Underestimation of lung regions on chest X-ray segmentation masks assessed by comparison with total lung volume evaluated on computed tomography
Feb 18, 2024Przemysław Bombiński, Patryk Szatkowski, Bartłomiej Sobieski, Tymoteusz Kwieciński, Szymon Płotka, Mariusz Adamek, Marcin Banasiuk, Mariusz I. Furmanek, Przemysław Biecek
Lung mask creation lacks well-defined criteria and standardized guidelines, leading to a high degree of subjectivity between annotators. In this study, we assess the underestimation of lung regions on chest X-ray segmentation masks created according to the current state-of-the-art method, by comparison with total lung volume evaluated on computed tomography (CT). We show, that lung X-ray masks created by following the contours of the heart, mediastinum, and diaphragm significantly underestimate lung regions and exclude substantial portions of the lungs from further assessment, which may result in numerous clinical errors.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024Kelly Payette, Céline Steger, Roxane Licandro, Priscille de Dumast, Hongwei Bran Li, Matthew Barkovich, Liu Li, Maik Dannecker, Chen Chen, Cheng Ouyang, Niccolò McConnell, Alina Miron, Yongmin Li, Alena Uus, Irina Grigorescu, Paula Ramirez Gilliland, Md Mahfuzur Rahman Siddiquee, Daguang Xu, Andriy Myronenko, Haoyu Wang, Ziyan Huang, Jin Ye, Mireia Alenyà, Valentin Comte, Oscar Camara, Jean-Baptiste Masson, Astrid Nilsson, Charlotte Godard, Moona Mazher, Abdul Qayyum, Yibo Gao, Hangqi Zhou, Shangqi Gao, Jia Fu, Guiming Dong, Guotai Wang, ZunHyan Rieu, HyeonSik Yang, Minwoo Lee, Szymon Płotka, Michal K. Grzeszczyk, Arkadiusz Sitek, Luisa Vargas Daza, Santiago Usma, Pablo Arbelaez, Wenying Lu, Wenhao Zhang, Jing Liang, Romain Valabregue, Anand A. Joshi, Krishna N. Nayak, Richard M. Leahy, Luca Wilhelmi, Aline Dändliker, Hui Ji, Antonio G. Gennari, Anton Jakovčić, Melita Klaić, Ana Adžić, Pavel Marković, Gracia Grabarić, Gregor Kasprian, Gregor Dovjak, Milan Rados, Lana Vasung, Meritxell Bach Cuadra, Andras Jakab
Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
TabAttention: Learning Attention Conditionally on Tabular Data
Oct 27, 2023Michal K. Grzeszczyk, Szymon Płotka, Beata Rebizant, Katarzyna Kosińska-Kaczyńska, Michał Lipa, Robert Brawura-Biskupski-Samaha, Przemysław Korzeniowski, Tomasz Trzciński, Arkadiusz Sitek
Medical data analysis often combines both imaging and tabular data processing using machine learning algorithms. While previous studies have investigated the impact of attention mechanisms on deep learning models, few have explored integrating attention modules and tabular data. In this paper, we introduce TabAttention, a novel module that enhances the performance of Convolutional Neural Networks (CNNs) with an attention mechanism that is trained conditionally on tabular data. Specifically, we extend the Convolutional Block Attention Module to 3D by adding a Temporal Attention Module that uses multi-head self-attention to learn attention maps. Furthermore, we enhance all attention modules by integrating tabular data embeddings. Our approach is demonstrated on the fetal birth weight (FBW) estimation task, using 92 fetal abdominal ultrasound video scans and fetal biometry measurements. Our results indicate that TabAttention outperforms clinicians and existing methods that rely on tabular and/or imaging data for FBW prediction. This novel approach has the potential to improve computer-aided diagnosis in various clinical workflows where imaging and tabular data are combined. We provide a source code for integrating TabAttention in CNNs at https://github.com/SanoScience/Tab-Attention.
Why is the winner the best?
Mar 30, 2023Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Sharib Ali, Vincent Andrearczyk, Marc Aubreville, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Veronika Cheplygina, Marie Daum, Marleen de Bruijne, Adrien Depeursinge, Reuben Dorent, Jan Egger, David G. Ellis, Sandy Engelhardt, Melanie Ganz, Noha Ghatwary, Gabriel Girard, Patrick Godau, Anubha Gupta, Lasse Hansen, Kanako Harada, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Pierre Jannin, Ali Emre Kavur, Oldřich Kodym, Michal Kozubek, Jianning Li, Hongwei Li, Jun Ma, Carlos Martín-Isla, Bjoern Menze, Alison Noble, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Tim Rädsch, Jonathan Rafael-Patiño, Vivek Singh Bawa, Stefanie Speidel, Carole H. Sudre, Kimberlin van Wijnen, Martin Wagner, Donglai Wei, Amine Yamlahi, Moi Hoon Yap, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Dogu Baran Aydogan, Binod Bhattarai, Louise Bloch, Raphael Brüngel, Jihoon Cho, Chanyeol Choi, Qi Dou, Ivan Ezhov, Christoph M. Friedrich, Clifton Fuller, Rebati Raman Gaire, Adrian Galdran, Álvaro García Faura, Maria Grammatikopoulou, SeulGi Hong, Mostafa Jahanifar, Ikbeom Jang, Abdolrahim Kadkhodamohammadi, Inha Kang, Florian Kofler, Satoshi Kondo, Hugo Kuijf, Mingxing Li, Minh Huan Luu, Tomaž Martinčič, Pedro Morais, Mohamed A. Naser, Bruno Oliveira, David Owen, Subeen Pang, Jinah Park, Sung-Hong Park, Szymon Płotka, Elodie Puybareau, Nasir Rajpoot, Kanghyun Ryu, Numan Saeed, Adam Shephard, Pengcheng Shi, Dejan Štepec, Ronast Subedi, Guillaume Tochon, Helena R. Torres, Helene Urien, João L. Vilaça, Kareem Abdul Wahid, Haojie Wang, Jiacheng Wang, Liansheng Wang, Xiyue Wang, Benedikt Wiestler, Marek Wodzinski, Fangfang Xia, Juanying Xie, Zhiwei Xiong, Sen Yang, Yanwu Yang, Zixuan Zhao, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
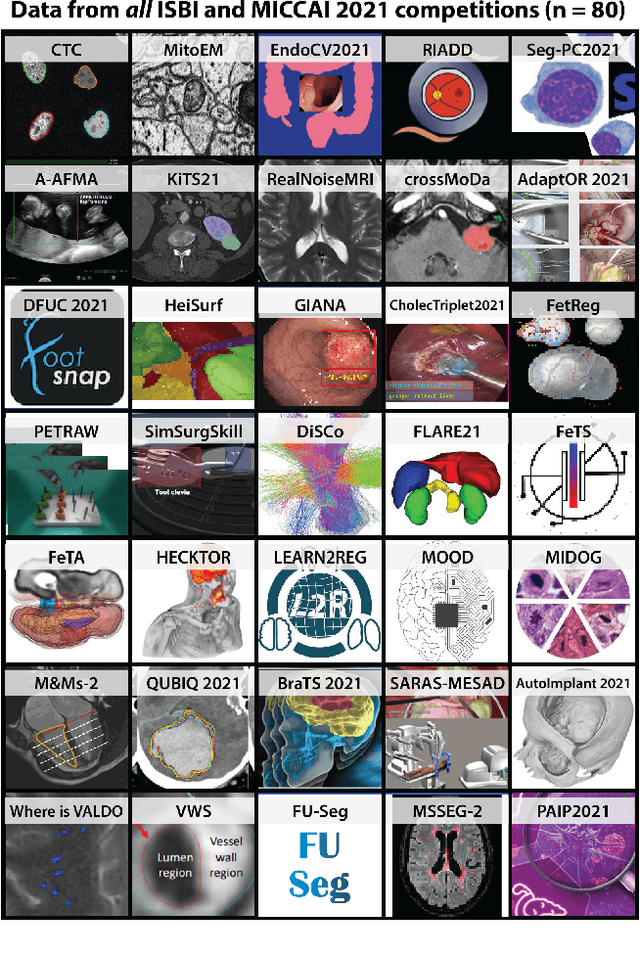
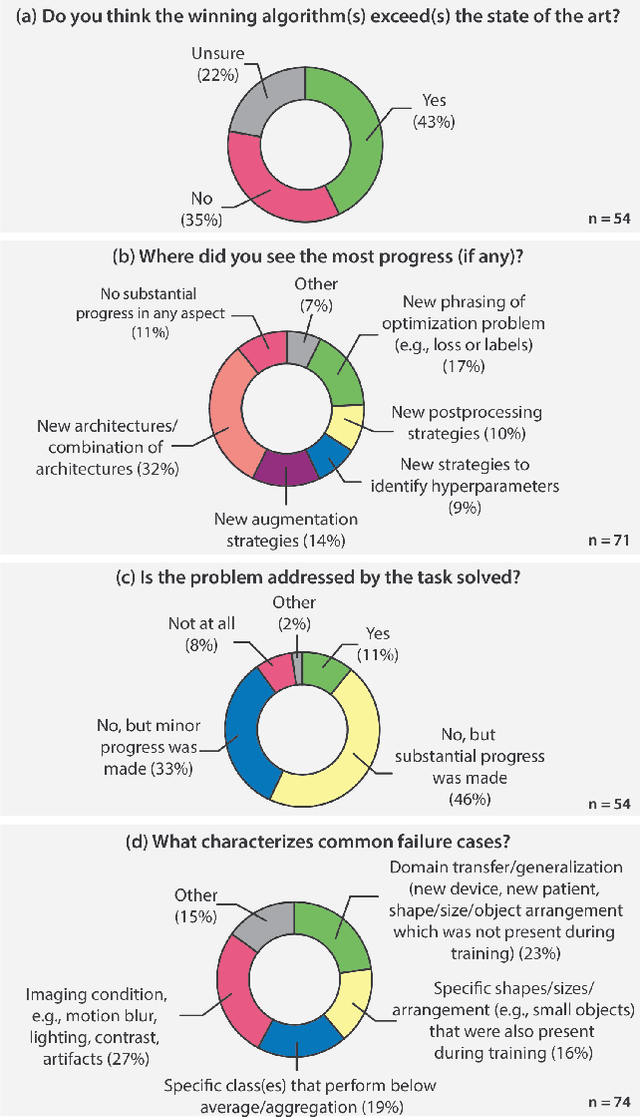
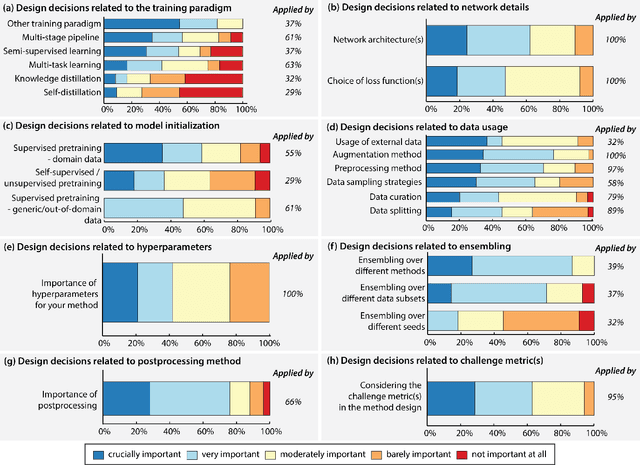
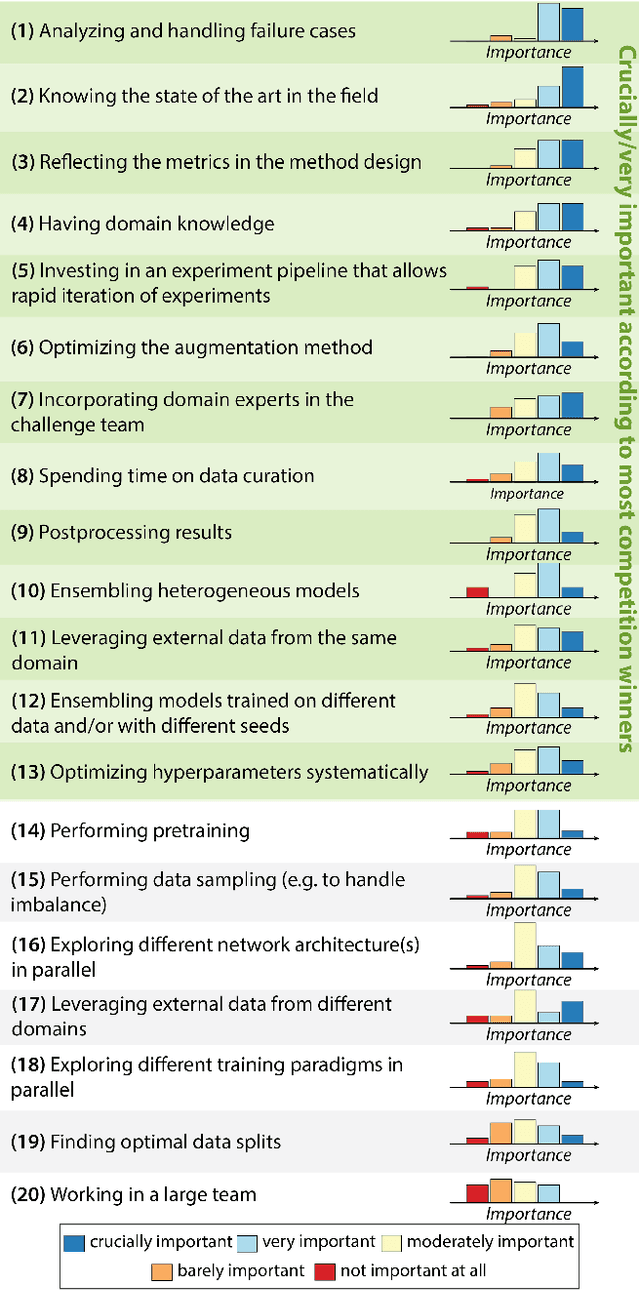
International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
Multi-task Swin Transformer for Motion Artifacts Classification and Cardiac Magnetic Resonance Image Segmentation
Sep 06, 2022Michal K. Grzeszczyk, Szymon Płotka, Arkadiusz Sitek
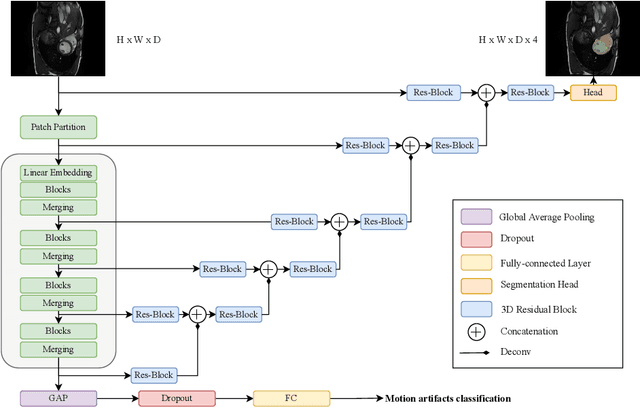
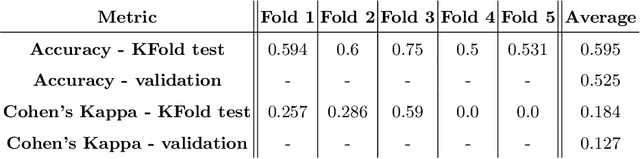

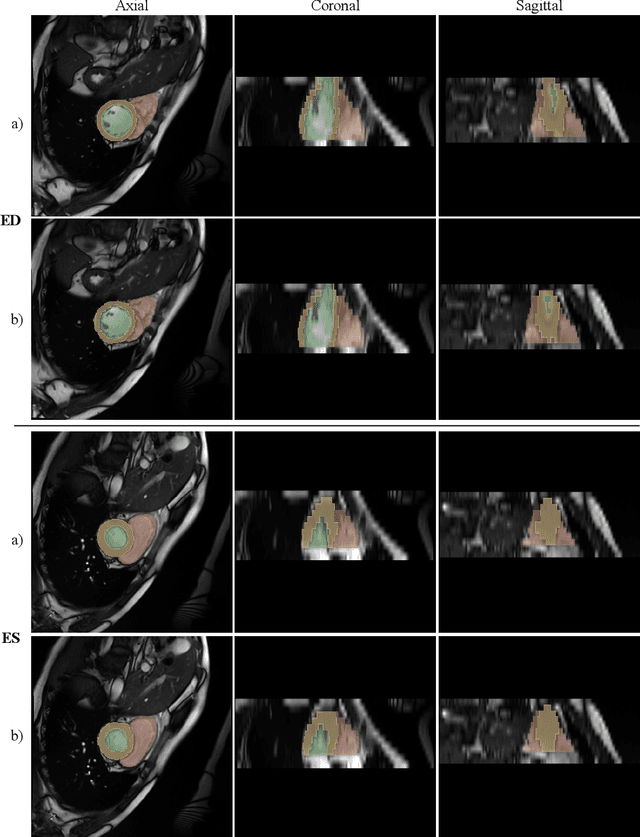
Cardiac Magnetic Resonance Imaging is commonly used for the assessment of the cardiac anatomy and function. The delineations of left and right ventricle blood pools and left ventricular myocardium are important for the diagnosis of cardiac diseases. Unfortunately, the movement of a patient during the CMR acquisition procedure may result in motion artifacts appearing in the final image. Such artifacts decrease the diagnostic quality of CMR images and force redoing of the procedure. In this paper, we present a Multi-task Swin UNEt TRansformer network for simultaneous solving of two tasks in the CMRxMotion challenge: CMR segmentation and motion artifacts classification. We utilize both segmentation and classification as a multi-task learning approach which allows us to determine the diagnostic quality of CMR and generate masks at the same time. CMR images are classified into three diagnostic quality classes, whereas, all samples with non-severe motion artifacts are being segmented. Ensemble of five networks trained using 5-Fold Cross-validation achieves segmentation performance of DICE coefficient of 0.871 and classification accuracy of 0.595.
Virtual Reality Simulator for Fetoscopic Spina Bifida Repair Surgery
Jul 30, 2022Przemysław Korzeniowski, Szymon Płotka, Robert Brawura-Biskupski-Samaha, Arkadiusz Sitek
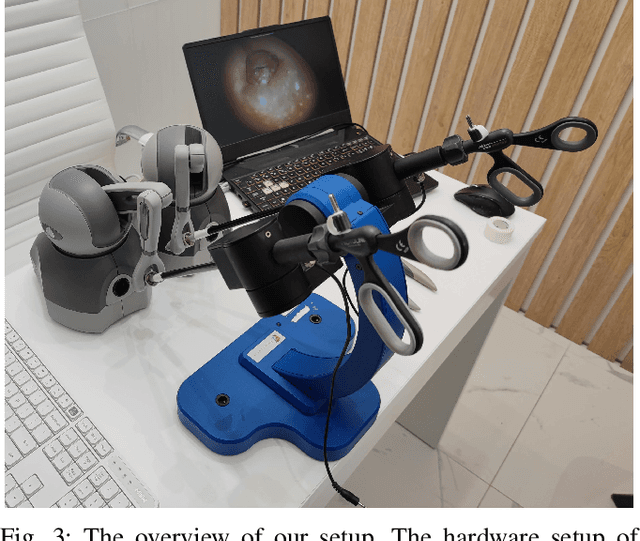
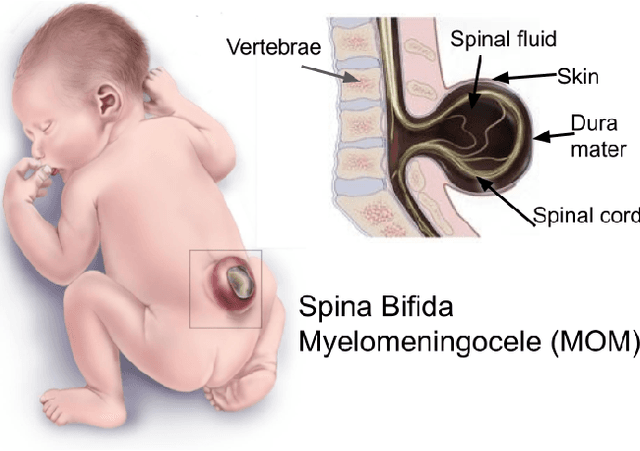


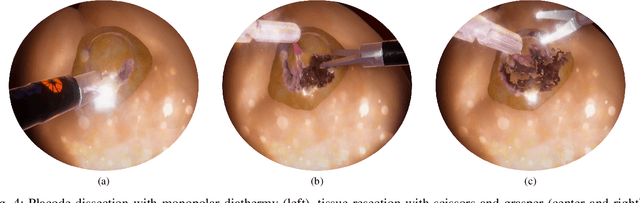
Spina Bifida (SB) is a birth defect developed during the early stage of pregnancy in which there is incomplete closing of the spine around the spinal cord. The growing interest in fetoscopic Spina-Bifida repair, which is performed in fetuses who are still in the pregnant uterus, prompts the need for appropriate training. The learning curve for such procedures is steep and requires excellent procedural skills. Computer-based virtual reality (VR) simulation systems offer a safe, cost-effective, and configurable training environment free from ethical and patient safety issues. However, to the best of our knowledge, there are currently no commercial or experimental VR training simulation systems available for fetoscopic SB-repair procedures. In this paper, we propose a novel VR simulator for core manual skills training for SB-repair. An initial simulation realism validation study was carried out by obtaining subjective feedback (face and content validity) from 14 clinicians. The overall simulation realism was on average marked 4.07 on a 5-point Likert scale (1 - very unrealistic, 5 - very realistic). Its usefulness as a training tool for SB-repair as well as in learning fundamental laparoscopic skills was marked 4.63 and 4.80, respectively. These results indicate that VR simulation of fetoscopic procedures may contribute to surgical training without putting fetuses and their mothers at risk. It could also facilitate wider adaptation of fetoscopic procedures in place of much more invasive open fetal surgeries.
FetReg2021: A Challenge on Placental Vessel Segmentation and Registration in Fetoscopy
Jun 30, 2022Sophia Bano, Alessandro Casella, Francisco Vasconcelos, Abdul Qayyum, Abdesslam Benzinou, Moona Mazher, Fabrice Meriaudeau, Chiara Lena, Ilaria Anita Cintorrino, Gaia Romana De Paolis, Jessica Biagioli, Daria Grechishnikova, Jing Jiao, Bizhe Bai, Yanyan Qiao, Binod Bhattarai, Rebati Raman Gaire, Ronast Subedi, Eduard Vazquez, Szymon Płotka, Aneta Lisowska, Arkadiusz Sitek, George Attilakos, Ruwan Wimalasundera, Anna L David, Dario Paladini, Jan Deprest, Elena De Momi, Leonardo S Mattos, Sara Moccia, Danail Stoyanov
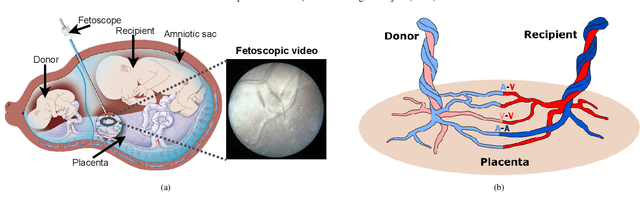
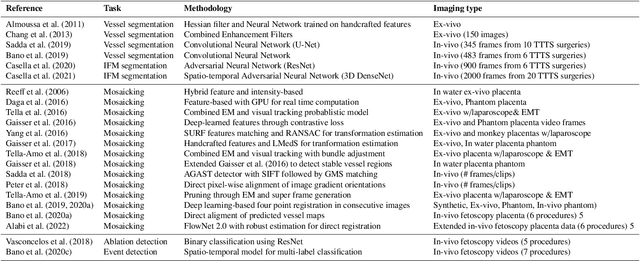
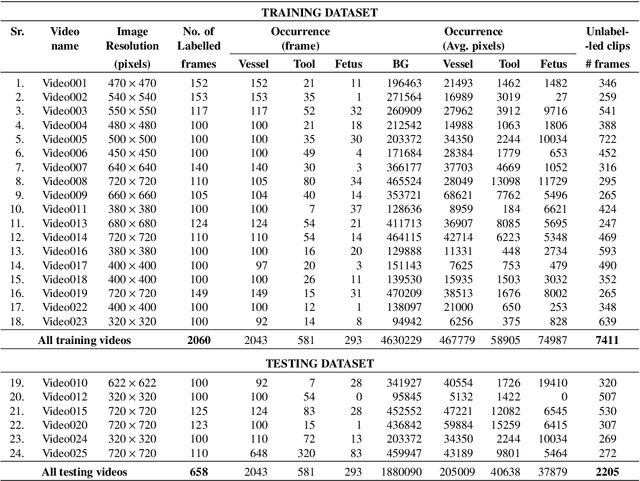
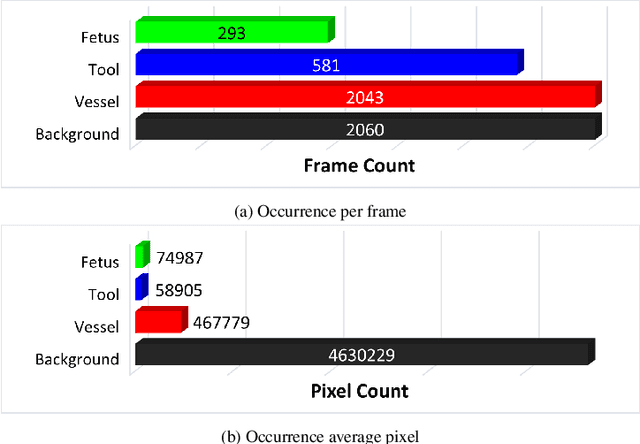
Fetoscopy laser photocoagulation is a widely adopted procedure for treating Twin-to-Twin Transfusion Syndrome (TTTS). The procedure involves photocoagulation pathological anastomoses to regulate blood exchange among twins. The procedure is particularly challenging due to the limited field of view, poor manoeuvrability of the fetoscope, poor visibility, and variability in illumination. These challenges may lead to increased surgery time and incomplete ablation. Computer-assisted intervention (CAI) can provide surgeons with decision support and context awareness by identifying key structures in the scene and expanding the fetoscopic field of view through video mosaicking. Research in this domain has been hampered by the lack of high-quality data to design, develop and test CAI algorithms. Through the Fetoscopic Placental Vessel Segmentation and Registration (FetReg2021) challenge, which was organized as part of the MICCAI2021 Endoscopic Vision challenge, we released the first largescale multicentre TTTS dataset for the development of generalized and robust semantic segmentation and video mosaicking algorithms. For this challenge, we released a dataset of 2060 images, pixel-annotated for vessels, tool, fetus and background classes, from 18 in-vivo TTTS fetoscopy procedures and 18 short video clips. Seven teams participated in this challenge and their model performance was assessed on an unseen test dataset of 658 pixel-annotated images from 6 fetoscopic procedures and 6 short clips. The challenge provided an opportunity for creating generalized solutions for fetoscopic scene understanding and mosaicking. In this paper, we present the findings of the FetReg2021 challenge alongside reporting a detailed literature review for CAI in TTTS fetoscopy. Through this challenge, its analysis and the release of multi-centre fetoscopic data, we provide a benchmark for future research in this field.
Deep Learning Fetal Ultrasound Video Model Match Human Observers in Biometric Measurements
May 27, 2022Szymon Płotka, Adam Klasa, Aneta Lisowska, Joanna Seliga-Siwecka, Michał Lipa, Tomasz Trzciński, Arkadiusz Sitek
Objective. This work investigates the use of deep convolutional neural networks (CNN) to automatically perform measurements of fetal body parts, including head circumference, biparietal diameter, abdominal circumference and femur length, and to estimate gestational age and fetal weight using fetal ultrasound videos. Approach. We developed a novel multi-task CNN-based spatio-temporal fetal US feature extraction and standard plane detection algorithm (called FUVAI) and evaluated the method on 50 freehand fetal US video scans. We compared FUVAI fetal biometric measurements with measurements made by five experienced sonographers at two time points separated by at least two weeks. Intra- and inter-observer variabilities were estimated. Main results. We found that automated fetal biometric measurements obtained by FUVAI were comparable to the measurements performed by experienced sonographers The observed differences in measurement values were within the range of inter- and intra-observer variability. Moreover, analysis has shown that these differences were not statistically significant when comparing any individual medical expert to our model. Significance. We argue that FUVAI has the potential to assist sonographers who perform fetal biometric measurements in clinical settings by providing them with suggestions regarding the best measuring frames, along with automated measurements. Moreover, FUVAI is able perform these tasks in just a few seconds, which is a huge difference compared to the average of six minutes taken by sonographers. This is significant, given the shortage of medical experts capable of interpreting fetal ultrasound images in numerous countries.
* Published at Physics in Medicine & Biology
BabyNet: Residual Transformer Module for Birth Weight Prediction on Fetal Ultrasound Video
May 19, 2022Szymon Płotka, Michał K. Grzeszczyk, Robert Brawura-Biskupski-Samaha, Paweł Gutaj, Michał Lipa, Tomasz Trzciński, Arkadiusz Sitek
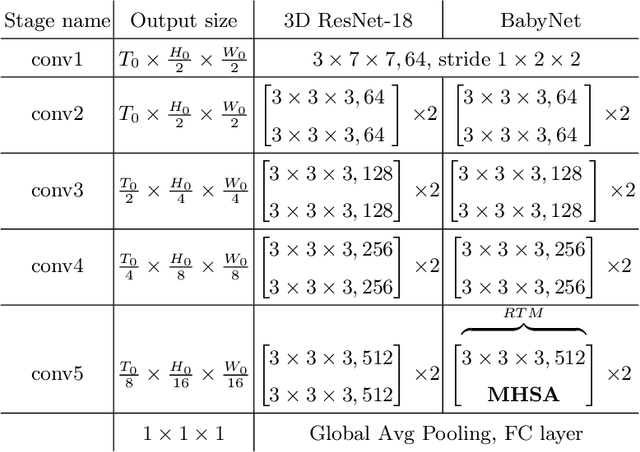
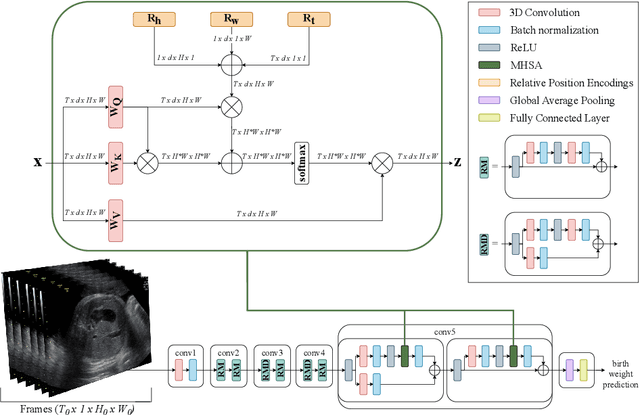

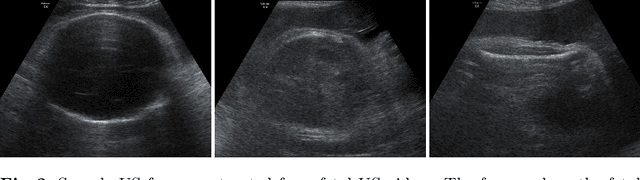

Predicting fetal weight at birth is an important aspect of perinatal care, particularly in the context of antenatal management, which includes the planned timing and the mode of delivery. Accurate prediction of weight using prenatal ultrasound is challenging as it requires images of specific fetal body parts during advanced pregnancy which is difficult to capture due to poor quality of images caused by the lack of amniotic fluid. As a consequence, predictions which rely on standard methods often suffer from significant errors. In this paper we propose the Residual Transformer Module which extends a 3D ResNet-based network for analysis of 2D+t spatio-temporal ultrasound video scans. Our end-to-end method, called BabyNet, automatically predicts fetal birth weight based on fetal ultrasound video scans. We evaluate BabyNet using a dedicated clinical set comprising 225 2D fetal ultrasound videos of pregnancies from 75 patients performed one day prior to delivery. Experimental results show that BabyNet outperforms several state-of-the-art methods and estimates the weight at birth with accuracy comparable to human experts. Furthermore, combining estimates provided by human experts with those computed by BabyNet yields the best results, outperforming either of other methods by a significant margin. The source code of BabyNet is available at https://github.com/SanoScience/BabyNet.
CXR-FL: Deep Learning-based Chest X-ray Image Analysis Using Federated Learning
Apr 11, 2022Filip Ślazyk, Przemysław Jabłecki, Aneta Lisowska, Maciej Malawski, Szymon Płotka
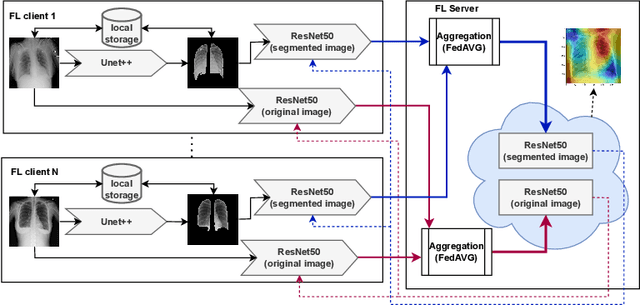

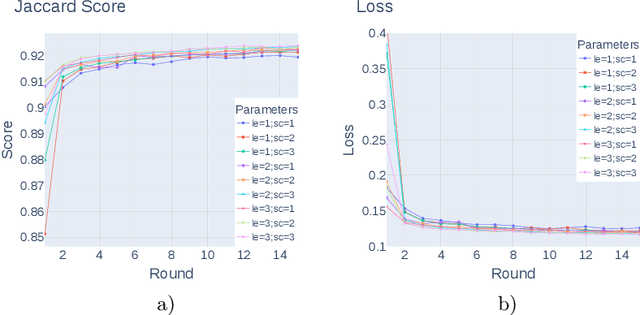
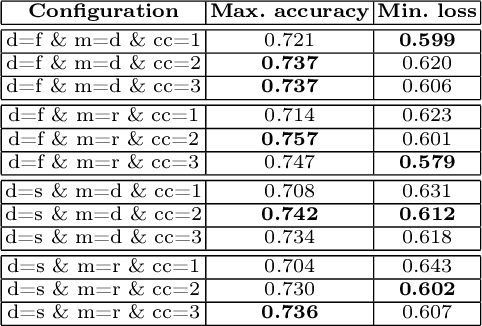
Federated learning enables building a shared model from multicentre data while storing the training data locally for privacy. In this paper, we present an evaluation (called CXR-FL) of deep learning-based models for chest X-ray image analysis using the federated learning method. We examine the impact of federated learning parameters on the performance of central models. Additionally, we show that classification models perform worse if trained on a region of interest reduced to segmentation of the lung compared to the full image. However, focusing training of the classification model on the lung area may result in improved pathology interpretability during inference. We also find that federated learning helps maintain model generalizability. The pre-trained weights and code are publicly available at (https://github.com/SanoScience/CXR-FL).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge