COMBO: Compositional World Models for Embodied Multi-Agent Cooperation
Apr 16, 2024Hongxin Zhang, Zeyuan Wang, Qiushi Lyu, Zheyuan Zhang, Sunli Chen, Tianmin Shu, Yilun Du, Chuang Gan
In this paper, we investigate the problem of embodied multi-agent cooperation, where decentralized agents must cooperate given only partial egocentric views of the world. To effectively plan in this setting, in contrast to learning world dynamics in a single-agent scenario, we must simulate world dynamics conditioned on an arbitrary number of agents' actions given only partial egocentric visual observations of the world. To address this issue of partial observability, we first train generative models to estimate the overall world state given partial egocentric observations. To enable accurate simulation of multiple sets of actions on this world state, we then propose to learn a compositional world model for multi-agent cooperation by factorizing the naturally composable joint actions of multiple agents and compositionally generating the video. By leveraging this compositional world model, in combination with Vision Language Models to infer the actions of other agents, we can use a tree search procedure to integrate these modules and facilitate online cooperative planning. To evaluate the efficacy of our methods, we create two challenging embodied multi-agent long-horizon cooperation tasks using the ThreeDWorld simulator and conduct experiments with 2-4 agents. The results show our compositional world model is effective and the framework enables the embodied agents to cooperate efficiently with different agents across various tasks and an arbitrary number of agents, showing the promising future of our proposed framework. More videos can be found at https://vis-www.cs.umass.edu/combo/.
Scientific Large Language Models: A Survey on Biological & Chemical Domains
Jan 26, 2024Qiang Zhang, Keyang Ding, Tianwen Lyv, Xinda Wang, Qingyu Yin, Yiwen Zhang, Jing Yu, Yuhao Wang, Xiaotong Li, Zhuoyi Xiang, Xiang Zhuang, Zeyuan Wang, Ming Qin, Mengyao Zhang, Jinlu Zhang, Jiyu Cui, Renjun Xu, Hongyang Chen, Xiaohui Fan, Huabin Xing, Huajun Chen
Large Language Models (LLMs) have emerged as a transformative power in enhancing natural language comprehension, representing a significant stride toward artificial general intelligence. The application of LLMs extends beyond conventional linguistic boundaries, encompassing specialized linguistic systems developed within various scientific disciplines. This growing interest has led to the advent of scientific LLMs, a novel subclass specifically engineered for facilitating scientific discovery. As a burgeoning area in the community of AI for Science, scientific LLMs warrant comprehensive exploration. However, a systematic and up-to-date survey introducing them is currently lacking. In this paper, we endeavor to methodically delineate the concept of "scientific language", whilst providing a thorough review of the latest advancements in scientific LLMs. Given the expansive realm of scientific disciplines, our analysis adopts a focused lens, concentrating on the biological and chemical domains. This includes an in-depth examination of LLMs for textual knowledge, small molecules, macromolecular proteins, genomic sequences, and their combinations, analyzing them in terms of model architectures, capabilities, datasets, and evaluation. Finally, we critically examine the prevailing challenges and point out promising research directions along with the advances of LLMs. By offering a comprehensive overview of technical developments in this field, this survey aspires to be an invaluable resource for researchers navigating the intricate landscape of scientific LLMs.
InstructProtein: Aligning Human and Protein Language via Knowledge Instruction
Oct 05, 2023Zeyuan Wang, Qiang Zhang, Keyan Ding, Ming Qin, Xiang Zhuang, Xiaotong Li, Huajun Chen
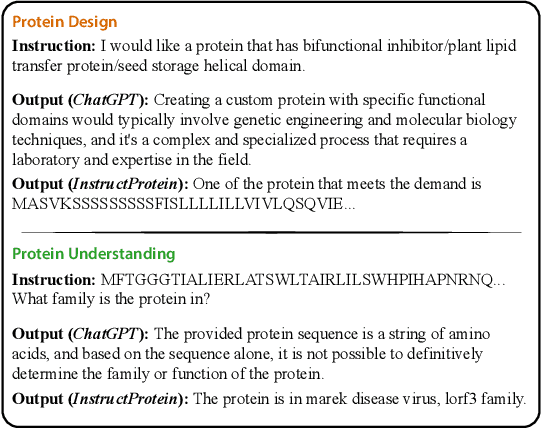

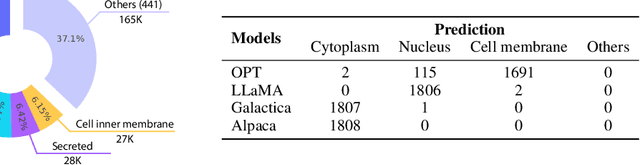
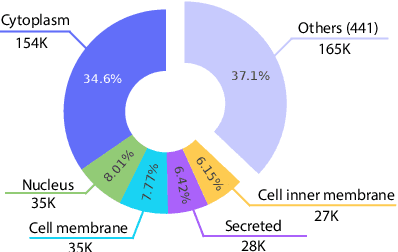
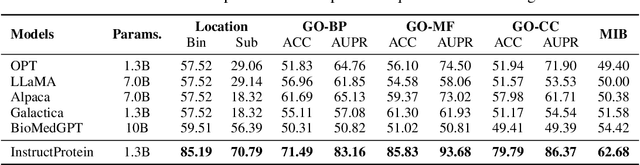
Large Language Models (LLMs) have revolutionized the field of natural language processing, but they fall short in comprehending biological sequences such as proteins. To address this challenge, we propose InstructProtein, an innovative LLM that possesses bidirectional generation capabilities in both human and protein languages: (i) taking a protein sequence as input to predict its textual function description and (ii) using natural language to prompt protein sequence generation. To achieve this, we first pre-train an LLM on both protein and natural language corpora, enabling it to comprehend individual languages. Then supervised instruction tuning is employed to facilitate the alignment of these two distinct languages. Herein, we introduce a knowledge graph-based instruction generation framework to construct a high-quality instruction dataset, addressing annotation imbalance and instruction deficits in existing protein-text corpus. In particular, the instructions inherit the structural relations between proteins and function annotations in knowledge graphs, which empowers our model to engage in the causal modeling of protein functions, akin to the chain-of-thought processes in natural languages. Extensive experiments on bidirectional protein-text generation tasks show that InstructProtein outperforms state-of-the-art LLMs by large margins. Moreover, InstructProtein serves as a pioneering step towards text-based protein function prediction and sequence design, effectively bridging the gap between protein and human language understanding.
Prompt-Guided Injection of Conformation to Pre-trained Protein Model
Feb 07, 2022Qiang Zhang, Zeyuan Wang, Yuqiang Han, Haoran Yu, Xurui Jin, Huajun Chen
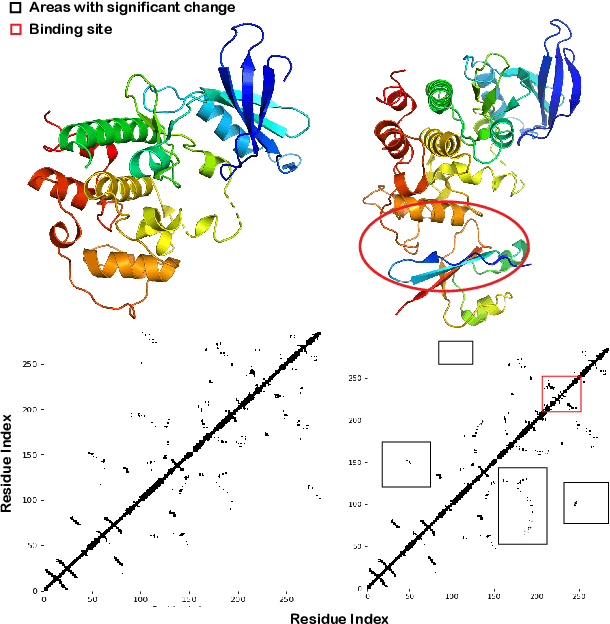
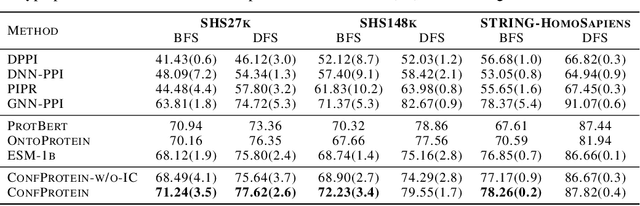
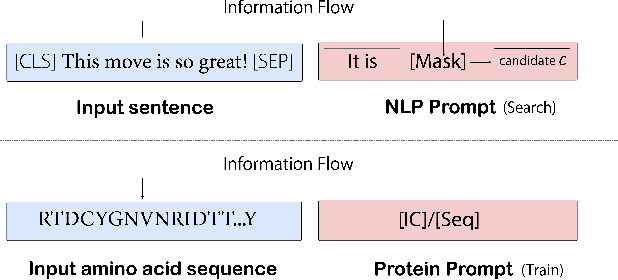

Pre-trained protein models (PTPMs) represent a protein with one fixed embedding and thus are not capable for diverse tasks. For example, protein structures can shift, namely protein folding, between several conformations in various biological processes. To enable PTPMs to produce task-aware representations, we propose to learn interpretable, pluggable and extensible protein prompts as a way of injecting task-related knowledge into PTPMs. In this regard, prior PTPM optimization with the masked language modeling task can be interpreted as learning a sequence prompt (Seq prompt) that enables PTPMs to capture the sequential dependency between amino acids. To incorporate conformational knowledge to PTPMs, we propose an interaction-conformation prompt (IC prompt) that is learned through back-propagation with the protein-protein interaction task. As an instantiation, we present a conformation-aware pre-trained protein model that learns both sequence and interaction-conformation prompts in a multi-task setting. We conduct comprehensive experiments on nine protein datasets. Results confirm our expectation that using the sequence prompt does not hurt PTPMs' performance on sequence-related tasks while incorporating the interaction-conformation prompt significantly improves PTPMs' performance on tasks where conformational knowledge counts. We also show the learned prompts can be combined and extended to deal with new complex tasks.
Reinforcement Learning Based Sparse Black-box Adversarial Attack on Video Recognition Models
Aug 29, 2021Zeyuan Wang, Chaofeng Sha, Su Yang


We explore the black-box adversarial attack on video recognition models. Attacks are only performed on selected key regions and key frames to reduce the high computation cost of searching adversarial perturbations on a video due to its high dimensionality. To select key frames, one way is to use heuristic algorithms to evaluate the importance of each frame and choose the essential ones. However, it is time inefficient on sorting and searching. In order to speed up the attack process, we propose a reinforcement learning based frame selection strategy. Specifically, the agent explores the difference between the original class and the target class of videos to make selection decisions. It receives rewards from threat models which indicate the quality of the decisions. Besides, we also use saliency detection to select key regions and only estimate the sign of gradient instead of the gradient itself in zeroth order optimization to further boost the attack process. We can use the trained model directly in the untargeted attack or with little fine-tune in the targeted attack, which saves computation time. A range of empirical results on real datasets demonstrate the effectiveness and efficiency of the proposed method.
* Accepted as a conference paper of IJCAI-21 (the 30th International Joint Conference on Artificial Intelligence)
Learning Order Parameters from Videos of Dynamical Phases for Skyrmions with Neural Networks
Dec 02, 2020Weidi Wang, Zeyuan Wang, Yinghui Zhang, Bo Sun, Ke Xia




The ability to recognize dynamical phenomena (e.g., dynamical phases) and dynamical processes in physical events from videos, then to abstract physical concepts and reveal physical laws, lies at the core of human intelligence. The main purposes of this paper are to use neural networks for classifying the dynamical phases of some videos and to demonstrate that neural networks can learn physical concepts from them. To this end, we employ multiple neural networks to recognize the static phases (image format) and dynamical phases (video format) of a particle-based skyrmion model. Our results show that neural networks, without any prior knowledge, can not only correctly classify these phases, but also predict the phase boundaries which agree with those obtained by simulation. We further propose a parameter visualization scheme to interpret what neural networks have learned. We show that neural networks can learn two order parameters from videos of dynamical phases and predict the critical values of two order parameters. Finally, we demonstrate that only two order parameters are needed to identify videos of skyrmion dynamical phases. It shows that this parameter visualization scheme can be used to determine how many order parameters are needed to fully recognize the input phases. Our work sheds light on the future use of neural networks in discovering new physical concepts and revealing unknown yet physical laws from videos.
Cooperative Bi-path Metric for Few-shot Learning
Aug 10, 2020Zeyuan Wang, Yifan Zhao, Jia Li, Yonghong Tian




Given base classes with sufficient labeled samples, the target of few-shot classification is to recognize unlabeled samples of novel classes with only a few labeled samples. Most existing methods only pay attention to the relationship between labeled and unlabeled samples of novel classes, which do not make full use of information within base classes. In this paper, we make two contributions to investigate the few-shot classification problem. First, we report a simple and effective baseline trained on base classes in the way of traditional supervised learning, which can achieve comparable results to the state of the art. Second, based on the baseline, we propose a cooperative bi-path metric for classification, which leverages the correlations between base classes and novel classes to further improve the accuracy. Experiments on two widely used benchmarks show that our method is a simple and effective framework, and a new state of the art is established in the few-shot classification field.
AMI-Net+: A Novel Multi-Instance Neural Network for Medical Diagnosis from Incomplete and Imbalanced Data
Jul 03, 2019Zeyuan Wang, Josiah Poon, Simon Poon




In medical real-world study (RWS), how to fully utilize the fragmentary and scarce information in model training to generate the solid diagnosis results is a challenging task. In this work, we introduce a novel multi-instance neural network, AMI-Net+, to train and predict from the incomplete and extremely imbalanced data. It is more effective than the state-of-art method, AMI-Net. First, we also implement embedding, multi-head attention and gated attention-based multi-instance pooling to capture the relations of symptoms themselves and with the given disease. Besides, we propose var-ious improvements to AMI-Net, that the cross-entropy loss is replaced by focal loss and we propose a novel self-adaptive multi-instance pooling method on instance-level to obtain the bag representation. We validate the performance of AMI-Net+ on two real-world datasets, from two different medical domains. Results show that our approach outperforms other base-line models by a considerable margin.
Attention-based Multi-instance Neural Network for Medical Diagnosis from Incomplete and Low Quality Data
Apr 09, 2019Zeyuan Wang, Josiah Poon, Shiding Sun, Simon Poon




One way to extract patterns from clinical records is to consider each patient record as a bag with various number of instances in the form of symptoms. Medical diagnosis is to discover informative ones first and then map them to one or more diseases. In many cases, patients are represented as vectors in some feature space and a classifier is applied after to generate diagnosis results. However, in many real-world cases, data is often of low-quality due to a variety of reasons, such as data consistency, integrity, completeness, accuracy, etc. In this paper, we propose a novel approach, attention based multi-instance neural network (AMI-Net), to make the single disease classification only based on the existing and valid information in the real-world outpatient records. In the context of a patient, it takes a bag of instances as input and output the bag label directly in end-to-end way. Embedding layer is adopted at the beginning, mapping instances into an embedding space which represents the individual patient condition. The correlations among instances and their importance for the final classification are captured by multi-head attention transformer, instance-level multi-instance pooling and bag-level multi-instance pooling. The proposed approach was test on two non-standardized and highly imbalanced datasets, one in the Traditional Chinese Medicine (TCM) domain and the other in the Western Medicine (WM) domain. Our preliminary results show that the proposed approach outperforms all baselines results by a significant margin.
CNN based Multi-Instance Multi-Task Learning for Syndrome Differentiation of Diabetic Patients
Jan 19, 2019Zeyuan Wang, Josiah Poon, Shiding Sun, Simon Poon




Syndrome differentiation in Traditional Chinese Medicine (TCM) is the process of understanding and reasoning body condition, which is the essential step and premise of effective treatments. However, due to its complexity and lack of standardization, it is challenging to achieve. In this study, we consider each patient's record as a one-dimensional image and symptoms as pixels, in which missing and negative values are represented by zero pixels. The objective is to find relevant symptoms first and then map them to proper syndromes, that is similar to the object detection problem in computer vision. Inspired from it, we employ multi-instance multi-task learning combined with the convolutional neural network (MIMT-CNN) for syndrome differentiation, which takes region proposals as input and output image labels directly. The neural network consists of region proposals generation, convolutional layer, fully connected layer, and max pooling (multi-instance pooling) layer followed by the sigmoid function in each syndrome prediction task for image representation learning and final results generation. On the diabetes dataset, it performs better than all other baseline methods. Moreover, it shows stability and reliability to generate results, even on the dataset with small sample size, a large number of missing values and noises.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge