Leave No Patient Behind: Enhancing Medication Recommendation for Rare Disease Patients
Mar 26, 2024Zihao Zhao, Yi Jing, Fuli Feng, Jiancan Wu, Chongming Gao, Xiangnan He
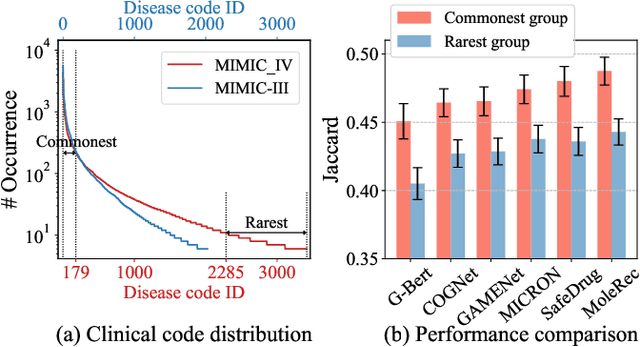

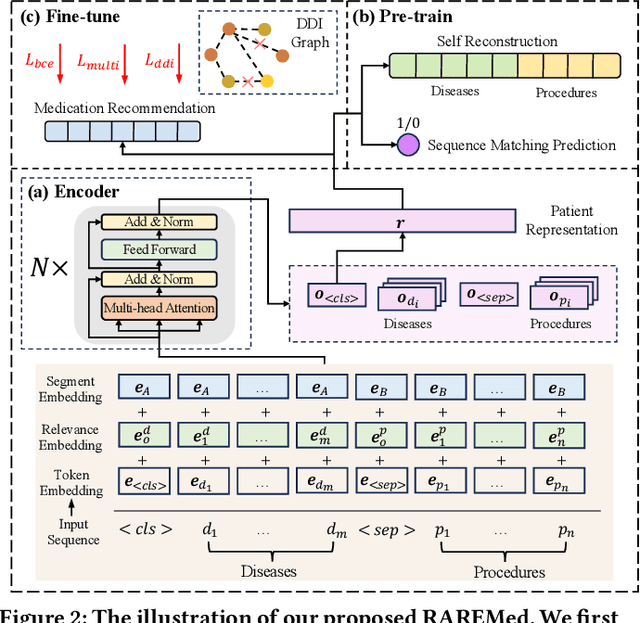
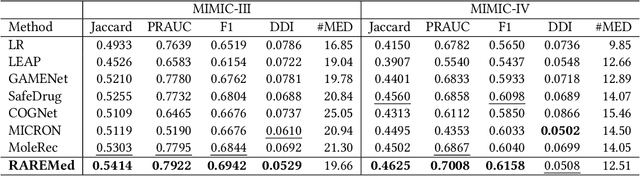
Medication recommendation systems have gained significant attention in healthcare as a means of providing tailored and effective drug combinations based on patients' clinical information. However, existing approaches often suffer from fairness issues, as recommendations tend to be more accurate for patients with common diseases compared to those with rare conditions. In this paper, we propose a novel model called Robust and Accurate REcommendations for Medication (RAREMed), which leverages the pretrain-finetune learning paradigm to enhance accuracy for rare diseases. RAREMed employs a transformer encoder with a unified input sequence approach to capture complex relationships among disease and procedure codes. Additionally, it introduces two self-supervised pre-training tasks, namely Sequence Matching Prediction (SMP) and Self Reconstruction (SR), to learn specialized medication needs and interrelations among clinical codes. Experimental results on two real-world datasets demonstrate that RAREMed provides accurate drug sets for both rare and common disease patients, thereby mitigating unfairness in medication recommendation systems.
An Incremental Update Framework for Online Recommenders with Data-Driven Prior
Dec 26, 2023Chen Yang, Jin Chen, Qian Yu, Xiangdong Wu, Kui Ma, Zihao Zhao, Zhiwei Fang, Wenlong Chen, Chaosheng Fan, Jie He, Changping Peng, Zhangang Lin, Jingping Shao
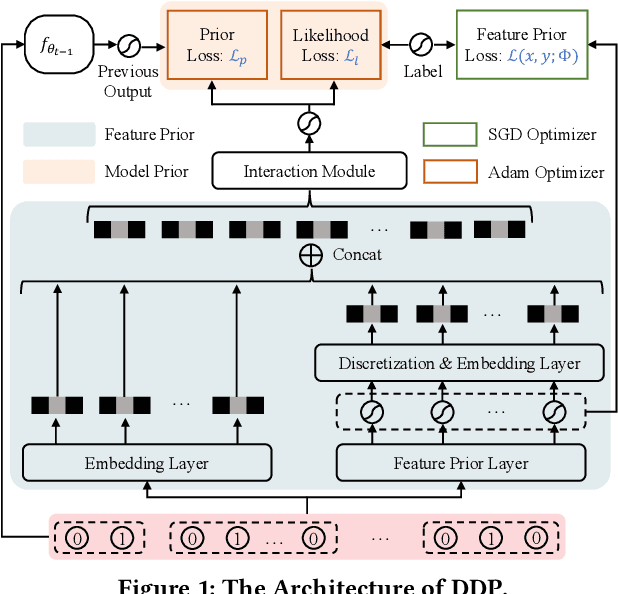
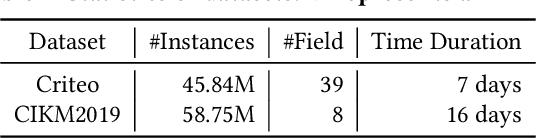
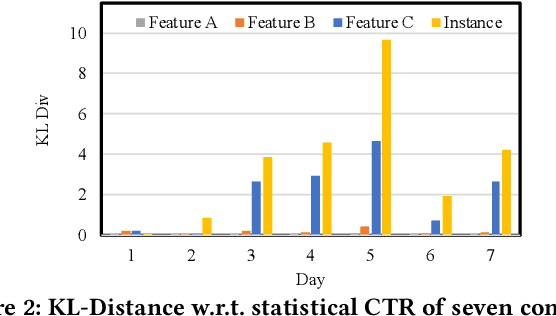
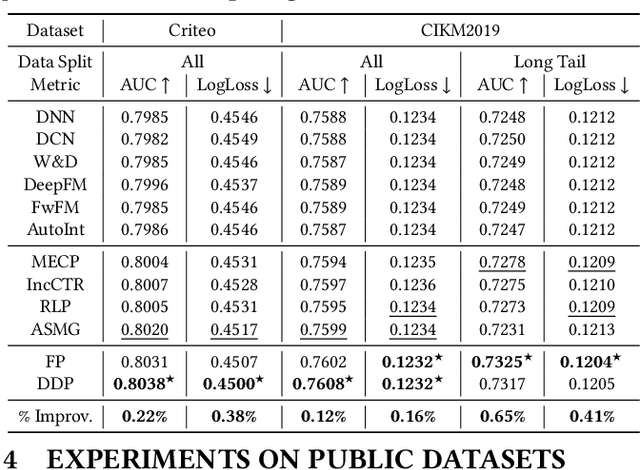
Online recommenders have attained growing interest and created great revenue for businesses. Given numerous users and items, incremental update becomes a mainstream paradigm for learning large-scale models in industrial scenarios, where only newly arrived data within a sliding window is fed into the model, meeting the strict requirements of quick response. However, this strategy would be prone to overfitting to newly arrived data. When there exists a significant drift of data distribution, the long-term information would be discarded, which harms the recommendation performance. Conventional methods address this issue through native model-based continual learning methods, without analyzing the data characteristics for online recommenders. To address the aforementioned issue, we propose an incremental update framework for online recommenders with Data-Driven Prior (DDP), which is composed of Feature Prior (FP) and Model Prior (MP). The FP performs the click estimation for each specific value to enhance the stability of the training process. The MP incorporates previous model output into the current update while strictly following the Bayes rules, resulting in a theoretically provable prior for the robust update. In this way, both the FP and MP are well integrated into the unified framework, which is model-agnostic and can accommodate various advanced interaction models. Extensive experiments on two publicly available datasets as well as an industrial dataset demonstrate the superior performance of the proposed framework.
CLIP in Medical Imaging: A Comprehensive Survey
Dec 26, 2023Zihao Zhao, Yuxiao Liu, Han Wu, Yonghao Li, Sheng Wang, Lin Teng, Disheng Liu, Zhiming Cui, Qian Wang, Dinggang Shen
Contrastive Language-Image Pre-training (CLIP), a simple yet effective pre-training paradigm, successfully introduces text supervision to vision models. It has shown promising results across various tasks, attributable to its generalizability and interpretability. The use of CLIP has recently gained increasing interest in the medical imaging domain, serving both as a pre-training paradigm for aligning medical vision and language, and as a critical component in diverse clinical tasks. With the aim of facilitating a deeper understanding of this promising direction, this survey offers an in-depth exploration of the CLIP paradigm within the domain of medical imaging, regarding both refined CLIP pre-training and CLIP-driven applications. In this study, We (1) start with a brief introduction to the fundamentals of CLIP methodology. (2) Then, we investigate the adaptation of CLIP pre-training in the medical domain, focusing on how to optimize CLIP given characteristics of medical images and reports. (3) Furthermore, we explore the practical utilization of CLIP pre-trained models in various tasks, including classification, dense prediction, and cross-modal tasks. (4) Finally, we discuss existing limitations of CLIP in the context of medical imaging and propose forward-looking directions to address the demands of medical imaging domain. We expect that this comprehensive survey will provide researchers in the field of medical image analysis with a holistic understanding of the CLIP paradigm and its potential implications. The project page can be found on https://github.com/zhaozh10/Awesome-CLIP-in-Medical-Imaging.
Mining Gaze for Contrastive Learning toward Computer-Assisted Diagnosis
Dec 12, 2023Zihao Zhao, Sheng Wang, Qian Wang, Dinggang Shen
Obtaining large-scale radiology reports can be difficult for medical images due to various reasons, limiting the effectiveness of contrastive pre-training in the medical image domain and underscoring the need for alternative methods. In this paper, we propose eye-tracking as an alternative to text reports, as it allows for the passive collection of gaze signals without disturbing radiologist's routine diagnosis process. By tracking the gaze of radiologists as they read and diagnose medical images, we can understand their visual attention and clinical reasoning. When a radiologist has similar gazes for two medical images, it may indicate semantic similarity for diagnosis, and these images should be treated as positive pairs when pre-training a computer-assisted diagnosis (CAD) network through contrastive learning. Accordingly, we introduce the Medical contrastive Gaze Image Pre-training (McGIP) as a plug-and-play module for contrastive learning frameworks. McGIP uses radiologist's gaze to guide contrastive pre-training. We evaluate our method using two representative types of medical images and two common types of gaze data. The experimental results demonstrate the practicality of McGIP, indicating its high potential for various clinical scenarios and applications.
MeLo: Low-rank Adaptation is Better than Fine-tuning for Medical Image Diagnosis
Nov 14, 2023Yitao Zhu, Zhenrong Shen, Zihao Zhao, Sheng Wang, Xin Wang, Xiangyu Zhao, Dinggang Shen, Qian Wang
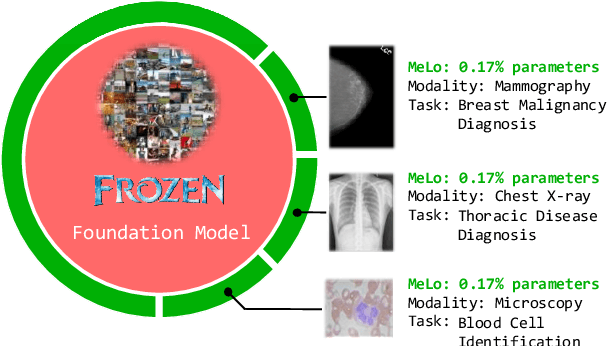
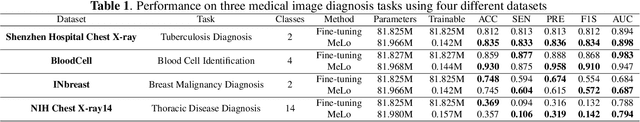

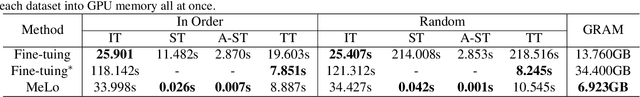
The common practice in developing computer-aided diagnosis (CAD) models based on transformer architectures usually involves fine-tuning from ImageNet pre-trained weights. However, with recent advances in large-scale pre-training and the practice of scaling laws, Vision Transformers (ViT) have become much larger and less accessible to medical imaging communities. Additionally, in real-world scenarios, the deployments of multiple CAD models can be troublesome due to problems such as limited storage space and time-consuming model switching. To address these challenges, we propose a new method MeLo (Medical image Low-rank adaptation), which enables the development of a single CAD model for multiple clinical tasks in a lightweight manner. It adopts low-rank adaptation instead of resource-demanding fine-tuning. By fixing the weight of ViT models and only adding small low-rank plug-ins, we achieve competitive results on various diagnosis tasks across different imaging modalities using only a few trainable parameters. Specifically, our proposed method achieves comparable performance to fully fine-tuned ViT models on four distinct medical imaging datasets using about 0.17% trainable parameters. Moreover, MeLo adds only about 0.5MB of storage space and allows for extremely fast model switching in deployment and inference. Our source code and pre-trained weights are available on our website (https://absterzhu.github.io/melo.github.io/).
Inclusive Data Representation in Federated Learning: A Novel Approach Integrating Textual and Visual Prompt
Oct 04, 2023Zihao Zhao, Zhenpeng Shi, Yang Liu, Wenbo Ding
Federated Learning (FL) is often impeded by communication overhead issues. Prompt tuning, as a potential solution, has been introduced to only adjust a few trainable parameters rather than the whole model. However, current single-modality prompt tuning approaches fail to comprehensively portray local clients' data. To overcome this limitation, we present Twin Prompt Federated learning (TPFL), a pioneering solution that integrates both visual and textual modalities, ensuring a more holistic representation of local clients' data characteristics. Furthermore, in order to tackle the data heterogeneity issues, we introduce the Augmented TPFL (ATPFL) employing the contrastive learning to TPFL, which not only enhances the global knowledge acquisition of client models but also fosters the development of robust, compact models. The effectiveness of TPFL and ATPFL is substantiated by our extensive evaluations, consistently showing superior performance compared to all baselines.
Federated PAC-Bayesian Learning on Non-IID data
Sep 13, 2023Zihao Zhao, Yang Liu, Wenbo Ding, Xiao-Ping Zhang
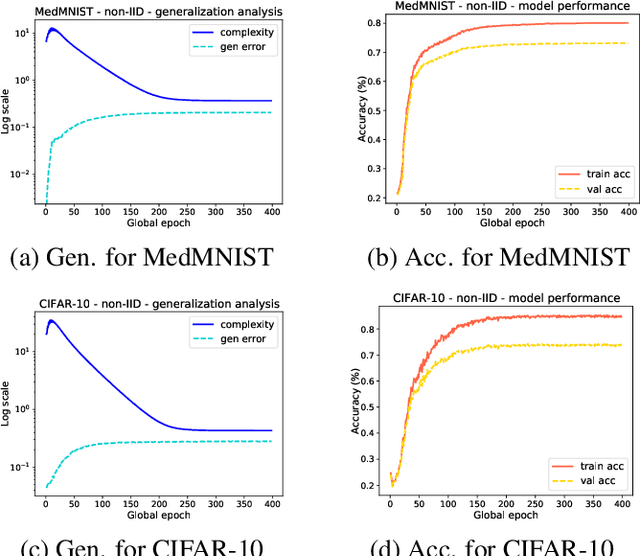
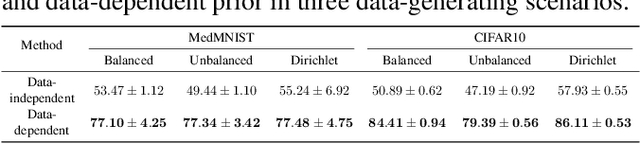
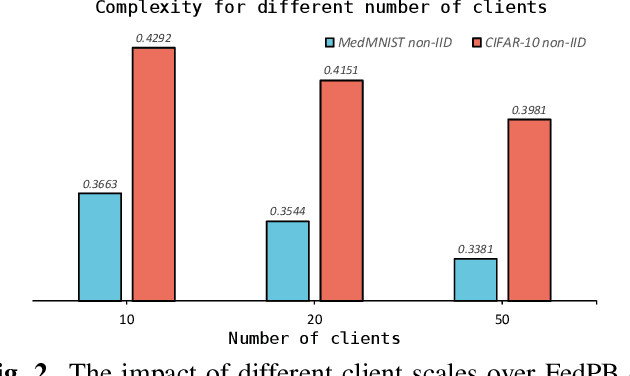
Existing research has either adapted the Probably Approximately Correct (PAC) Bayesian framework for federated learning (FL) or used information-theoretic PAC-Bayesian bounds while introducing their theorems, but few considering the non-IID challenges in FL. Our work presents the first non-vacuous federated PAC-Bayesian bound tailored for non-IID local data. This bound assumes unique prior knowledge for each client and variable aggregation weights. We also introduce an objective function and an innovative Gibbs-based algorithm for the optimization of the derived bound. The results are validated on real-world datasets.
AQUILA: Communication Efficient Federated Learning with Adaptive Quantization of Lazily-Aggregated Gradients
Aug 01, 2023Zihao Zhao, Yuzhu Mao, Zhenpeng Shi, Yang Liu, Tian Lan, Wenbo Ding, Xiao-Ping Zhang
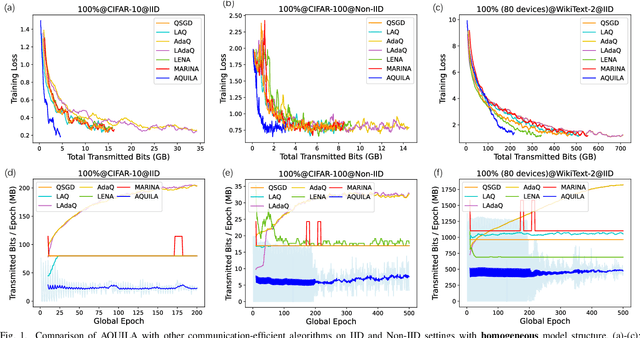
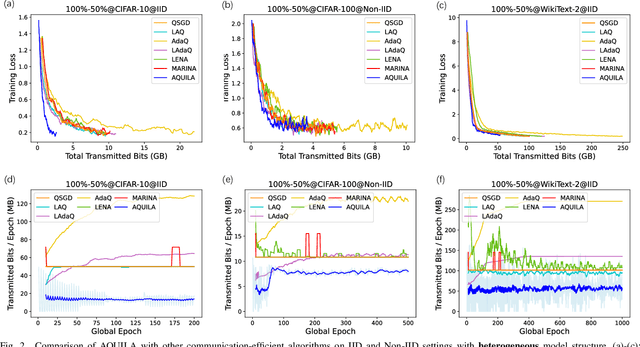

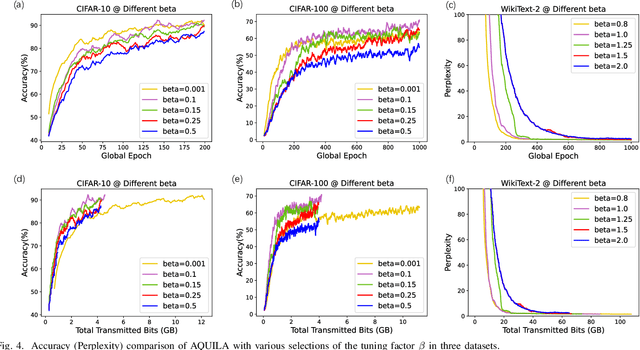
The widespread adoption of Federated Learning (FL), a privacy-preserving distributed learning methodology, has been impeded by the challenge of high communication overheads, typically arising from the transmission of large-scale models. Existing adaptive quantization methods, designed to mitigate these overheads, operate under the impractical assumption of uniform device participation in every training round. Additionally, these methods are limited in their adaptability due to the necessity of manual quantization level selection and often overlook biases inherent in local devices' data, thereby affecting the robustness of the global model. In response, this paper introduces AQUILA (adaptive quantization of lazily-aggregated gradients), a novel adaptive framework devised to effectively handle these issues, enhancing the efficiency and robustness of FL. AQUILA integrates a sophisticated device selection method that prioritizes the quality and usefulness of device updates. Utilizing the exact global model stored by devices, it enables a more precise device selection criterion, reduces model deviation, and limits the need for hyperparameter adjustments. Furthermore, AQUILA presents an innovative quantization criterion, optimized to improve communication efficiency while assuring model convergence. Our experiments demonstrate that AQUILA significantly decreases communication costs compared to existing methods, while maintaining comparable model performance across diverse non-homogeneous FL settings, such as Non-IID data and heterogeneous model architectures.
Evaluating Large Language Models for Radiology Natural Language Processing
Jul 27, 2023Zhengliang Liu, Tianyang Zhong, Yiwei Li, Yutong Zhang, Yi Pan, Zihao Zhao, Peixin Dong, Chao Cao, Yuxiao Liu, Peng Shu, Yaonai Wei, Zihao Wu, Chong Ma, Jiaqi Wang, Sheng Wang, Mengyue Zhou, Zuowei Jiang, Chunlin Li, Jason Holmes, Shaochen Xu, Lu Zhang, Haixing Dai, Kai Zhang, Lin Zhao, Yuanhao Chen, Xu Liu, Peilong Wang, Pingkun Yan, Jun Liu, Bao Ge, Lichao Sun, Dajiang Zhu, Xiang Li, Wei Liu, Xiaoyan Cai, Xintao Hu, Xi Jiang, Shu Zhang, Xin Zhang, Tuo Zhang, Shijie Zhao, Quanzheng Li, Hongtu Zhu, Dinggang Shen, Tianming Liu
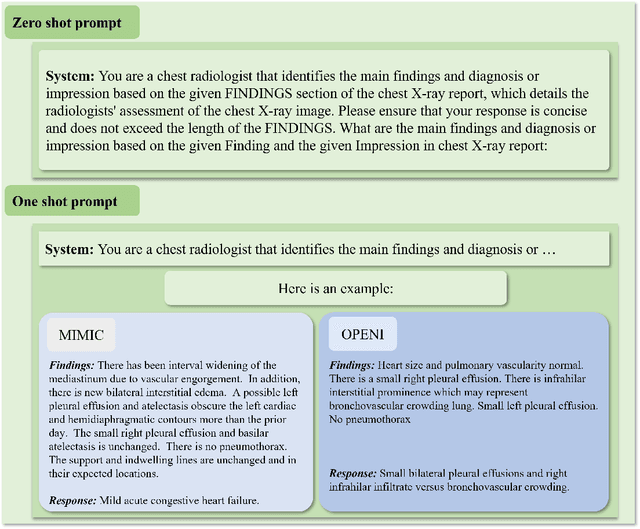

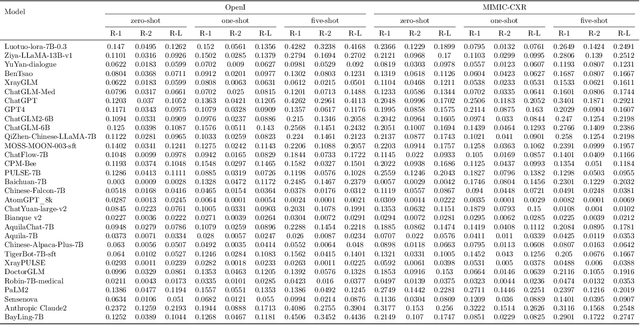
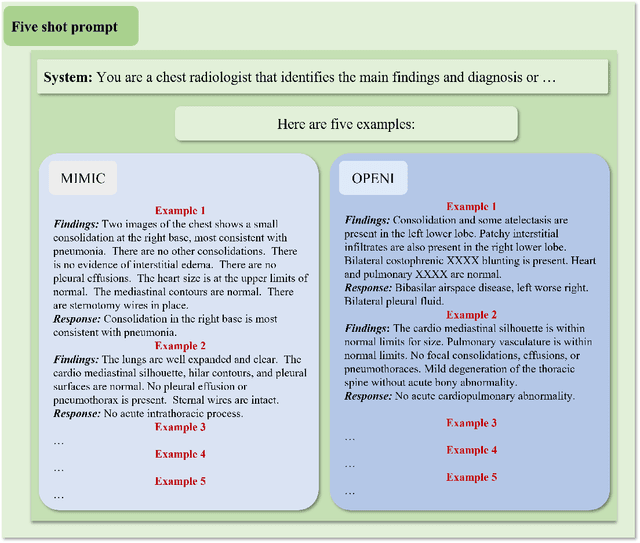
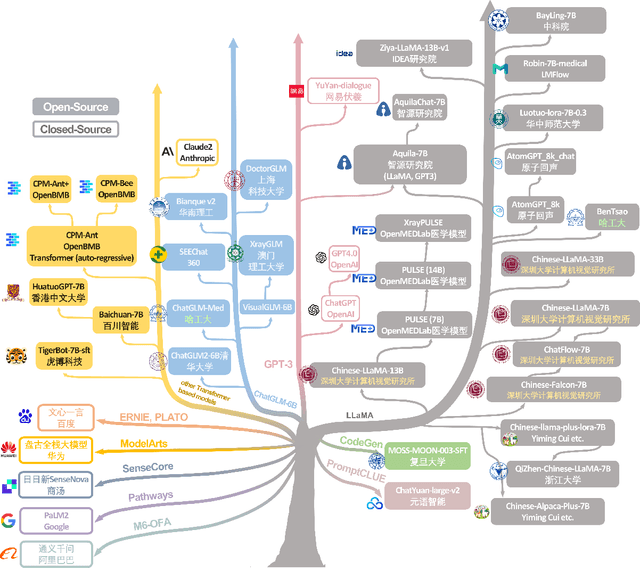
The rise of large language models (LLMs) has marked a pivotal shift in the field of natural language processing (NLP). LLMs have revolutionized a multitude of domains, and they have made a significant impact in the medical field. Large language models are now more abundant than ever, and many of these models exhibit bilingual capabilities, proficient in both English and Chinese. However, a comprehensive evaluation of these models remains to be conducted. This lack of assessment is especially apparent within the context of radiology NLP. This study seeks to bridge this gap by critically evaluating thirty two LLMs in interpreting radiology reports, a crucial component of radiology NLP. Specifically, the ability to derive impressions from radiologic findings is assessed. The outcomes of this evaluation provide key insights into the performance, strengths, and weaknesses of these LLMs, informing their practical applications within the medical domain.
ChatCAD+: Towards a Universal and Reliable Interactive CAD using LLMs
May 26, 2023Zihao Zhao, Sheng Wang, Jinchen Gu, Yitao Zhu, Lanzhuju Mei, Zixu Zhuang, Zhiming Cui, Qian Wang, Dinggang Shen
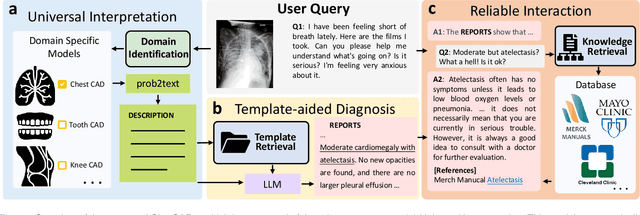


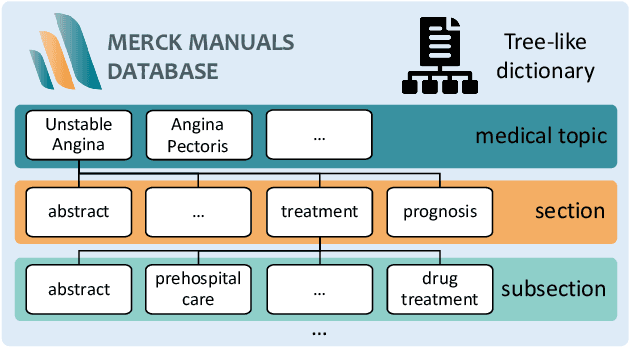
The potential of integrating Computer-Assisted Diagnosis (CAD) with Large Language Models (LLMs) in clinical applications, particularly in digital family doctor and clinic assistant roles, shows promise. However, existing works have limitations in terms of reliability, effectiveness, and their narrow applicability to specific image domains, which restricts their overall processing capabilities. Moreover, the mismatch in writing style between LLMs and radiologists undermines their practical utility. To address these challenges, we present ChatCAD+, an interactive CAD system that is universal, reliable, and capable of handling medical images from diverse domains. ChatCAD+ utilizes current information obtained from reputable medical websites to offer precise medical advice. Additionally, it incorporates a template retrieval system that emulates real-world diagnostic reporting, thereby improving its seamless integration into existing clinical workflows. The source code is available at https://github.com/zhaozh10/ChatCAD. The online demo will be available soon.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge