Gaussian Pancakes: Geometrically-Regularized 3D Gaussian Splatting for Realistic Endoscopic Reconstruction
Apr 09, 2024Sierra Bonilla, Shuai Zhang, Dimitrios Psychogyios, Danail Stoyanov, Francisco Vasconcelos, Sophia Bano
Within colorectal cancer diagnostics, conventional colonoscopy techniques face critical limitations, including a limited field of view and a lack of depth information, which can impede the detection of precancerous lesions. Current methods struggle to provide comprehensive and accurate 3D reconstructions of the colonic surface which can help minimize the missing regions and reinspection for pre-cancerous polyps. Addressing this, we introduce 'Gaussian Pancakes', a method that leverages 3D Gaussian Splatting (3D GS) combined with a Recurrent Neural Network-based Simultaneous Localization and Mapping (RNNSLAM) system. By introducing geometric and depth regularization into the 3D GS framework, our approach ensures more accurate alignment of Gaussians with the colon surface, resulting in smoother 3D reconstructions with novel viewing of detailed textures and structures. Evaluations across three diverse datasets show that Gaussian Pancakes enhances novel view synthesis quality, surpassing current leading methods with a 18% boost in PSNR and a 16% improvement in SSIM. It also delivers over 100X faster rendering and more than 10X shorter training times, making it a practical tool for real-time applications. Hence, this holds promise for achieving clinical translation for better detection and diagnosis of colorectal cancer.
Investigation of Federated Learning Algorithms for Retinal Optical Coherence Tomography Image Classification with Statistical Heterogeneity
Feb 15, 2024Sanskar Amgain, Prashant Shrestha, Sophia Bano, Ignacio del Valle Torres, Michael Cunniffe, Victor Hernandez, Phil Beales, Binod Bhattarai
Purpose: We apply federated learning to train an OCT image classifier simulating a realistic scenario with multiple clients and statistical heterogeneous data distribution where data in the clients lack samples of some categories entirely. Methods: We investigate the effectiveness of FedAvg and FedProx to train an OCT image classification model in a decentralized fashion, addressing privacy concerns associated with centralizing data. We partitioned a publicly available OCT dataset across multiple clients under IID and Non-IID settings and conducted local training on the subsets for each client. We evaluated two federated learning methods, FedAvg and FedProx for these settings. Results: Our experiments on the dataset suggest that under IID settings, both methods perform on par with training on a central data pool. However, the performance of both algorithms declines as we increase the statistical heterogeneity across the client data, while FedProx consistently performs better than FedAvg in the increased heterogeneity settings. Conclusion: Despite the effectiveness of federated learning in the utilization of private data across multiple medical institutions, the large number of clients and heterogeneous distribution of labels deteriorate the performance of both algorithms. Notably, FedProx appears to be more robust to the increased heterogeneity.
SimCol3D -- 3D Reconstruction during Colonoscopy Challenge
Jul 20, 2023Anita Rau, Sophia Bano, Yueming Jin, Pablo Azagra, Javier Morlana, Edward Sanderson, Bogdan J. Matuszewski, Jae Young Lee, Dong-Jae Lee, Erez Posner, Netanel Frank, Varshini Elangovan, Sista Raviteja, Zhengwen Li, Jiquan Liu, Seenivasan Lalithkumar, Mobarakol Islam, Hongliang Ren, José M. M. Montiel, Danail Stoyanov
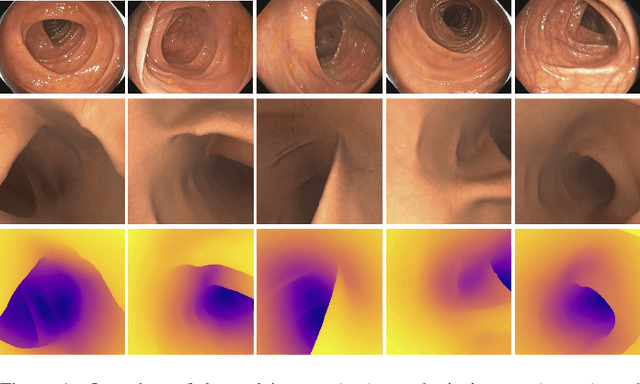
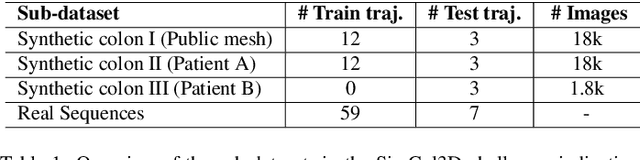
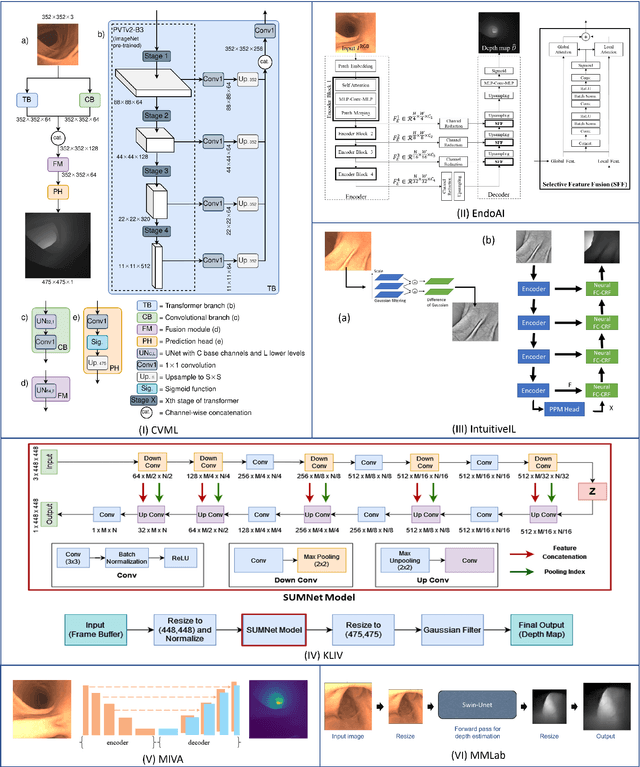
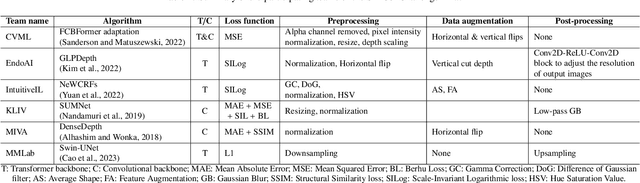
Colorectal cancer is one of the most common cancers in the world. While colonoscopy is an effective screening technique, navigating an endoscope through the colon to detect polyps is challenging. A 3D map of the observed surfaces could enhance the identification of unscreened colon tissue and serve as a training platform. However, reconstructing the colon from video footage remains unsolved due to numerous factors such as self-occlusion, reflective surfaces, lack of texture, and tissue deformation that limit feature-based methods. Learning-based approaches hold promise as robust alternatives, but necessitate extensive datasets. By establishing a benchmark, the 2022 EndoVis sub-challenge SimCol3D aimed to facilitate data-driven depth and pose prediction during colonoscopy. The challenge was hosted as part of MICCAI 2022 in Singapore. Six teams from around the world and representatives from academia and industry participated in the three sub-challenges: synthetic depth prediction, synthetic pose prediction, and real pose prediction. This paper describes the challenge, the submitted methods, and their results. We show that depth prediction in virtual colonoscopy is robustly solvable, while pose estimation remains an open research question.
Why is the winner the best?
Mar 30, 2023Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Sharib Ali, Vincent Andrearczyk, Marc Aubreville, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Veronika Cheplygina, Marie Daum, Marleen de Bruijne, Adrien Depeursinge, Reuben Dorent, Jan Egger, David G. Ellis, Sandy Engelhardt, Melanie Ganz, Noha Ghatwary, Gabriel Girard, Patrick Godau, Anubha Gupta, Lasse Hansen, Kanako Harada, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Pierre Jannin, Ali Emre Kavur, Oldřich Kodym, Michal Kozubek, Jianning Li, Hongwei Li, Jun Ma, Carlos Martín-Isla, Bjoern Menze, Alison Noble, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Tim Rädsch, Jonathan Rafael-Patiño, Vivek Singh Bawa, Stefanie Speidel, Carole H. Sudre, Kimberlin van Wijnen, Martin Wagner, Donglai Wei, Amine Yamlahi, Moi Hoon Yap, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Dogu Baran Aydogan, Binod Bhattarai, Louise Bloch, Raphael Brüngel, Jihoon Cho, Chanyeol Choi, Qi Dou, Ivan Ezhov, Christoph M. Friedrich, Clifton Fuller, Rebati Raman Gaire, Adrian Galdran, Álvaro García Faura, Maria Grammatikopoulou, SeulGi Hong, Mostafa Jahanifar, Ikbeom Jang, Abdolrahim Kadkhodamohammadi, Inha Kang, Florian Kofler, Satoshi Kondo, Hugo Kuijf, Mingxing Li, Minh Huan Luu, Tomaž Martinčič, Pedro Morais, Mohamed A. Naser, Bruno Oliveira, David Owen, Subeen Pang, Jinah Park, Sung-Hong Park, Szymon Płotka, Elodie Puybareau, Nasir Rajpoot, Kanghyun Ryu, Numan Saeed, Adam Shephard, Pengcheng Shi, Dejan Štepec, Ronast Subedi, Guillaume Tochon, Helena R. Torres, Helene Urien, João L. Vilaça, Kareem Abdul Wahid, Haojie Wang, Jiacheng Wang, Liansheng Wang, Xiyue Wang, Benedikt Wiestler, Marek Wodzinski, Fangfang Xia, Juanying Xie, Zhiwei Xiong, Sen Yang, Yanwu Yang, Zixuan Zhao, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
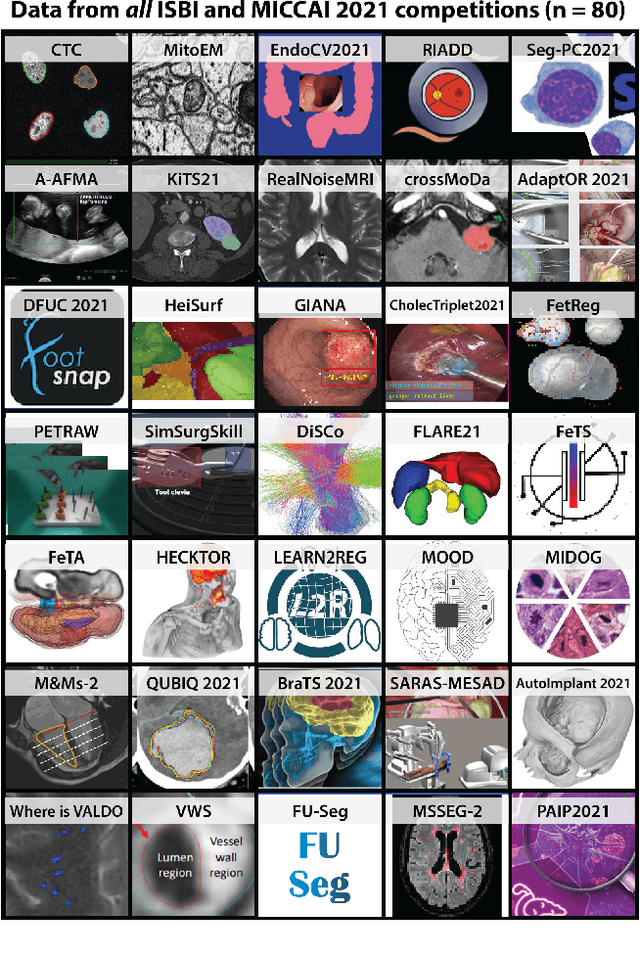
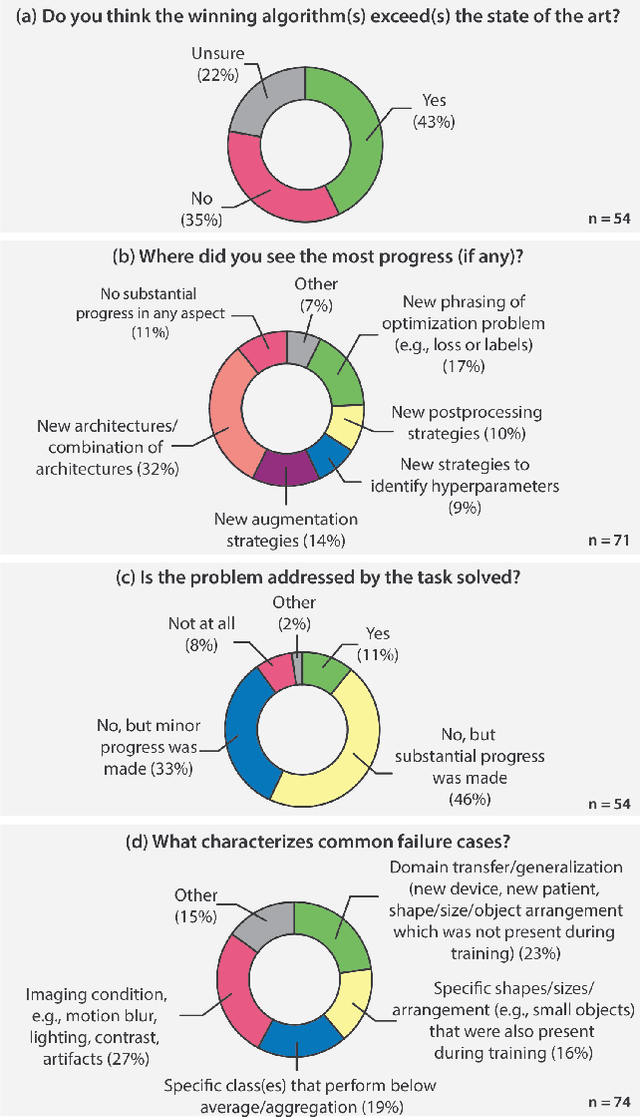
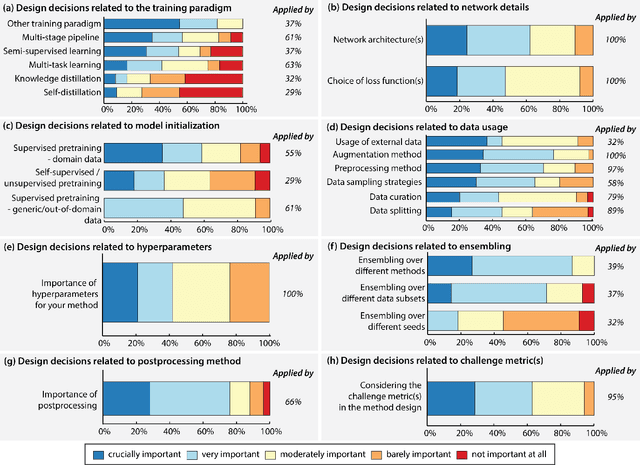
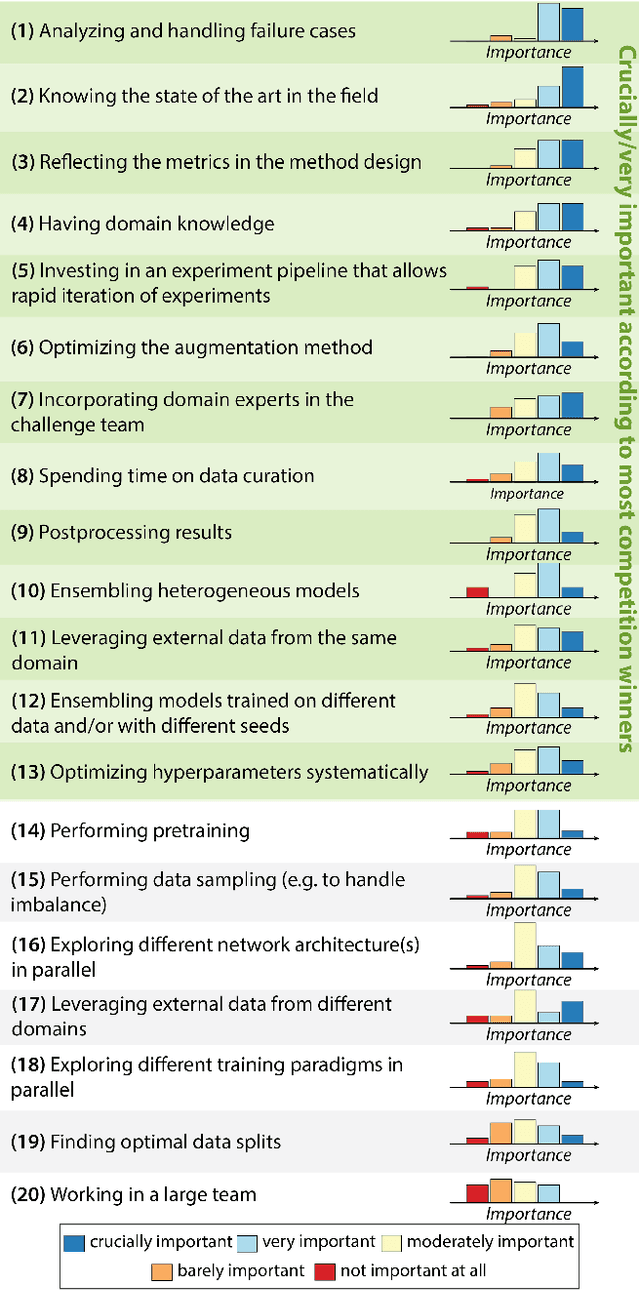
International benchmarking competitions have become fundamental for the comparative performance assessment of image analysis methods. However, little attention has been given to investigating what can be learnt from these competitions. Do they really generate scientific progress? What are common and successful participation strategies? What makes a solution superior to a competing method? To address this gap in the literature, we performed a multi-center study with all 80 competitions that were conducted in the scope of IEEE ISBI 2021 and MICCAI 2021. Statistical analyses performed based on comprehensive descriptions of the submitted algorithms linked to their rank as well as the underlying participation strategies revealed common characteristics of winning solutions. These typically include the use of multi-task learning (63%) and/or multi-stage pipelines (61%), and a focus on augmentation (100%), image preprocessing (97%), data curation (79%), and postprocessing (66%). The "typical" lead of a winning team is a computer scientist with a doctoral degree, five years of experience in biomedical image analysis, and four years of experience in deep learning. Two core general development strategies stood out for highly-ranked teams: the reflection of the metrics in the method design and the focus on analyzing and handling failure cases. According to the organizers, 43% of the winning algorithms exceeded the state of the art but only 11% completely solved the respective domain problem. The insights of our study could help researchers (1) improve algorithm development strategies when approaching new problems, and (2) focus on open research questions revealed by this work.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Patrick Godau, Veronika Cheplygina, Michal Kozubek, Sharib Ali, Anubha Gupta, Jan Kybic, Alison Noble, Carlos Ortiz de Solórzano, Samiksha Pachade, Caroline Petitjean, Daniel Sage, Donglai Wei, Elizabeth Wilden, Deepak Alapatt, Vincent Andrearczyk, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Vivek Singh Bawa, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Jinwook Choi, Olivier Commowick, Marie Daum, Adrien Depeursinge, Reuben Dorent, Jan Egger, Hannah Eichhorn, Sandy Engelhardt, Melanie Ganz, Gabriel Girard, Lasse Hansen, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Hyunjeong Kim, Bennett Landman, Hongwei Bran Li, Jianning Li, Jun Ma, Anne Martel, Carlos Martín-Isla, Bjoern Menze, Chinedu Innocent Nwoye, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Carole Sudre, Kimberlin van Wijnen, Armine Vardazaryan, Tom Vercauteren, Martin Wagner, Chuanbo Wang, Moi Hoon Yap, Zeyun Yu, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Rina Bao, Chanyeol Choi, Andrew Cohen, Oleh Dzyubachyk, Adrian Galdran, Tianyuan Gan, Tianqi Guo, Pradyumna Gupta, Mahmood Haithami, Edward Ho, Ikbeom Jang, Zhili Li, Zhengbo Luo, Filip Lux, Sokratis Makrogiannis, Dominik Müller, Young-tack Oh, Subeen Pang, Constantin Pape, Gorkem Polat, Charlotte Rosalie Reed, Kanghyun Ryu, Tim Scherr, Vajira Thambawita, Haoyu Wang, Xinliang Wang, Kele Xu, Hung Yeh, Doyeob Yeo, Yixuan Yuan, Yan Zeng, Xin Zhao, Julian Abbing, Jannes Adam, Nagesh Adluru, Niklas Agethen, Salman Ahmed, Yasmina Al Khalil, Mireia Alenyà, Esa Alhoniemi, Chengyang An, Talha Anwar, Tewodros Weldebirhan Arega, Netanell Avisdris, Dogu Baran Aydogan, Yingbin Bai, Maria Baldeon Calisto, Berke Doga Basaran, Marcel Beetz, Cheng Bian, Hao Bian, Kevin Blansit, Louise Bloch, Robert Bohnsack, Sara Bosticardo, Jack Breen, Mikael Brudfors, Raphael Brüngel, Mariano Cabezas, Alberto Cacciola, Zhiwei Chen, Yucong Chen, Daniel Tianming Chen, Minjeong Cho, Min-Kook Choi, Chuantao Xie Chuantao Xie, Dana Cobzas, Julien Cohen-Adad, Jorge Corral Acero, Sujit Kumar Das, Marcela de Oliveira, Hanqiu Deng, Guiming Dong, Lars Doorenbos, Cory Efird, Di Fan, Mehdi Fatan Serj, Alexandre Fenneteau, Lucas Fidon, Patryk Filipiak, René Finzel, Nuno R. Freitas, Christoph M. Friedrich, Mitchell Fulton, Finn Gaida, Francesco Galati, Christoforos Galazis, Chang Hee Gan, Zheyao Gao, Shengbo Gao, Matej Gazda, Beerend Gerats, Neil Getty, Adam Gibicar, Ryan Gifford, Sajan Gohil, Maria Grammatikopoulou, Daniel Grzech, Orhun Güley, Timo Günnemann, Chunxu Guo, Sylvain Guy, Heonjin Ha, Luyi Han, Il Song Han, Ali Hatamizadeh, Tian He, Jimin Heo, Sebastian Hitziger, SeulGi Hong, SeungBum Hong, Rian Huang, Ziyan Huang, Markus Huellebrand, Stephan Huschauer, Mustaffa Hussain, Tomoo Inubushi, Ece Isik Polat, Mojtaba Jafaritadi, SeongHun Jeong, Bailiang Jian, Yuanhong Jiang, Zhifan Jiang, Yueming Jin, Smriti Joshi, Abdolrahim Kadkhodamohammadi, Reda Abdellah Kamraoui, Inha Kang, Junghwa Kang, Davood Karimi, April Khademi, Muhammad Irfan Khan, Suleiman A. Khan, Rishab Khantwal, Kwang-Ju Kim, Timothy Kline, Satoshi Kondo, Elina Kontio, Adrian Krenzer, Artem Kroviakov, Hugo Kuijf, Satyadwyoom Kumar, Francesco La Rosa, Abhi Lad, Doohee Lee, Minho Lee, Chiara Lena, Hao Li, Ling Li, Xingyu Li, Fuyuan Liao, KuanLun Liao, Arlindo Limede Oliveira, Chaonan Lin, Shan Lin, Akis Linardos, Marius George Linguraru, Han Liu, Tao Liu, Di Liu, Yanling Liu, João Lourenço-Silva, Jingpei Lu, Jiangshan Lu, Imanol Luengo, Christina B. Lund, Huan Minh Luu, Yi Lv, Uzay Macar, Leon Maechler, Sina Mansour L., Kenji Marshall, Moona Mazher, Richard McKinley, Alfonso Medela, Felix Meissen, Mingyuan Meng, Dylan Miller, Seyed Hossein Mirjahanmardi, Arnab Mishra, Samir Mitha, Hassan Mohy-ud-Din, Tony Chi Wing Mok, Gowtham Krishnan Murugesan, Enamundram Naga Karthik, Sahil Nalawade, Jakub Nalepa, Mohamed Naser, Ramin Nateghi, Hammad Naveed, Quang-Minh Nguyen, Cuong Nguyen Quoc, Brennan Nichyporuk, Bruno Oliveira, David Owen, Jimut Bahan Pal, Junwen Pan, Wentao Pan, Winnie Pang, Bogyu Park, Vivek Pawar, Kamlesh Pawar, Michael Peven, Lena Philipp, Tomasz Pieciak, Szymon Plotka, Marcel Plutat, Fattaneh Pourakpour, Domen Preložnik, Kumaradevan Punithakumar, Abdul Qayyum, Sandro Queirós, Arman Rahmim, Salar Razavi, Jintao Ren, Mina Rezaei, Jonathan Adam Rico, ZunHyan Rieu, Markus Rink, Johannes Roth, Yusely Ruiz-Gonzalez, Numan Saeed, Anindo Saha, Mostafa Salem, Ricardo Sanchez-Matilla, Kurt Schilling, Wei Shao, Zhiqiang Shen, Ruize Shi, Pengcheng Shi, Daniel Sobotka, Théodore Soulier, Bella Specktor Fadida, Danail Stoyanov, Timothy Sum Hon Mun, Xiaowu Sun, Rong Tao, Franz Thaler, Antoine Théberge, Felix Thielke, Helena Torres, Kareem A. Wahid, Jiacheng Wang, YiFei Wang, Wei Wang, Xiong Wang, Jianhui Wen, Ning Wen, Marek Wodzinski, Ye Wu, Fangfang Xia, Tianqi Xiang, Chen Xiaofei, Lizhan Xu, Tingting Xue, Yuxuan Yang, Lin Yang, Kai Yao, Huifeng Yao, Amirsaeed Yazdani, Michael Yip, Hwanseung Yoo, Fereshteh Yousefirizi, Shunkai Yu, Lei Yu, Jonathan Zamora, Ramy Ashraf Zeineldin, Dewen Zeng, Jianpeng Zhang, Bokai Zhang, Jiapeng Zhang, Fan Zhang, Huahong Zhang, Zhongchen Zhao, Zixuan Zhao, Jiachen Zhao, Can Zhao, Qingshuo Zheng, Yuheng Zhi, Ziqi Zhou, Baosheng Zou, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Retrieval of surgical phase transitions using reinforcement learning
Aug 01, 2022Yitong Zhang, Sophia Bano, Ann-Sophie Page, Jan Deprest, Danail Stoyanov, Francisco Vasconcelos
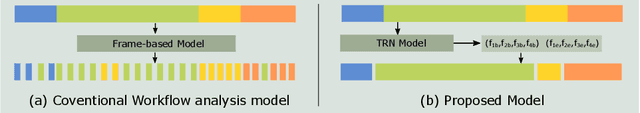
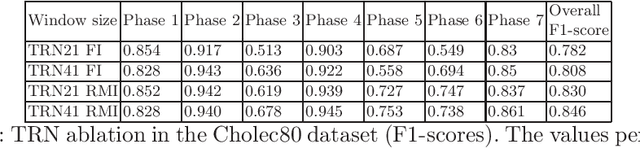
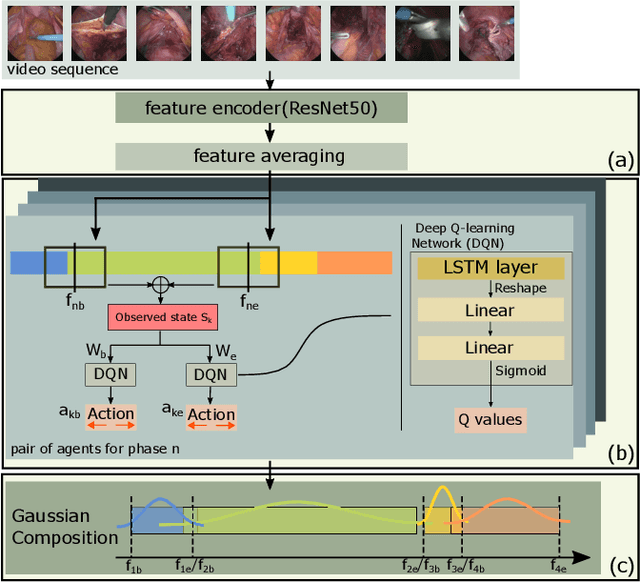
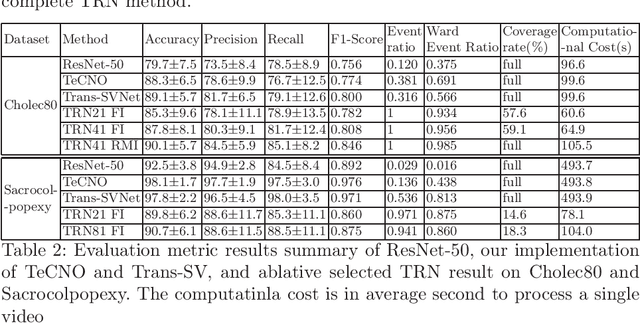
In minimally invasive surgery, surgical workflow segmentation from video analysis is a well studied topic. The conventional approach defines it as a multi-class classification problem, where individual video frames are attributed a surgical phase label. We introduce a novel reinforcement learning formulation for offline phase transition retrieval. Instead of attempting to classify every video frame, we identify the timestamp of each phase transition. By construction, our model does not produce spurious and noisy phase transitions, but contiguous phase blocks. We investigate two different configurations of this model. The first does not require processing all frames in a video (only <60% and <20% of frames in 2 different applications), while producing results slightly under the state-of-the-art accuracy. The second configuration processes all video frames, and outperforms the state-of-the art at a comparable computational cost. We compare our method against the recent top-performing frame-based approaches TeCNO and Trans-SVNet on the public dataset Cholec80 and also on an in-house dataset of laparoscopic sacrocolpopexy. We perform both a frame-based (accuracy, precision, recall and F1-score) and an event-based (event ratio) evaluation of our algorithms.
Learning-Based Keypoint Registration for Fetoscopic Mosaicking
Jul 26, 2022Alessandro Casella, Sophia Bano, Francisco Vasconcelos, Anna L. David, Dario Paladini, Jan Deprest, Elena De Momi, Leonardo S. Mattos, Sara Moccia, Danail Stoyanov
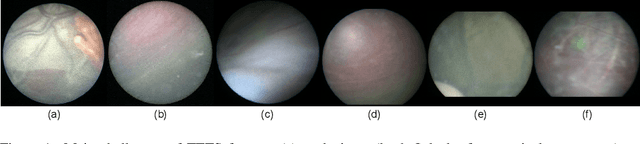
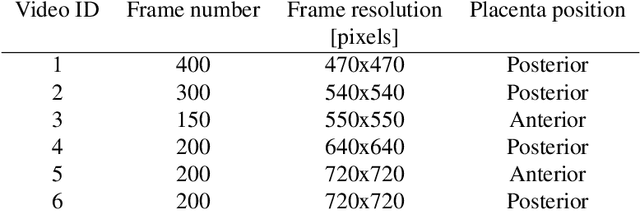
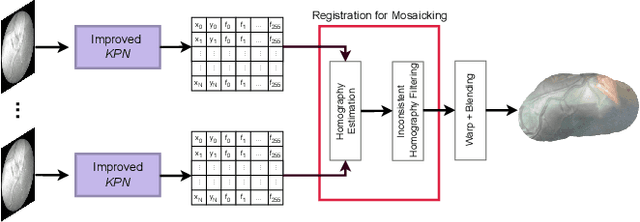

In Twin-to-Twin Transfusion Syndrome (TTTS), abnormal vascular anastomoses in the monochorionic placenta can produce uneven blood flow between the two fetuses. In the current practice, TTTS is treated surgically by closing abnormal anastomoses using laser ablation. This surgery is minimally invasive and relies on fetoscopy. Limited field of view makes anastomosis identification a challenging task for the surgeon. To tackle this challenge, we propose a learning-based framework for in-vivo fetoscopy frame registration for field-of-view expansion. The novelties of this framework relies on a learning-based keypoint proposal network and an encoding strategy to filter (i) irrelevant keypoints based on fetoscopic image segmentation and (ii) inconsistent homographies. We validate of our framework on a dataset of 6 intraoperative sequences from 6 TTTS surgeries from 6 different women against the most recent state of the art algorithm, which relies on the segmentation of placenta vessels. The proposed framework achieves higher performance compared to the state of the art, paving the way for robust mosaicking to provide surgeons with context awareness during TTTS surgery.
FetReg2021: A Challenge on Placental Vessel Segmentation and Registration in Fetoscopy
Jun 30, 2022Sophia Bano, Alessandro Casella, Francisco Vasconcelos, Abdul Qayyum, Abdesslam Benzinou, Moona Mazher, Fabrice Meriaudeau, Chiara Lena, Ilaria Anita Cintorrino, Gaia Romana De Paolis, Jessica Biagioli, Daria Grechishnikova, Jing Jiao, Bizhe Bai, Yanyan Qiao, Binod Bhattarai, Rebati Raman Gaire, Ronast Subedi, Eduard Vazquez, Szymon Płotka, Aneta Lisowska, Arkadiusz Sitek, George Attilakos, Ruwan Wimalasundera, Anna L David, Dario Paladini, Jan Deprest, Elena De Momi, Leonardo S Mattos, Sara Moccia, Danail Stoyanov
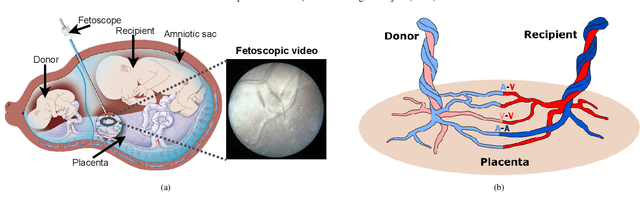
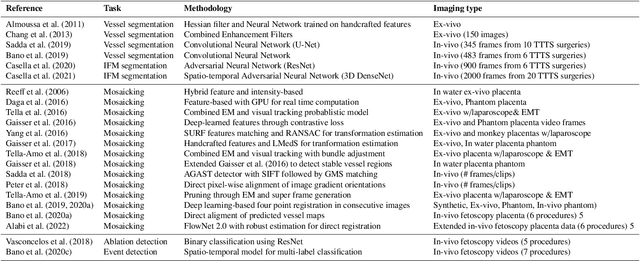
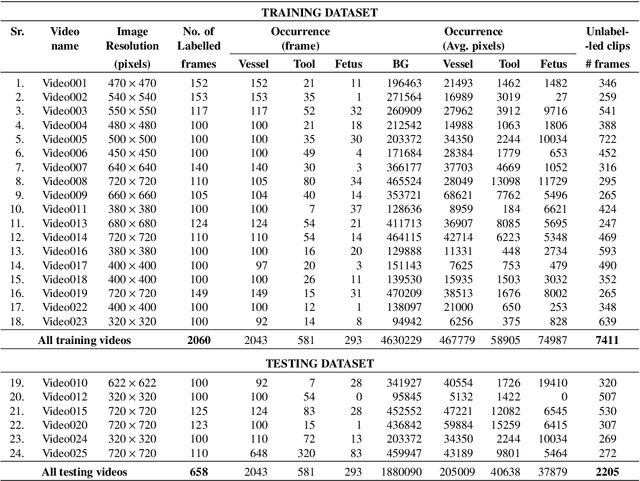
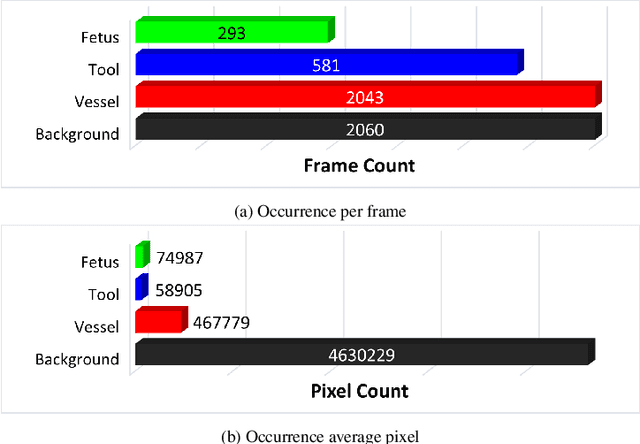
Fetoscopy laser photocoagulation is a widely adopted procedure for treating Twin-to-Twin Transfusion Syndrome (TTTS). The procedure involves photocoagulation pathological anastomoses to regulate blood exchange among twins. The procedure is particularly challenging due to the limited field of view, poor manoeuvrability of the fetoscope, poor visibility, and variability in illumination. These challenges may lead to increased surgery time and incomplete ablation. Computer-assisted intervention (CAI) can provide surgeons with decision support and context awareness by identifying key structures in the scene and expanding the fetoscopic field of view through video mosaicking. Research in this domain has been hampered by the lack of high-quality data to design, develop and test CAI algorithms. Through the Fetoscopic Placental Vessel Segmentation and Registration (FetReg2021) challenge, which was organized as part of the MICCAI2021 Endoscopic Vision challenge, we released the first largescale multicentre TTTS dataset for the development of generalized and robust semantic segmentation and video mosaicking algorithms. For this challenge, we released a dataset of 2060 images, pixel-annotated for vessels, tool, fetus and background classes, from 18 in-vivo TTTS fetoscopy procedures and 18 short video clips. Seven teams participated in this challenge and their model performance was assessed on an unseen test dataset of 658 pixel-annotated images from 6 fetoscopic procedures and 6 short clips. The challenge provided an opportunity for creating generalized solutions for fetoscopic scene understanding and mosaicking. In this paper, we present the findings of the FetReg2021 challenge alongside reporting a detailed literature review for CAI in TTTS fetoscopy. Through this challenge, its analysis and the release of multi-centre fetoscopic data, we provide a benchmark for future research in this field.
BiometryNet: Landmark-based Fetal Biometry Estimation from Standard Ultrasound Planes
Jun 29, 2022Netanell Avisdris, Leo Joskowicz, Brian Dromey, Anna L. David, Donald M. Peebles, Danail Stoyanov, Dafna Ben Bashat, Sophia Bano
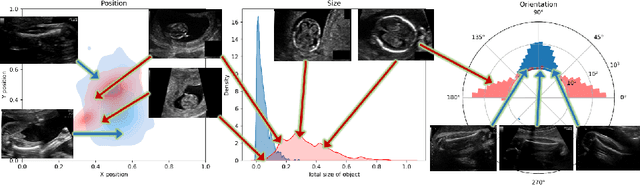
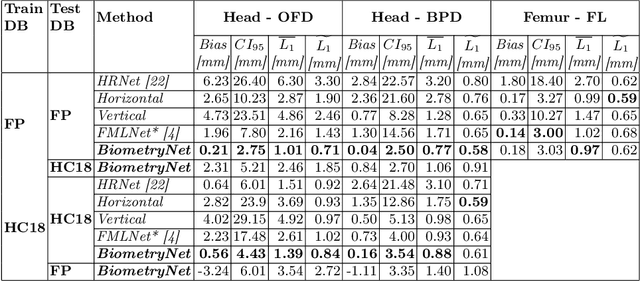
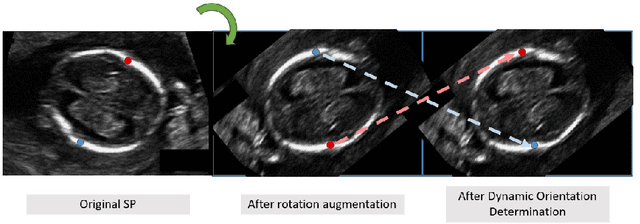
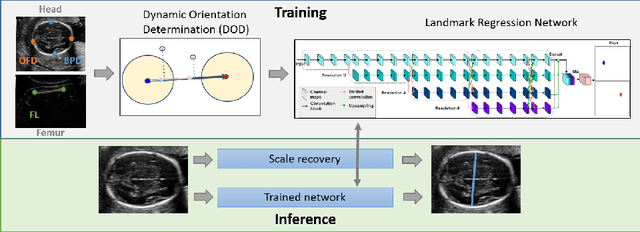
Fetal growth assessment from ultrasound is based on a few biometric measurements that are performed manually and assessed relative to the expected gestational age. Reliable biometry estimation depends on the precise detection of landmarks in standard ultrasound planes. Manual annotation can be time-consuming and operator dependent task, and may results in high measurements variability. Existing methods for automatic fetal biometry rely on initial automatic fetal structure segmentation followed by geometric landmark detection. However, segmentation annotations are time-consuming and may be inaccurate, and landmark detection requires developing measurement-specific geometric methods. This paper describes BiometryNet, an end-to-end landmark regression framework for fetal biometry estimation that overcomes these limitations. It includes a novel Dynamic Orientation Determination (DOD) method for enforcing measurement-specific orientation consistency during network training. DOD reduces variabilities in network training, increases landmark localization accuracy, thus yields accurate and robust biometric measurements. To validate our method, we assembled a dataset of 3,398 ultrasound images from 1,829 subjects acquired in three clinical sites with seven different ultrasound devices. Comparison and cross-validation of three different biometric measurements on two independent datasets shows that BiometryNet is robust and yields accurate measurements whose errors are lower than the clinically permissible errors, outperforming other existing automated biometry estimation methods. Code is available at https://github.com/netanellavisdris/fetalbiometry.
Fluorescence angiography classification in colorectal surgery -- A preliminary report
Jun 13, 2022Antonio S Soares, Sophia Bano, Neil T Clancy, Laurence B Lovat, Danail Stoyanov, Manish Chand
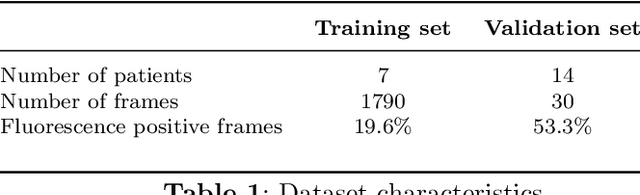

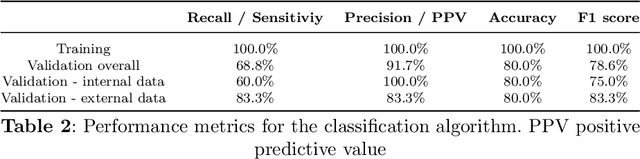
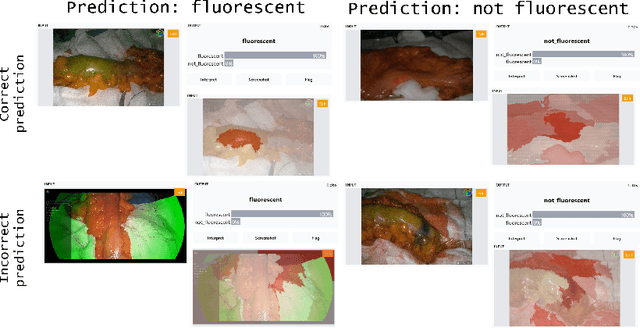
Background: Fluorescence angiography has shown very promising results in reducing anastomotic leaks by allowing the surgeon to select optimally perfused tissue. However, subjective interpretation of the fluorescent signal still hinders broad application of the technique, as significant variation between different surgeons exists. Our aim is to develop an artificial intelligence algorithm to classify colonic tissue as 'perfused' or 'not perfused' based on intraoperative fluorescence angiography data. Methods: A classification model with a Resnet architecture was trained on a dataset of fluorescence angiography videos of colorectal resections at a tertiary referral centre. Frames corresponding to fluorescent and non-fluorescent segments of colon were used to train a classification algorithm. Validation using frames from patients not used in the training set was performed, including both data collected using the same equipment and data collected using a different camera. Performance metrics were calculated, and saliency maps used to further analyse the output. A decision boundary was identified based on the tissue classification. Results: A convolutional neural network was successfully trained on 1790 frames from 7 patients and validated in 24 frames from 14 patients. The accuracy on the training set was 100%, on the validation set was 80%. Recall and precision were respectively 100% and 100% on the training set and 68.8% and 91.7% on the validation set. Conclusion: Automated classification of intraoperative fluorescence angiography with a high degree of accuracy is possible and allows automated decision boundary identification. This will enable surgeons to standardise the technique of fluorescence angiography. A web based app was made available to deploy the algorithm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge