Scale-aware competition network for palmprint recognition
Nov 21, 2023Chengrui Gao, Ziyuan Yang, Min Zhu, Andrew Beng Jin Teoh
Palmprint biometrics garner heightened attention in palm-scanning payment and social security due to their distinctive attributes. However, prevailing methodologies singularly prioritize texture orientation, neglecting the significant texture scale dimension. We design an innovative network for concurrently extracting intra-scale and inter-scale features to redress this limitation. This paper proposes a scale-aware competitive network (SAC-Net), which includes the Inner-Scale Competition Module (ISCM) and the Across-Scale Competition Module (ASCM) to capture texture characteristics related to orientation and scale. ISCM efficiently integrates learnable Gabor filters and a self-attention mechanism to extract rich orientation data and discern textures with long-range discriminative properties. Subsequently, ASCM leverages a competitive strategy across various scales to effectively encapsulate the competitive texture scale elements. By synergizing ISCM and ASCM, our method adeptly characterizes palmprint features. Rigorous experimentation across three benchmark datasets unequivocally demonstrates our proposed approach's exceptional recognition performance and resilience relative to state-of-the-art alternatives.
SA-Med2D-20M Dataset: Segment Anything in 2D Medical Imaging with 20 Million masks
Nov 20, 2023Jin Ye, Junlong Cheng, Jianpin Chen, Zhongying Deng, Tianbin Li, Haoyu Wang, Yanzhou Su, Ziyan Huang, Jilong Chen, Lei Jiang, Hui Sun, Min Zhu, Shaoting Zhang, Junjun He, Yu Qiao
Segment Anything Model (SAM) has achieved impressive results for natural image segmentation with input prompts such as points and bounding boxes. Its success largely owes to massive labeled training data. However, directly applying SAM to medical image segmentation cannot perform well because SAM lacks medical knowledge -- it does not use medical images for training. To incorporate medical knowledge into SAM, we introduce SA-Med2D-20M, a large-scale segmentation dataset of 2D medical images built upon numerous public and private datasets. It consists of 4.6 million 2D medical images and 19.7 million corresponding masks, covering almost the whole body and showing significant diversity. This paper describes all the datasets collected in SA-Med2D-20M and details how to process these datasets. Furthermore, comprehensive statistics of SA-Med2D-20M are presented to facilitate the better use of our dataset, which can help the researchers build medical vision foundation models or apply their models to downstream medical applications. We hope that the large scale and diversity of SA-Med2D-20M can be leveraged to develop medical artificial intelligence for enhancing diagnosis, medical image analysis, knowledge sharing, and education. The data with the redistribution license is publicly available at https://github.com/OpenGVLab/SAM-Med2D.
SAM-Med2D
Aug 30, 2023Junlong Cheng, Jin Ye, Zhongying Deng, Jianpin Chen, Tianbin Li, Haoyu Wang, Yanzhou Su, Ziyan Huang, Jilong Chen, Lei Jiang, Hui Sun, Junjun He, Shaoting Zhang, Min Zhu, Yu Qiao


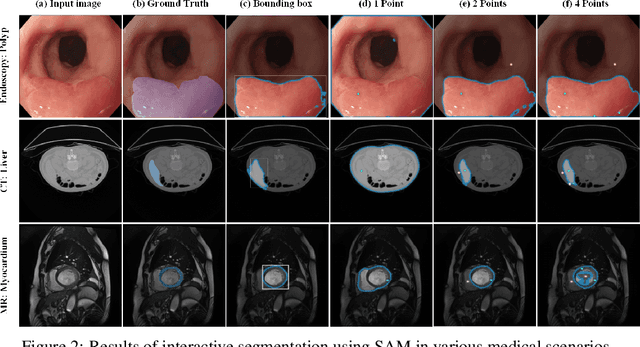
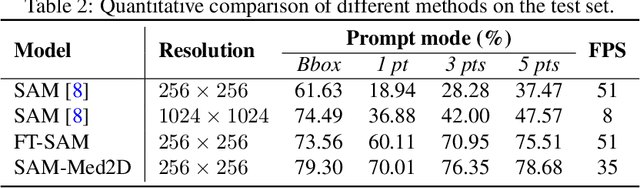
The Segment Anything Model (SAM) represents a state-of-the-art research advancement in natural image segmentation, achieving impressive results with input prompts such as points and bounding boxes. However, our evaluation and recent research indicate that directly applying the pretrained SAM to medical image segmentation does not yield satisfactory performance. This limitation primarily arises from significant domain gap between natural images and medical images. To bridge this gap, we introduce SAM-Med2D, the most comprehensive studies on applying SAM to medical 2D images. Specifically, we first collect and curate approximately 4.6M images and 19.7M masks from public and private datasets, constructing a large-scale medical image segmentation dataset encompassing various modalities and objects. Then, we comprehensively fine-tune SAM on this dataset and turn it into SAM-Med2D. Unlike previous methods that only adopt bounding box or point prompts as interactive segmentation approach, we adapt SAM to medical image segmentation through more comprehensive prompts involving bounding boxes, points, and masks. We additionally fine-tune the encoder and decoder of the original SAM to obtain a well-performed SAM-Med2D, leading to the most comprehensive fine-tuning strategies to date. Finally, we conducted a comprehensive evaluation and analysis to investigate the performance of SAM-Med2D in medical image segmentation across various modalities, anatomical structures, and organs. Concurrently, we validated the generalization capability of SAM-Med2D on 9 datasets from MICCAI 2023 challenge. Overall, our approach demonstrated significantly superior performance and generalization capability compared to SAM.
VideoPro: A Visual Analytics Approach for Interactive Video Programming
Aug 01, 2023Jianben He, Xingbo Wang, Kam Kwai Wong, Xijie Huang, Changjian Chen, Zixin Chen, Fengjie Wang, Min Zhu, Huamin Qu
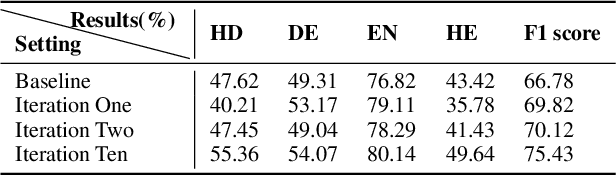
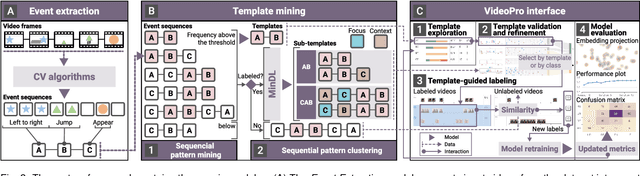
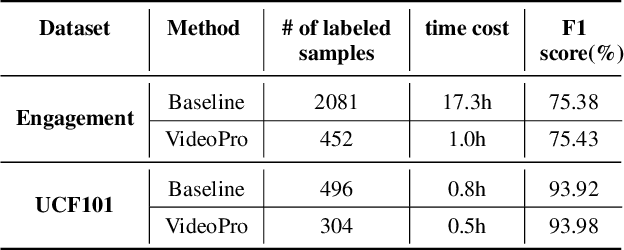
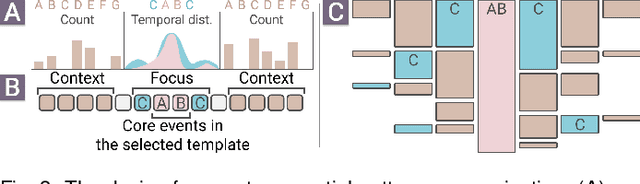
Constructing supervised machine learning models for real-world video analysis require substantial labeled data, which is costly to acquire due to scarce domain expertise and laborious manual inspection. While data programming shows promise in generating labeled data at scale with user-defined labeling functions, the high dimensional and complex temporal information in videos poses additional challenges for effectively composing and evaluating labeling functions. In this paper, we propose VideoPro, a visual analytics approach to support flexible and scalable video data programming for model steering with reduced human effort. We first extract human-understandable events from videos using computer vision techniques and treat them as atomic components of labeling functions. We further propose a two-stage template mining algorithm that characterizes the sequential patterns of these events to serve as labeling function templates for efficient data labeling. The visual interface of VideoPro facilitates multifaceted exploration, examination, and application of the labeling templates, allowing for effective programming of video data at scale. Moreover, users can monitor the impact of programming on model performance and make informed adjustments during the iterative programming process. We demonstrate the efficiency and effectiveness of our approach with two case studies and expert interviews.
SegNetr: Rethinking the local-global interactions and skip connections in U-shaped networks
Jul 21, 2023Junlong Cheng, Chengrui Gao, Fengjie Wang, Min Zhu
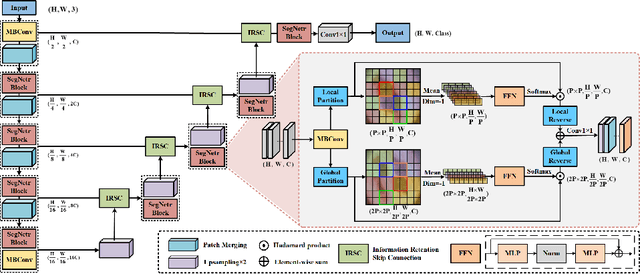
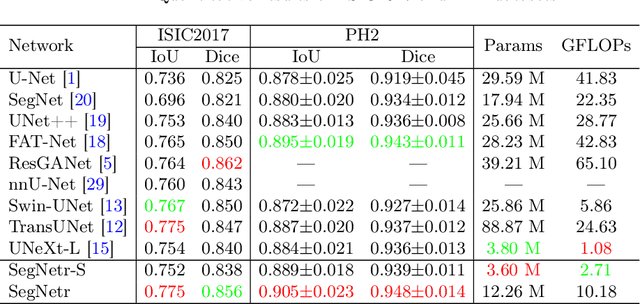


Recently, U-shaped networks have dominated the field of medical image segmentation due to their simple and easily tuned structure. However, existing U-shaped segmentation networks: 1) mostly focus on designing complex self-attention modules to compensate for the lack of long-term dependence based on convolution operation, which increases the overall number of parameters and computational complexity of the network; 2) simply fuse the features of encoder and decoder, ignoring the connection between their spatial locations. In this paper, we rethink the above problem and build a lightweight medical image segmentation network, called SegNetr. Specifically, we introduce a novel SegNetr block that can perform local-global interactions dynamically at any stage and with only linear complexity. At the same time, we design a general information retention skip connection (IRSC) to preserve the spatial location information of encoder features and achieve accurate fusion with the decoder features. We validate the effectiveness of SegNetr on four mainstream medical image segmentation datasets, with 59\% and 76\% fewer parameters and GFLOPs than vanilla U-Net, while achieving segmentation performance comparable to state-of-the-art methods. Notably, the components proposed in this paper can be applied to other U-shaped networks to improve their segmentation performance.
Accurate 3D Prediction of Missing Teeth in Diverse Patterns for Precise Dental Implant Planning
Jul 16, 2023Lei Ma, Peng Xue, Yuning Gu, Yue Zhao, Min Zhu, Zhongxiang Ding, Dinggang Shen



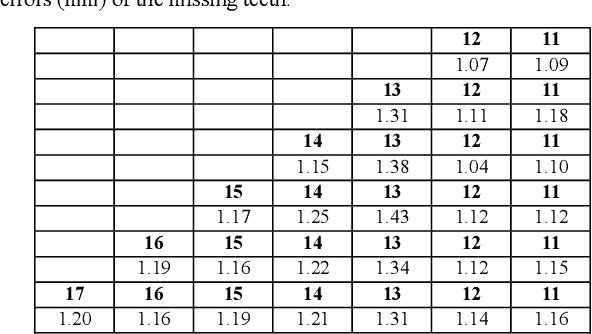
In recent years, the demand for dental implants has surged, driven by their high success rates and esthetic advantages. However, accurate prediction of missing teeth for precise digital implant planning remains a challenge due to the intricate nature of dental structures and the variability in tooth loss patterns. This study presents a novel framework for accurate prediction of missing teeth in different patterns, facilitating digital implant planning. The proposed framework begins by estimating point-to-point correspondence among a dataset of dental mesh models reconstructed from CBCT images of healthy subjects. Subsequently, tooth dictionaries are constructed for each tooth type, encoding their position and shape information based on the established point-to-point correspondence. To predict missing teeth in a given dental mesh model, sparse coefficients are learned by sparsely representing adjacent teeth of the missing teeth using the corresponding tooth dictionaries. These coefficients are then applied to the dictionaries of the missing teeth to generate accurate predictions of their positions and shapes. The evaluation results on real subjects shows that our proposed framework achieves an average prediction error of 1.04mm for predictions of single missing tooth and an average prediction error of 1.33mm for the prediction of 14 missing teeth, which demonstrates its capability of accurately predicting missing teeth in various patterns. By accurately predicting missing teeth, dental professionals can improve the planning and placement of dental implants, leading to better esthetic and functional outcomes for patients undergoing dental implant procedures.
$\mathbf{\mathbb{E}^{FWI}}$: Multi-parameter Benchmark Datasets for Elastic Full Waveform Inversion of Geophysical Properties
Jun 21, 2023Shihang Feng, Hanchen Wang, Chengyuan Deng, Yinan Feng, Yanhua Liu, Min Zhu, Peng Jin, Yinpeng Chen, Youzuo Lin
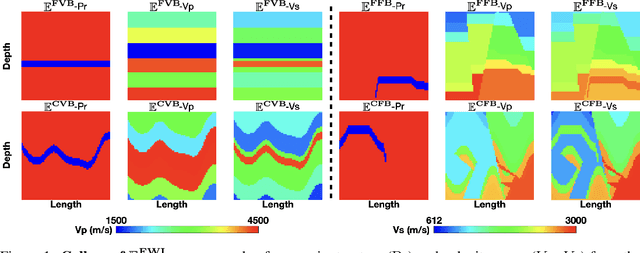
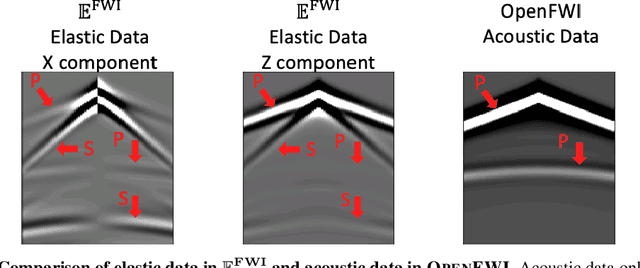
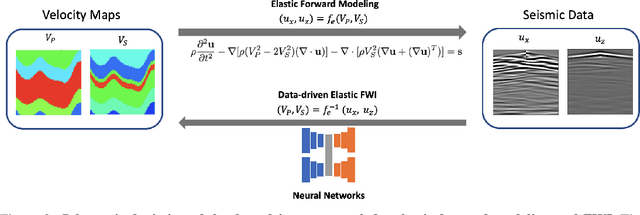
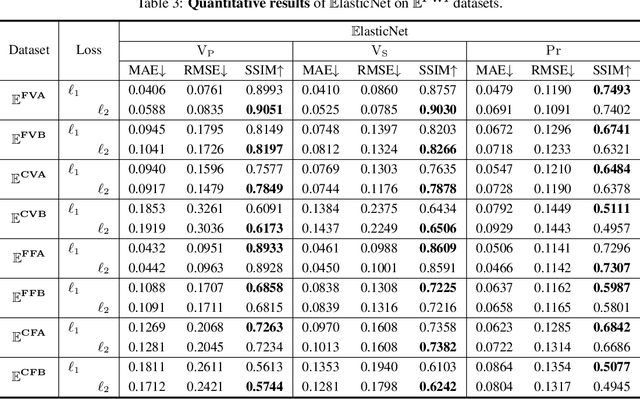
Elastic geophysical properties (such as P- and S-wave velocities) are of great importance to various subsurface applications like CO$_2$ sequestration and energy exploration (e.g., hydrogen and geothermal). Elastic full waveform inversion (FWI) is widely applied for characterizing reservoir properties. In this paper, we introduce $\mathbf{\mathbb{E}^{FWI}}$, a comprehensive benchmark dataset that is specifically designed for elastic FWI. $\mathbf{\mathbb{E}^{FWI}}$ encompasses 8 distinct datasets that cover diverse subsurface geologic structures (flat, curve, faults, etc). The benchmark results produced by three different deep learning methods are provided. In contrast to our previously presented dataset (pressure recordings) for acoustic FWI (referred to as OpenFWI), the seismic dataset in $\mathbf{\mathbb{E}^{FWI}}$ has both vertical and horizontal components. Moreover, the velocity maps in $\mathbf{\mathbb{E}^{FWI}}$ incorporate both P- and S-wave velocities. While the multicomponent data and the added S-wave velocity make the data more realistic, more challenges are introduced regarding the convergence and computational cost of the inversion. We conduct comprehensive numerical experiments to explore the relationship between P-wave and S-wave velocities in seismic data. The relation between P- and S-wave velocities provides crucial insights into the subsurface properties such as lithology, porosity, fluid content, etc. We anticipate that $\mathbf{\mathbb{E}^{FWI}}$ will facilitate future research on multiparameter inversions and stimulate endeavors in several critical research topics of carbon-zero and new energy exploration. All datasets, codes and relevant information can be accessed through our website at https://efwi-lanl.github.io/
Fourier-DeepONet: Fourier-enhanced deep operator networks for full waveform inversion with improved accuracy, generalizability, and robustness
May 26, 2023Min Zhu, Shihang Feng, Youzuo Lin, Lu Lu
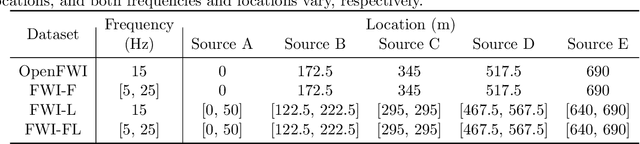
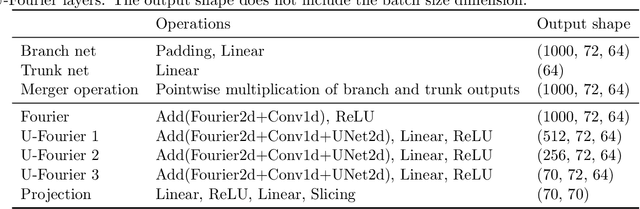
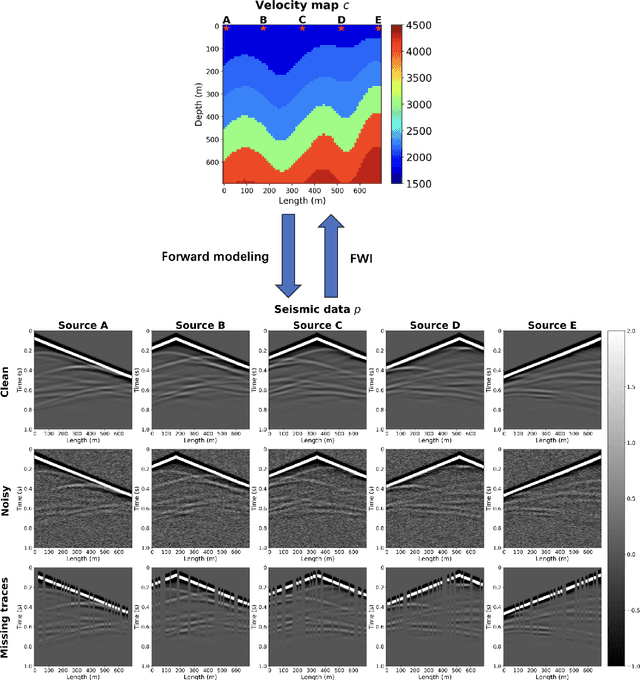

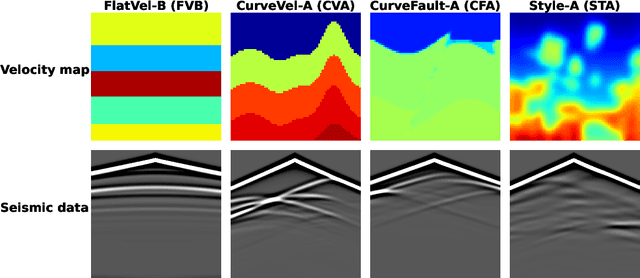

Full waveform inversion (FWI) infers the subsurface structure information from seismic waveform data by solving a non-convex optimization problem. Data-driven FWI has been increasingly studied with various neural network architectures to improve accuracy and computational efficiency. Nevertheless, the applicability of pre-trained neural networks is severely restricted by potential discrepancies between the source function used in the field survey and the one utilized during training. Here, we develop a Fourier-enhanced deep operator network (Fourier-DeepONet) for FWI with the generalization of seismic sources, including the frequencies and locations of sources. Specifically, we employ the Fourier neural operator as the decoder of DeepONet, and we utilize source parameters as one input of Fourier-DeepONet, facilitating the resolution of FWI with variable sources. To test Fourier-DeepONet, we develop two new and realistic FWI benchmark datasets (FWI-F and FWI-L) with varying source frequencies and locations. Our experiments demonstrate that compared with existing data-driven FWI methods, Fourier-DeepONet obtains more accurate predictions of subsurface structures in a wide range of source parameters. Moreover, the proposed Fourier-DeepONet exhibits superior robustness when dealing with noisy inputs or inputs with missing traces, paving the way for more reliable and accurate subsurface imaging across diverse real conditions.
Construction of unbiased dental template and parametric dental model for precision digital dentistry
Apr 07, 2023Lei Ma, Jingyang Zhang, Ke Deng, Peng Xue, Zhiming Cui, Yu Fang, Minhui Tang, Yue Zhao, Min Zhu, Zhongxiang Ding, Dinggang Shen
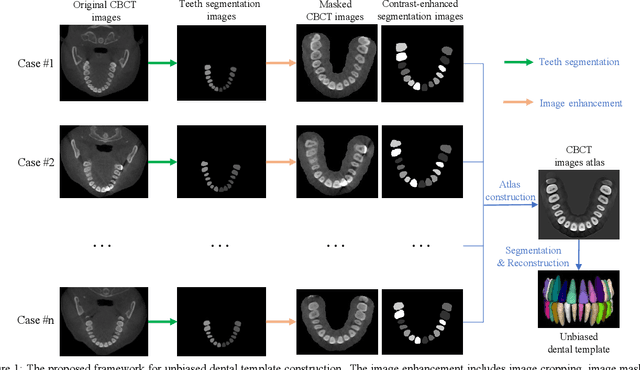
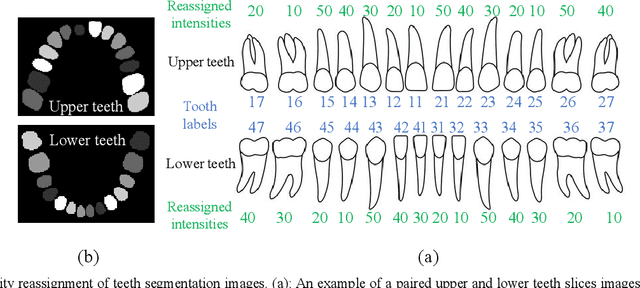
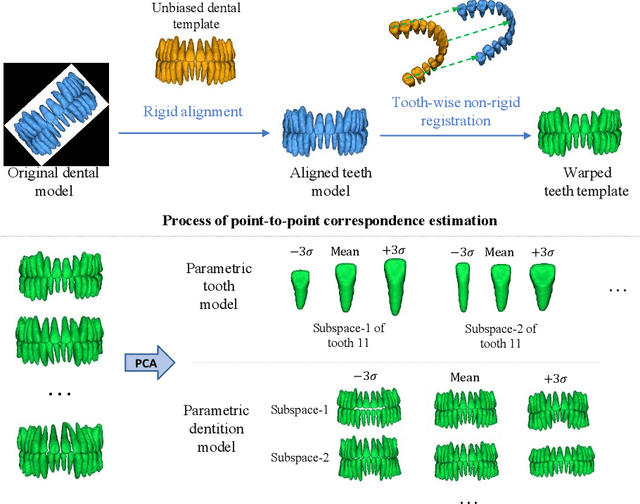
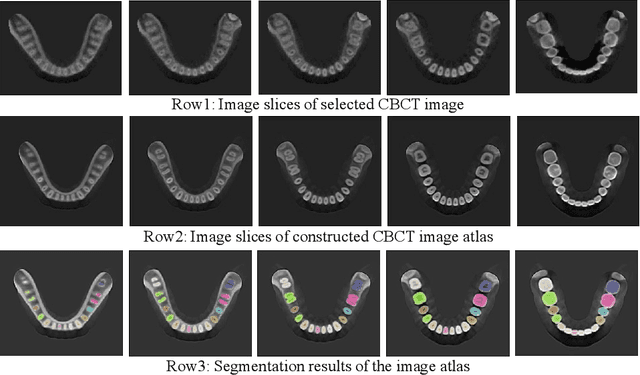
Dental template and parametric dental models are important tools for various applications in digital dentistry. However, constructing an unbiased dental template and accurate parametric dental models remains a challenging task due to the complex anatomical and morphological dental structures and also low volume ratio of the teeth. In this study, we develop an unbiased dental template by constructing an accurate dental atlas from CBCT images with guidance of teeth segmentation. First, to address the challenges, we propose to enhance the CBCT images and their segmentation images, including image cropping, image masking and segmentation intensity reassigning. Then, we further use the segmentation images to perform co-registration with the CBCT images to generate an accurate dental atlas, from which an unbiased dental template can be generated. By leveraging the unbiased dental template, we construct parametric dental models by estimating point-to-point correspondences between the dental models and employing Principal Component Analysis to determine shape subspaces of the parametric dental models. A total of 159 CBCT images of real subjects are collected to perform the constructions. Experimental results demonstrate effectiveness of our proposed method in constructing unbiased dental template and parametric dental model. The developed dental template and parametric dental models are available at https://github.com/Marvin0724/Teeth_template.
Fourier-MIONet: Fourier-enhanced multiple-input neural operators for multiphase modeling of geological carbon sequestration
Mar 08, 2023Zhongyi Jiang, Min Zhu, Dongzhuo Li, Qiuzi Li, Yanhua O. Yuan, Lu Lu
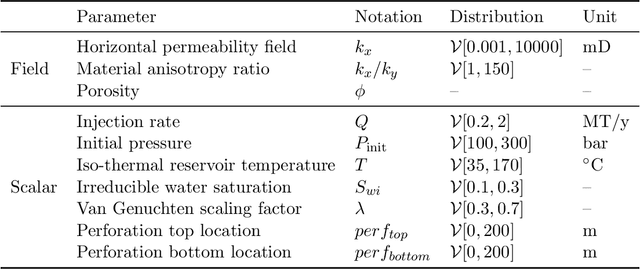
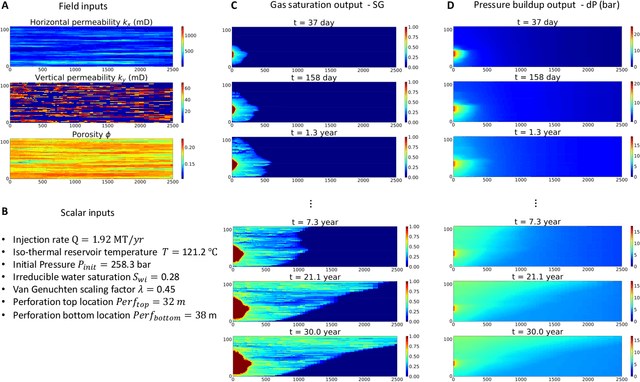

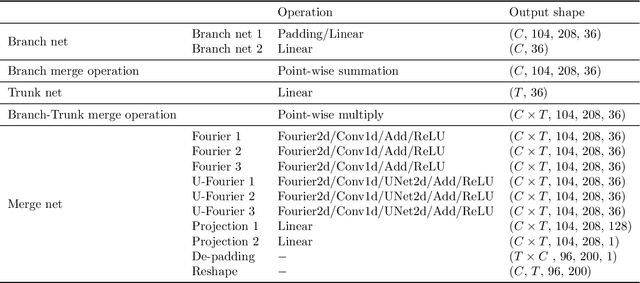
Geologic Carbon Storage (GCS) is an important technology that aims to reduce the amount of carbon dioxide in the atmosphere. Multiphase flow in porous media is essential to understand CO2 migration and pressure fields in the subsurface associated with GCS. However, numerical simulation for such problems in 4D is computationally challenging and expensive, due to the multiphysics and multiscale nature of the highly nonlinear governing partial differential equations (PDEs). It prevents us from considering multiple subsurface scenarios and conducting real-time optimization. Here, we develop a Fourier-enhanced multiple-input neural operator (Fourier-MIONet) to learn the solution operator of the problem of multiphase flow in porous media. Fourier-MIONet utilizes the recently developed framework of the multiple-input deep neural operators (MIONet) and incorporates the Fourier neural operator (FNO) in the network architecture. Once Fourier-MIONet is trained, it can predict the evolution of saturation and pressure of the multiphase flow under various reservoir conditions, such as permeability and porosity heterogeneity, anisotropy, injection configurations, and multiphase flow properties. Compared to the enhanced FNO (U-FNO), the proposed Fourier-MIONet has 90% fewer unknown parameters, and it can be trained in significantly less time (about 3.5 times faster) with much lower CPU memory (< 15%) and GPU memory (< 35%) requirements, to achieve similar prediction accuracy. In addition to the lower computational cost, Fourier-MIONet can be trained with only 6 snapshots of time to predict the PDE solutions for 30 years. The excellent generalizability of Fourier-MIONet is enabled by its adherence to the physical principle that the solution to a PDE is continuous over time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge