RESTORE: Towards Feature Shift for Vision-Language Prompt Learning
Mar 10, 2024Yuncheng Yang, Chuyan Zhang, Zuopeng Yang, Yuting Gao, Yulei Qin, Ke Li, Xing Sun, Jie Yang, Yun Gu
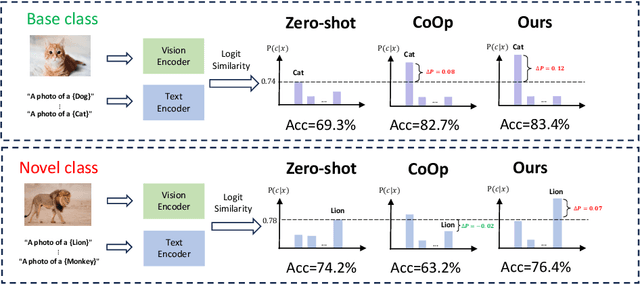
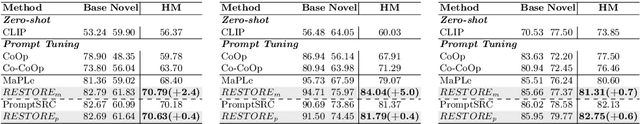
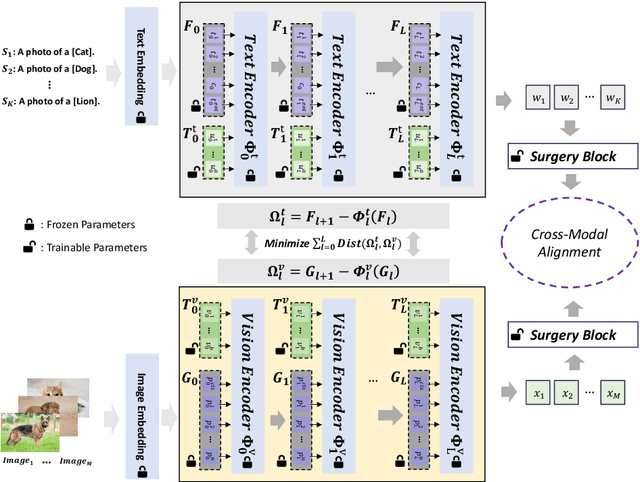
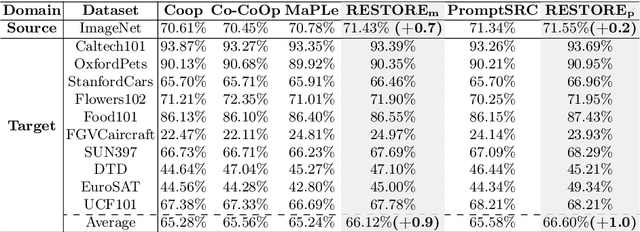
Prompt learning is effective for fine-tuning foundation models to improve their generalization across a variety of downstream tasks. However, the prompts that are independently optimized along a single modality path, may sacrifice the vision-language alignment of pre-trained models in return for improved performance on specific tasks and classes, leading to poorer generalization. In this paper, we first demonstrate that prompt tuning along only one single branch of CLIP (e.g., language or vision) is the reason why the misalignment occurs. Without proper regularization across the learnable parameters in different modalities, prompt learning violates the original pre-training constraints inherent in the two-tower architecture. To address such misalignment, we first propose feature shift, which is defined as the variation of embeddings after introducing the learned prompts, to serve as an explanatory tool. We dive into its relation with generalizability and thereafter propose RESTORE, a multi-modal prompt learning method that exerts explicit constraints on cross-modal consistency. To be more specific, to prevent feature misalignment, a feature shift consistency is introduced to synchronize inter-modal feature shifts by measuring and regularizing the magnitude of discrepancy during prompt tuning. In addition, we propose a "surgery" block to avoid short-cut hacking, where cross-modal misalignment can still be severe if the feature shift of each modality varies drastically at the same rate. It is implemented as feed-forward adapters upon both modalities to alleviate the misalignment problem. Extensive experiments on 15 datasets demonstrate that our method outperforms the state-of-the-art prompt tuning methods without compromising feature alignment.
AC-Norm: Effective Tuning for Medical Image Analysis via Affine Collaborative Normalization
Jul 28, 2023Chuyan Zhang, Yuncheng Yang, Hao Zheng, Yun Gu
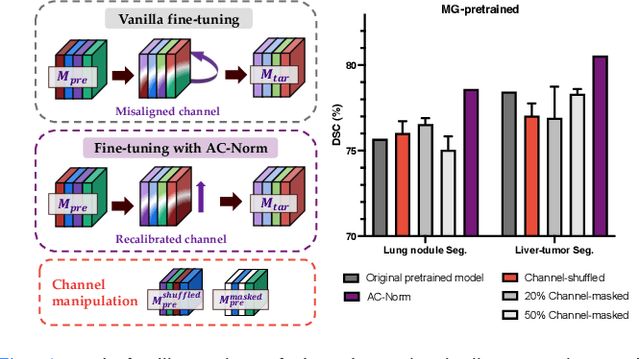
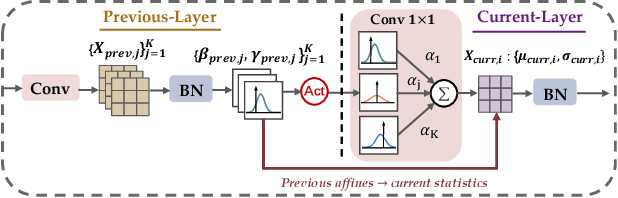
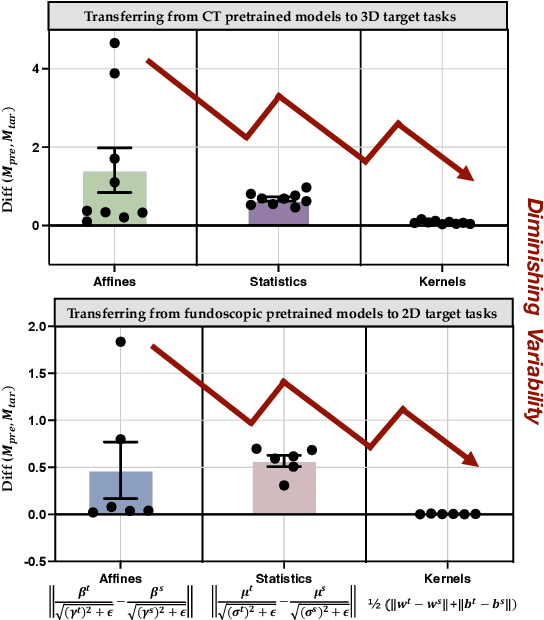
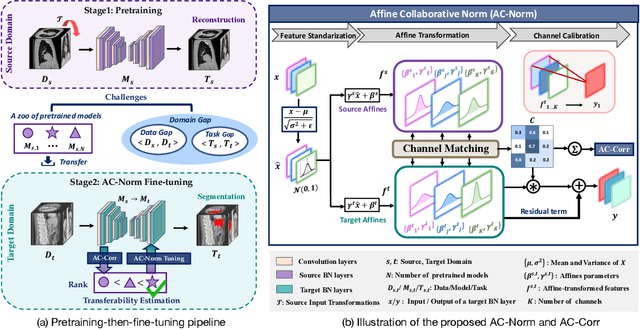
Driven by the latest trend towards self-supervised learning (SSL), the paradigm of "pretraining-then-finetuning" has been extensively explored to enhance the performance of clinical applications with limited annotations. Previous literature on model finetuning has mainly focused on regularization terms and specific policy models, while the misalignment of channels between source and target models has not received sufficient attention. In this work, we revisited the dynamics of batch normalization (BN) layers and observed that the trainable affine parameters of BN serve as sensitive indicators of domain information. Therefore, Affine Collaborative Normalization (AC-Norm) is proposed for finetuning, which dynamically recalibrates the channels in the target model according to the cross-domain channel-wise correlations without adding extra parameters. Based on a single-step backpropagation, AC-Norm can also be utilized to measure the transferability of pretrained models. We evaluated AC-Norm against the vanilla finetuning and state-of-the-art fine-tuning methods on transferring diverse pretrained models to the diabetic retinopathy grade classification, retinal vessel segmentation, CT lung nodule segmentation/classification, CT liver-tumor segmentation and MRI cardiac segmentation tasks. Extensive experiments demonstrate that AC-Norm unanimously outperforms the vanilla finetuning by up to 4% improvement, even under significant domain shifts where the state-of-the-art methods bring no gains. We also prove the capability of AC-Norm in fast transferability estimation. Our code is available at https://github.com/EndoluminalSurgicalVision-IMR/ACNorm.
Pick the Best Pre-trained Model: Towards Transferability Estimation for Medical Image Segmentation
Jul 22, 2023Yuncheng Yang, Meng Wei, Junjun He, Jie Yang, Jin Ye, Yun Gu
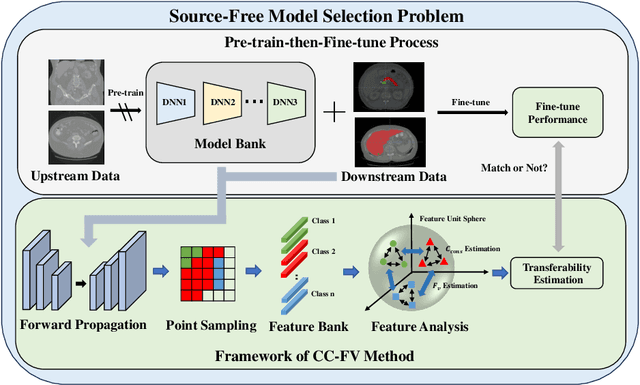
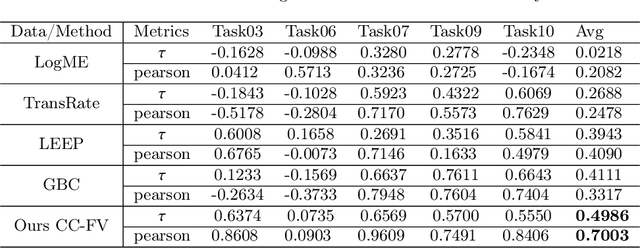
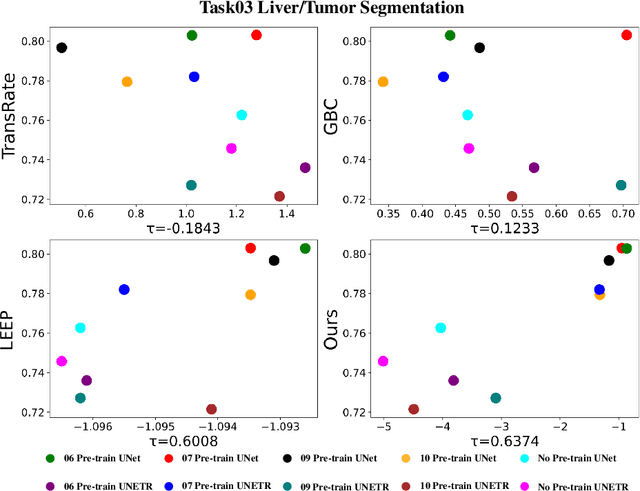
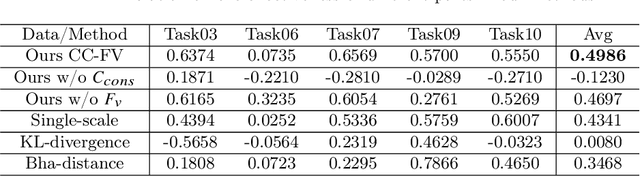
Transfer learning is a critical technique in training deep neural networks for the challenging medical image segmentation task that requires enormous resources. With the abundance of medical image data, many research institutions release models trained on various datasets that can form a huge pool of candidate source models to choose from. Hence, it's vital to estimate the source models' transferability (i.e., the ability to generalize across different downstream tasks) for proper and efficient model reuse. To make up for its deficiency when applying transfer learning to medical image segmentation, in this paper, we therefore propose a new Transferability Estimation (TE) method. We first analyze the drawbacks of using the existing TE algorithms for medical image segmentation and then design a source-free TE framework that considers both class consistency and feature variety for better estimation. Extensive experiments show that our method surpasses all current algorithms for transferability estimation in medical image segmentation. Code is available at https://github.com/EndoluminalSurgicalVision-IMR/CCFV
Exploring Vanilla U-Net for Lesion Segmentation from Whole-body FDG-PET/CT Scans
Oct 14, 2022Jin Ye, Haoyu Wang, Ziyan Huang, Zhongying Deng, Yanzhou Su, Can Tu, Qian Wu, Yuncheng Yang, Meng Wei, Jingqi Niu, Junjun He
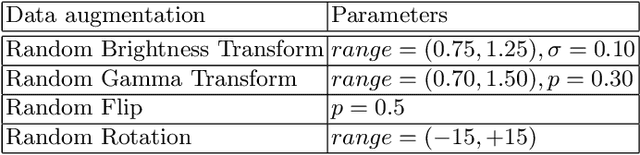
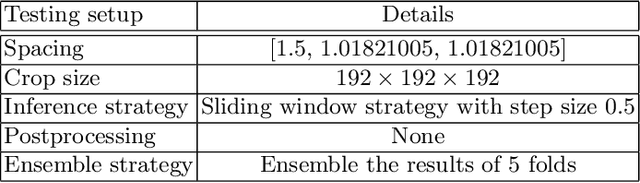


Tumor lesion segmentation is one of the most important tasks in medical image analysis. In clinical practice, Fluorodeoxyglucose Positron-Emission Tomography~(FDG-PET) is a widely used technique to identify and quantify metabolically active tumors. However, since FDG-PET scans only provide metabolic information, healthy tissue or benign disease with irregular glucose consumption may be mistaken for cancer. To handle this challenge, PET is commonly combined with Computed Tomography~(CT), with the CT used to obtain the anatomic structure of the patient. The combination of PET-based metabolic and CT-based anatomic information can contribute to better tumor segmentation results. %Computed tomography~(CT) is a popular modality to illustrate the anatomic structure of the patient. The combination of PET and CT is promising to handle this challenge by utilizing metabolic and anatomic information. In this paper, we explore the potential of U-Net for lesion segmentation in whole-body FDG-PET/CT scans from three aspects, including network architecture, data preprocessing, and data augmentation. The experimental results demonstrate that the vanilla U-Net with proper input shape can achieve satisfactory performance. Specifically, our method achieves first place in both preliminary and final leaderboards of the autoPET 2022 challenge. Our code is available at https://github.com/Yejin0111/autoPET2022_Blackbean.
An evaluation of U-Net in Renal Structure Segmentation
Sep 06, 2022Haoyu Wang, Ziyan Huang, Jin Ye, Can Tu, Yuncheng Yang, Shiyi Du, Zhongying Deng, Chenglong Ma, Jingqi Niu, Junjun He
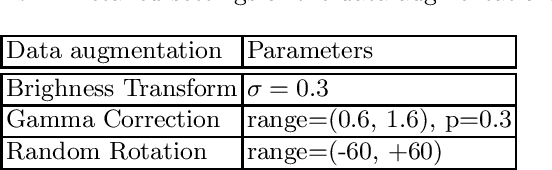
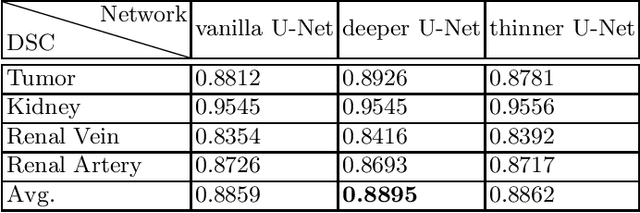
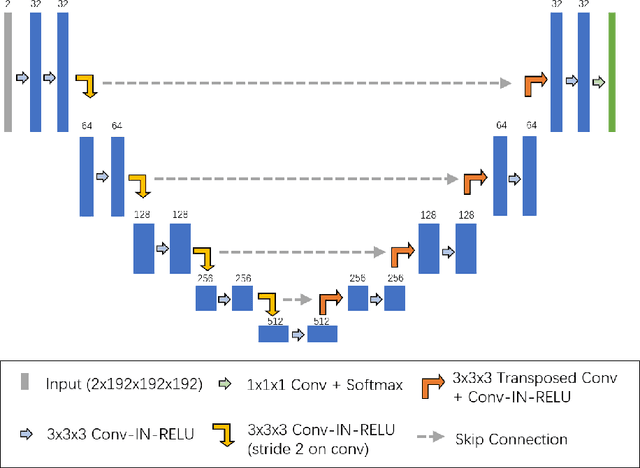
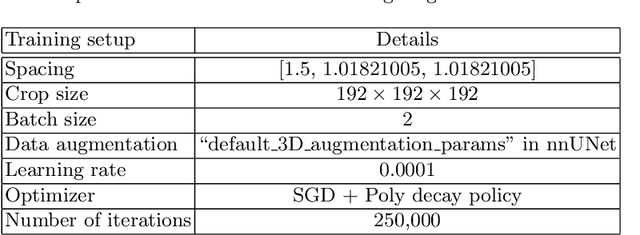
Renal structure segmentation from computed tomography angiography~(CTA) is essential for many computer-assisted renal cancer treatment applications. Kidney PArsing~(KiPA 2022) Challenge aims to build a fine-grained multi-structure dataset and improve the segmentation of multiple renal structures. Recently, U-Net has dominated the medical image segmentation. In the KiPA challenge, we evaluated several U-Net variants and selected the best models for the final submission.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge