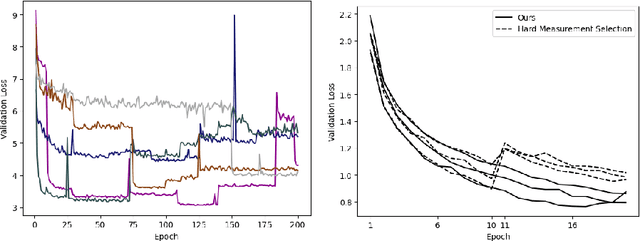
We present PROSUB: PROgressive SUBsampling, a deep learning based, automated methodology that subsamples an oversampled data set (e.g. multi-channeled 3D images) with minimal loss of information. We build upon a recent dual-network approach that won the MICCAI MUlti-DIffusion (MUDI) quantitative MRI measurement sampling-reconstruction challenge, but suffers from deep learning training instability, by subsampling with a hard decision boundary. PROSUB uses the paradigm of recursive feature elimination (RFE) and progressively subsamples measurements during deep learning training, improving optimization stability. PROSUB also integrates a neural architecture search (NAS) paradigm, allowing the network architecture hyperparameters to respond to the subsampling process. We show PROSUB outperforms the winner of the MUDI MICCAI challenge, producing large improvements >18% MSE on the MUDI challenge sub-tasks and qualitative improvements on downstream processes useful for clinical applications. We also show the benefits of incorporating NAS and analyze the effect of PROSUB's components. As our method generalizes to other problems beyond MRI measurement selection-reconstruction, our code is https://github.com/sbb-gh/PROSUB
VAFO-Loss: VAscular Feature Optimised Loss Function for Retinal Artery/Vein Segmentation
Mar 12, 2022Yukun Zhou, Moucheng Xu, Yipeng Hu, Stefano B. Blumberg, An Zhao, Siegfried K. Wagner, Pearse A. Keane, Daniel C. Alexander
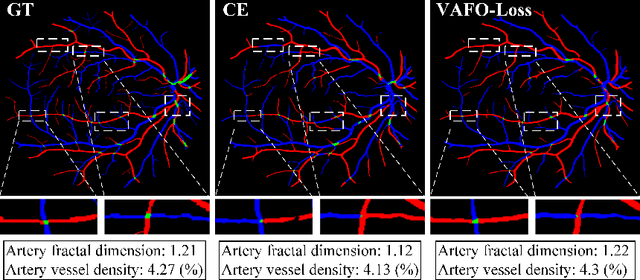
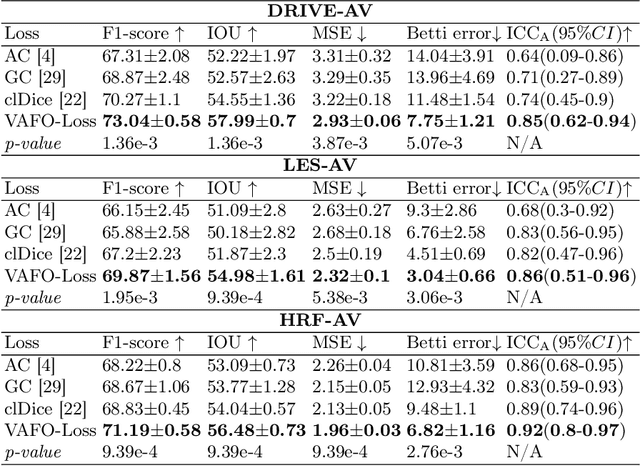
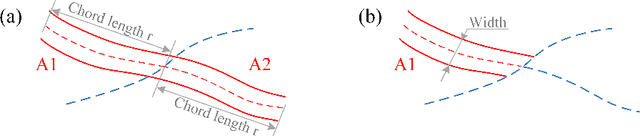
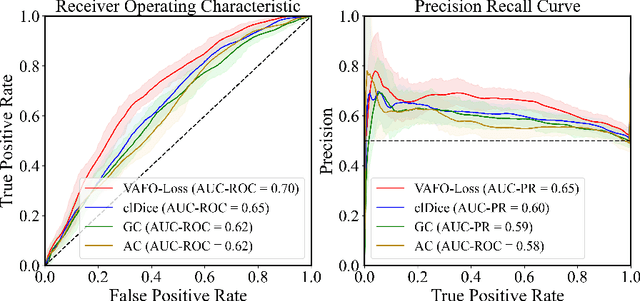
Estimating clinically-relevant vascular features following vessel segmentation is a standard pipeline for retinal vessel analysis, which provides potential ocular biomarkers for both ophthalmic disease and systemic disease. In this work, we integrate these clinical features into a novel vascular feature optimised loss function (VAFO-Loss), in order to regularise networks to produce segmentation maps, with which more accurate vascular features can be derived. Two common vascular features, vessel density and fractal dimension, are identified to be sensitive to intra-segment misclassification, which is a well-recognised problem in multi-class artery/vein segmentation particularly hindering the estimation of these vascular features. Thus we encode these two features into VAFO-Loss. We first show that incorporating our end-to-end VAFO-Loss in standard segmentation networks indeed improves vascular feature estimation, yielding quantitative improvement in stroke incidence prediction, a clinical downstream task. We also report a technically interesting finding that the trained segmentation network, albeit biased by the feature optimised loss VAFO-Loss, shows statistically significant improvement in segmentation metrics, compared to those trained with other state-of-the-art segmentation losses.
Learning to Address Intra-segment Misclassification in Retinal Imaging
Apr 25, 2021Yukun Zhou, Moucheng Xu, Yipeng Hu, Hongxiang Lin, Joseph Jacob, Pearse Keane, Daniel Alexander
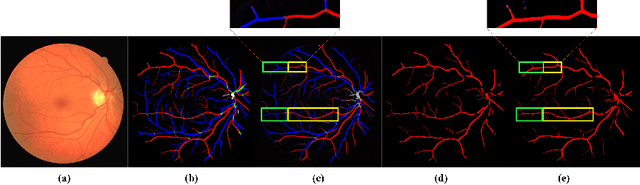
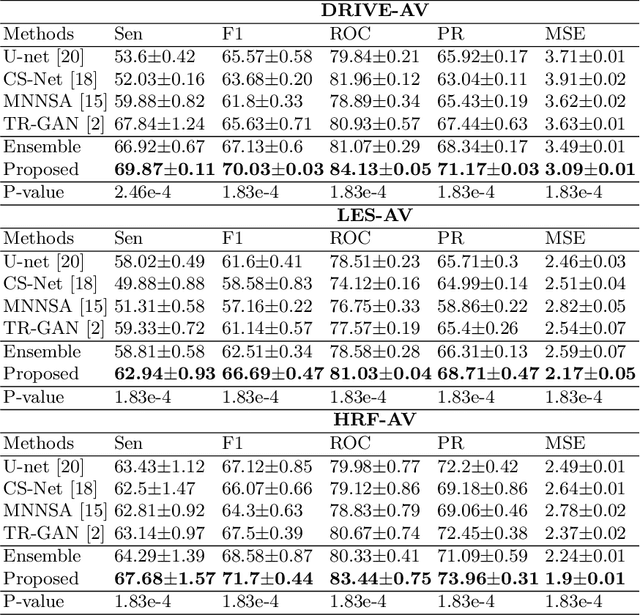
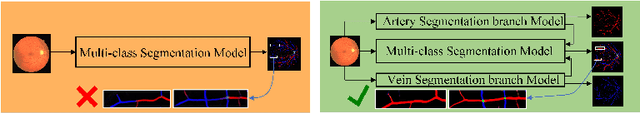
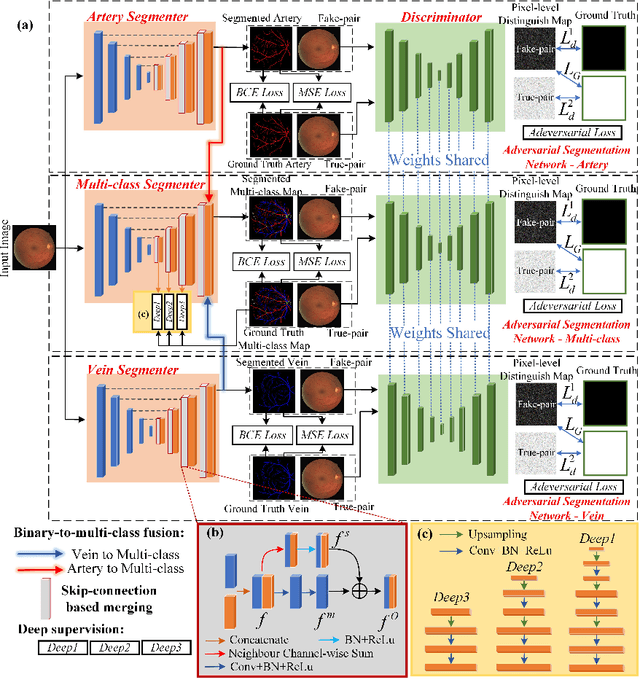
Accurate multi-class segmentation is a long-standing challenge in medical imaging, especially in scenarios where classes share strong similarity. Segmenting retinal blood vessels in retinal photographs is one such scenario, in which arteries and veins need to be identified and differentiated from each other and from the background. Intra-segment misclassification, i.e. veins classified as arteries or vice versa, frequently occurs when arteries and veins intersect, whereas in binary retinal vessel segmentation, error rates are much lower. We thus propose a new approach that decomposes multi-class segmentation into multiple binary, followed by a binary-to-multi-class fusion network. The network merges representations of artery, vein, and multi-class feature maps, each of which are supervised by expert vessel annotation in adversarial training. A skip-connection based merging process explicitly maintains class-specific gradients to avoid gradient vanishing in deep layers, to favor the discriminative features. The results show that, our model respectively improves F1-score by 4.4\%, 5.1\%, and 4.2\% compared with three state-of-the-art deep learning based methods on DRIVE-AV, LES-AV, and HRF-AV data sets.
Deep learning to estimate the physical proportion of infected region of lung for COVID-19 pneumonia with CT image set
Jun 09, 2020Wei Wu, Yu Shi, Xukun Li, Yukun Zhou, Peng Du, Shuangzhi Lv, Tingbo Liang, Jifang Sheng
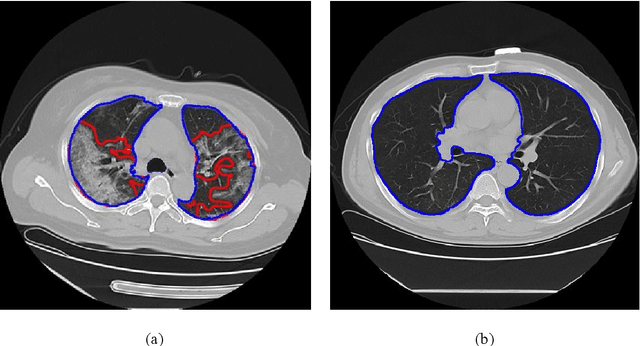
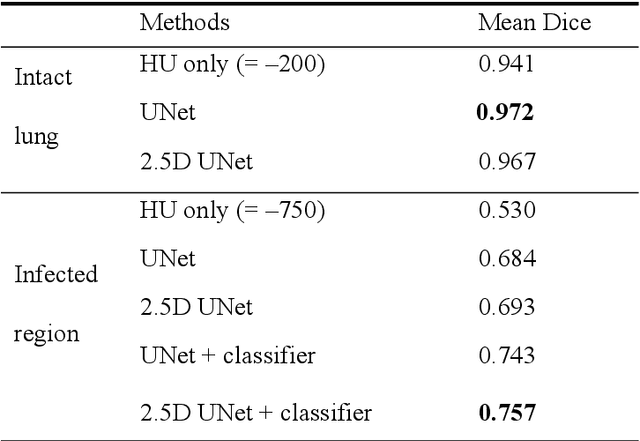
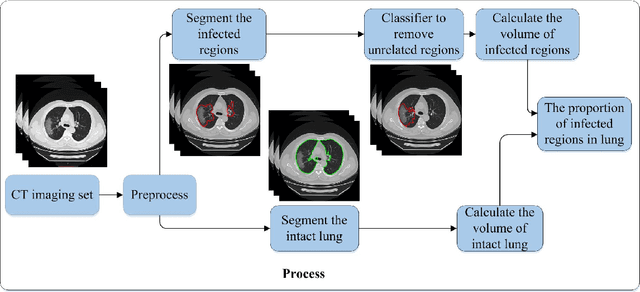
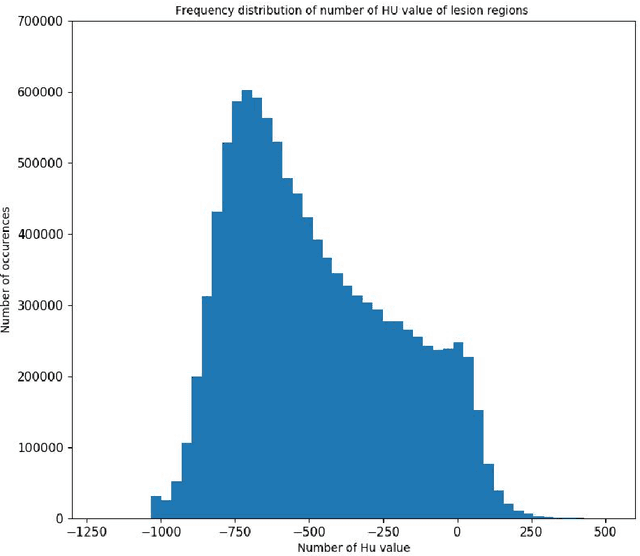
Utilizing computed tomography (CT) images to quickly estimate the severity of cases with COVID-19 is one of the most straightforward and efficacious methods. Two tasks were studied in this present paper. One was to segment the mask of intact lung in case of pneumonia. Another was to generate the masks of regions infected by COVID-19. The masks of these two parts of images then were converted to corresponding volumes to calculate the physical proportion of infected region of lung. A total of 129 CT image set were herein collected and studied. The intrinsic Hounsfiled value of CT images was firstly utilized to generate the initial dirty version of labeled masks both for intact lung and infected regions. Then, the samples were carefully adjusted and improved by two professional radiologists to generate the final training set and test benchmark. Two deep learning models were evaluated: UNet and 2.5D UNet. For the segment of infected regions, a deep learning based classifier was followed to remove unrelated blur-edged regions that were wrongly segmented out such as air tube and blood vessel tissue etc. For the segmented masks of intact lung and infected regions, the best method could achieve 0.972 and 0.757 measure in mean Dice similarity coefficient on our test benchmark. As the overall proportion of infected region of lung, the final result showed 0.961 (Pearson's correlation coefficient) and 11.7% (mean absolute percent error). The instant proportion of infected regions of lung could be used as a visual evidence to assist clinical physician to determine the severity of the case. Furthermore, a quantified report of infected regions can help predict the prognosis for COVID-19 cases which were scanned periodically within the treatment cycle.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge