A Split-and-Privatize Framework for Large Language Model Fine-Tuning
Dec 25, 2023Xicong Shen, Yang Liu, Huiqi Liu, Jue Hong, Bing Duan, Zirui Huang, Yunlong Mao, Ye Wu, Di Wu
Fine-tuning is a prominent technique to adapt a pre-trained language model to downstream scenarios. In parameter-efficient fine-tuning, only a small subset of modules are trained over the downstream datasets, while leaving the rest of the pre-trained model frozen to save computation resources. In recent years, a popular productization form arises as Model-as-a-Service (MaaS), in which vendors provide abundant pre-trained language models, server resources and core functions, and customers can fine-tune, deploy and invoke their customized model by accessing the one-stop MaaS with their own private dataset. In this paper, we identify the model and data privacy leakage risks in MaaS fine-tuning, and propose a Split-and-Privatize (SAP) framework, which manage to mitigate the privacy issues by adapting the existing split learning architecture. The proposed SAP framework is sufficiently investigated by experiments, and the results indicate that it can enhance the empirical privacy by 62% at the cost of 1% model performance degradation on the Stanford Sentiment Treebank dataset.
Segment Anything Model-guided Collaborative Learning Network for Scribble-supervised Polyp Segmentation
Dec 01, 2023Yiming Zhao, Tao Zhou, Yunqi Gu, Yi Zhou, Yizhe Zhang, Ye Wu, Huazhu Fu
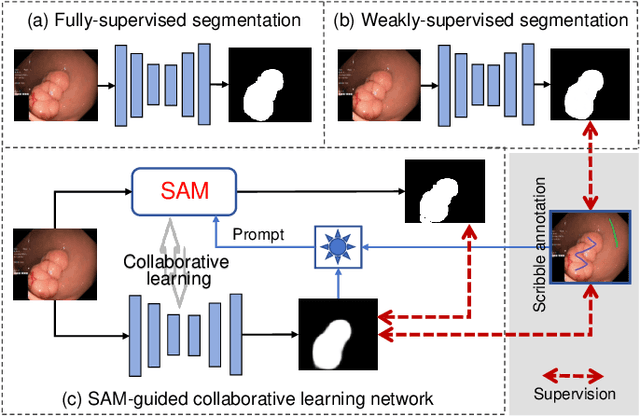
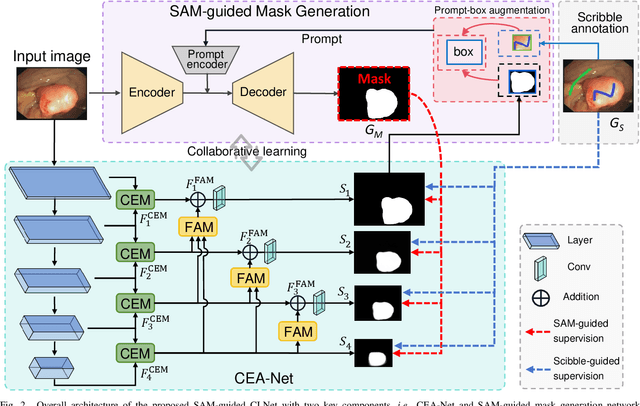
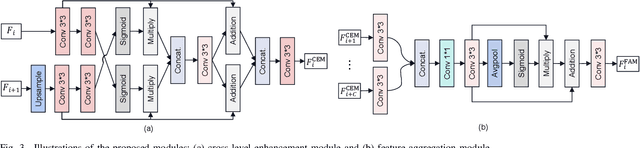
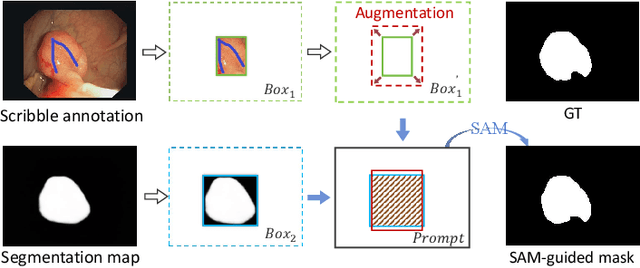
Polyp segmentation plays a vital role in accurately locating polyps at an early stage, which holds significant clinical importance for the prevention of colorectal cancer. Various polyp segmentation methods have been developed using fully-supervised deep learning techniques. However, pixel-wise annotation for polyp images by physicians during the diagnosis is both time-consuming and expensive. Moreover, visual foundation models such as the Segment Anything Model (SAM) have shown remarkable performance. Nevertheless, directly applying SAM to medical segmentation may not produce satisfactory results due to the inherent absence of medical knowledge. In this paper, we propose a novel SAM-guided Collaborative Learning Network (SAM-CLNet) for scribble-supervised polyp segmentation, enabling a collaborative learning process between our segmentation network and SAM to boost the model performance. Specifically, we first propose a Cross-level Enhancement and Aggregation Network (CEA-Net) for weakly-supervised polyp segmentation. Within CEA-Net, we propose a Cross-level Enhancement Module (CEM) that integrates the adjacent features to enhance the representation capabilities of different resolution features. Additionally, a Feature Aggregation Module (FAM) is employed to capture richer features across multiple levels. Moreover, we present a box-augmentation strategy that combines the segmentation maps generated by CEA-Net with scribble annotations to create more precise prompts. These prompts are then fed into SAM, generating segmentation SAM-guided masks, which can provide additional supervision to train CEA-Net effectively. Furthermore, we present an Image-level Filtering Mechanism to filter out unreliable SAM-guided masks. Extensive experimental results show that our SAM-CLNet outperforms state-of-the-art weakly-supervised segmentation methods.
A Survey on Deep Learning for Polyp Segmentation: Techniques, Challenges and Future Trends
Nov 30, 2023Jiaxin Mei, Tao Zhou, Kaiwen Huang, Yizhe Zhang, Yi Zhou, Ye Wu, Huazhu Fu
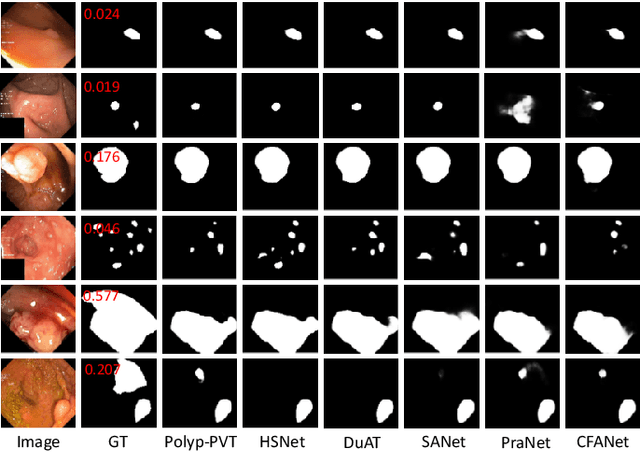
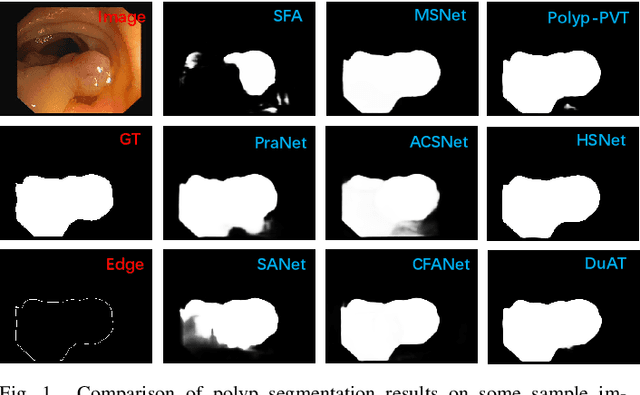
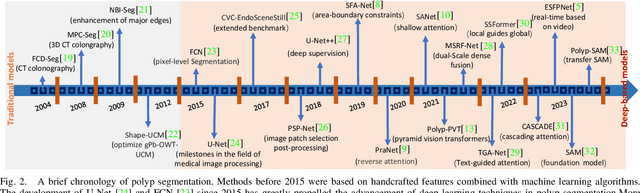
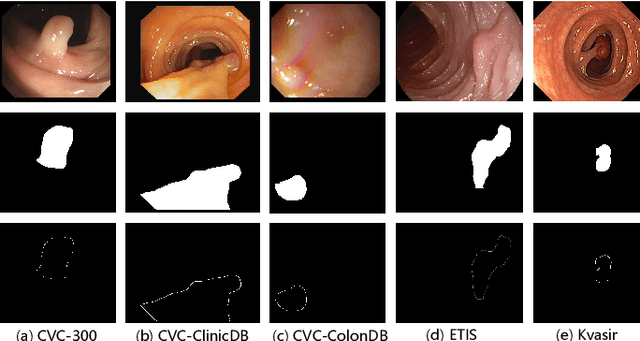

Early detection and assessment of polyps play a crucial role in the prevention and treatment of colorectal cancer (CRC). Polyp segmentation provides an effective solution to assist clinicians in accurately locating and segmenting polyp regions. In the past, people often relied on manually extracted lower-level features such as color, texture, and shape, which often had issues capturing global context and lacked robustness to complex scenarios. With the advent of deep learning, more and more outstanding medical image segmentation algorithms based on deep learning networks have emerged, making significant progress in this field. This paper provides a comprehensive review of polyp segmentation algorithms. We first review some traditional algorithms based on manually extracted features and deep segmentation algorithms, then detail benchmark datasets related to the topic. Specifically, we carry out a comprehensive evaluation of recent deep learning models and results based on polyp sizes, considering the pain points of research topics and differences in network structures. Finally, we discuss the challenges of polyp segmentation and future trends in this field. The models, benchmark datasets, and source code links we collected are all published at https://github.com/taozh2017/Awesome-Polyp-Segmentation.
Hawkeye: A PyTorch-based Library for Fine-Grained Image Recognition with Deep Learning
Oct 14, 2023Jiabei He, Yang Shen, Xiu-Shen Wei, Ye Wu
Fine-Grained Image Recognition (FGIR) is a fundamental and challenging task in computer vision and multimedia that plays a crucial role in Intellectual Economy and Industrial Internet applications. However, the absence of a unified open-source software library covering various paradigms in FGIR poses a significant challenge for researchers and practitioners in the field. To address this gap, we present Hawkeye, a PyTorch-based library for FGIR with deep learning. Hawkeye is designed with a modular architecture, emphasizing high-quality code and human-readable configuration, providing a comprehensive solution for FGIR tasks. In Hawkeye, we have implemented 16 state-of-the-art fine-grained methods, covering 6 different paradigms, enabling users to explore various approaches for FGIR. To the best of our knowledge, Hawkeye represents the first open-source PyTorch-based library dedicated to FGIR. It is publicly available at https://github.com/Hawkeye-FineGrained/Hawkeye/, providing researchers and practitioners with a powerful tool to advance their research and development in the field of FGIR.
Edge-aware Feature Aggregation Network for Polyp Segmentation
Sep 19, 2023Tao Zhou, Yizhe Zhang, Geng Chen, Yi Zhou, Ye Wu, Deng-Ping Fan




Precise polyp segmentation is vital for the early diagnosis and prevention of colorectal cancer (CRC) in clinical practice. However, due to scale variation and blurry polyp boundaries, it is still a challenging task to achieve satisfactory segmentation performance with different scales and shapes. In this study, we present a novel Edge-aware Feature Aggregation Network (EFA-Net) for polyp segmentation, which can fully make use of cross-level and multi-scale features to enhance the performance of polyp segmentation. Specifically, we first present an Edge-aware Guidance Module (EGM) to combine the low-level features with the high-level features to learn an edge-enhanced feature, which is incorporated into each decoder unit using a layer-by-layer strategy. Besides, a Scale-aware Convolution Module (SCM) is proposed to learn scale-aware features by using dilated convolutions with different ratios, in order to effectively deal with scale variation. Further, a Cross-level Fusion Module (CFM) is proposed to effectively integrate the cross-level features, which can exploit the local and global contextual information. Finally, the outputs of CFMs are adaptively weighted by using the learned edge-aware feature, which are then used to produce multiple side-out segmentation maps. Experimental results on five widely adopted colonoscopy datasets show that our EFA-Net outperforms state-of-the-art polyp segmentation methods in terms of generalization and effectiveness.
VideoGen: A Reference-Guided Latent Diffusion Approach for High Definition Text-to-Video Generation
Sep 07, 2023Xin Li, Wenqing Chu, Ye Wu, Weihang Yuan, Fanglong Liu, Qi Zhang, Fu Li, Haocheng Feng, Errui Ding, Jingdong Wang
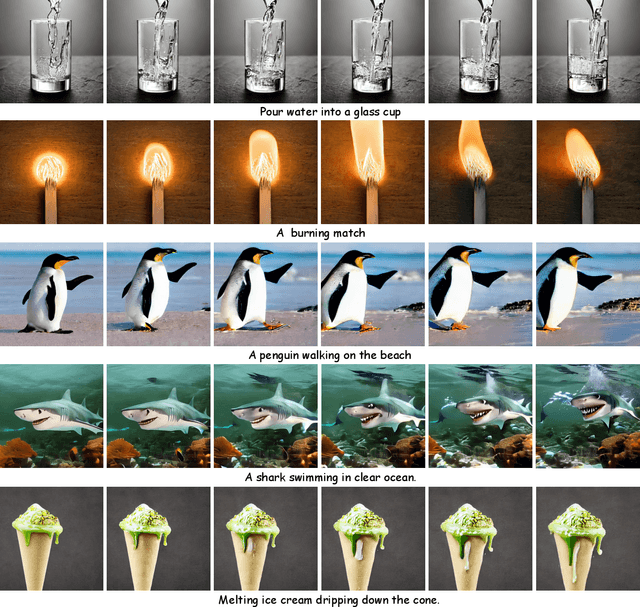

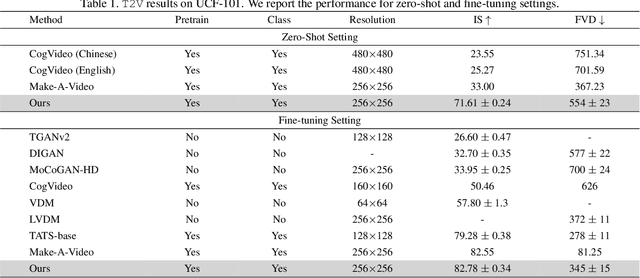
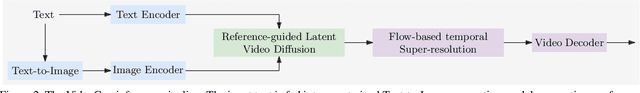
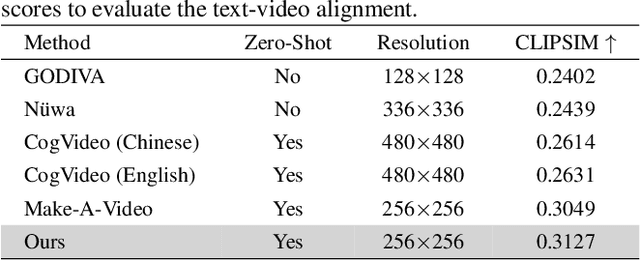
In this paper, we present VideoGen, a text-to-video generation approach, which can generate a high-definition video with high frame fidelity and strong temporal consistency using reference-guided latent diffusion. We leverage an off-the-shelf text-to-image generation model, e.g., Stable Diffusion, to generate an image with high content quality from the text prompt, as a reference image to guide video generation. Then, we introduce an efficient cascaded latent diffusion module conditioned on both the reference image and the text prompt, for generating latent video representations, followed by a flow-based temporal upsampling step to improve the temporal resolution. Finally, we map latent video representations into a high-definition video through an enhanced video decoder. During training, we use the first frame of a ground-truth video as the reference image for training the cascaded latent diffusion module. The main characterises of our approach include: the reference image generated by the text-to-image model improves the visual fidelity; using it as the condition makes the diffusion model focus more on learning the video dynamics; and the video decoder is trained over unlabeled video data, thus benefiting from high-quality easily-available videos. VideoGen sets a new state-of-the-art in text-to-video generation in terms of both qualitative and quantitative evaluation. See \url{https://videogen.github.io/VideoGen/} for more samples.
SamDSK: Combining Segment Anything Model with Domain-Specific Knowledge for Semi-Supervised Learning in Medical Image Segmentation
Aug 26, 2023Yizhe Zhang, Tao Zhou, Shuo Wang, Ye Wu, Pengfei Gu, Danny Z. Chen




The Segment Anything Model (SAM) exhibits a capability to segment a wide array of objects in natural images, serving as a versatile perceptual tool for various downstream image segmentation tasks. In contrast, medical image segmentation tasks often rely on domain-specific knowledge (DSK). In this paper, we propose a novel method that combines the segmentation foundation model (i.e., SAM) with domain-specific knowledge for reliable utilization of unlabeled images in building a medical image segmentation model. Our new method is iterative and consists of two main stages: (1) segmentation model training; (2) expanding the labeled set by using the trained segmentation model, an unlabeled set, SAM, and domain-specific knowledge. These two stages are repeated until no more samples are added to the labeled set. A novel optimal-matching-based method is developed for combining the SAM-generated segmentation proposals and pixel-level and image-level DSK for constructing annotations of unlabeled images in the iterative stage (2). In experiments, we demonstrate the effectiveness of our proposed method for breast cancer segmentation in ultrasound images, polyp segmentation in endoscopic images, and skin lesion segmentation in dermoscopic images. Our work initiates a new direction of semi-supervised learning for medical image segmentation: the segmentation foundation model can be harnessed as a valuable tool for label-efficient segmentation learning in medical image segmentation.
Can SAM Segment Polyps?
Apr 15, 2023Tao Zhou, Yizhe Zhang, Yi Zhou, Ye Wu, Chen Gong




Recently, Meta AI Research releases a general Segment Anything Model (SAM), which has demonstrated promising performance in several segmentation tasks. As we know, polyp segmentation is a fundamental task in the medical imaging field, which plays a critical role in the diagnosis and cure of colorectal cancer. In particular, applying SAM to the polyp segmentation task is interesting. In this report, we evaluate the performance of SAM in segmenting polyps, in which SAM is under unprompted settings. We hope this report will provide insights to advance this polyp segmentation field and promote more interesting works in the future. This project is publicly at https://github.com/taozh2017/SAMPolyp.
Brain Tissue Segmentation Across the Human Lifespan via Supervised Contrastive Learning
Jan 03, 2023Xiaoyang Chen, Jinjian Wu, Wenjiao Lyu, Yicheng Zou, Kim-Han Thung, Siyuan Liu, Ye Wu, Sahar Ahmad, Pew-Thian Yap
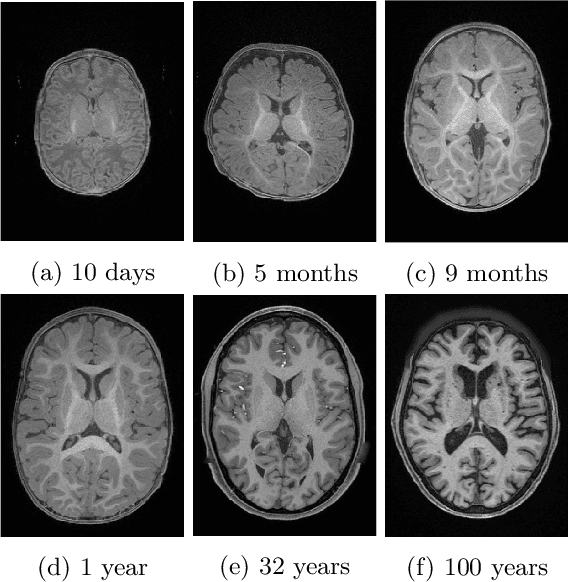
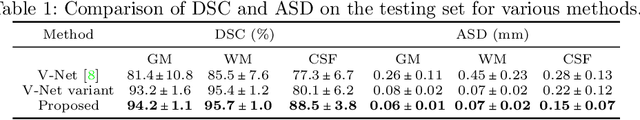
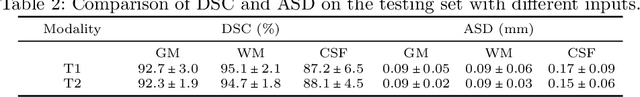
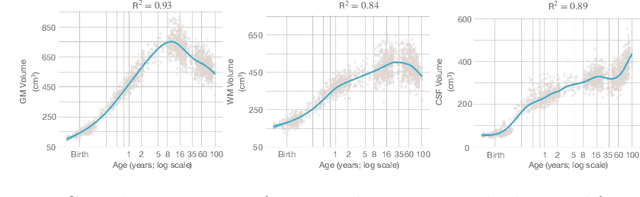
Automatic segmentation of brain MR images into white matter (WM), gray matter (GM), and cerebrospinal fluid (CSF) is critical for tissue volumetric analysis and cortical surface reconstruction. Due to dramatic structural and appearance changes associated with developmental and aging processes, existing brain tissue segmentation methods are only viable for specific age groups. Consequently, methods developed for one age group may fail for another. In this paper, we make the first attempt to segment brain tissues across the entire human lifespan (0-100 years of age) using a unified deep learning model. To overcome the challenges related to structural variability underpinned by biological processes, intensity inhomogeneity, motion artifacts, scanner-induced differences, and acquisition protocols, we propose to use contrastive learning to improve the quality of feature representations in a latent space for effective lifespan tissue segmentation. We compared our approach with commonly used segmentation methods on a large-scale dataset of 2,464 MR images. Experimental results show that our model accurately segments brain tissues across the lifespan and outperforms existing methods.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Patrick Godau, Veronika Cheplygina, Michal Kozubek, Sharib Ali, Anubha Gupta, Jan Kybic, Alison Noble, Carlos Ortiz de Solórzano, Samiksha Pachade, Caroline Petitjean, Daniel Sage, Donglai Wei, Elizabeth Wilden, Deepak Alapatt, Vincent Andrearczyk, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Vivek Singh Bawa, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Jinwook Choi, Olivier Commowick, Marie Daum, Adrien Depeursinge, Reuben Dorent, Jan Egger, Hannah Eichhorn, Sandy Engelhardt, Melanie Ganz, Gabriel Girard, Lasse Hansen, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Hyunjeong Kim, Bennett Landman, Hongwei Bran Li, Jianning Li, Jun Ma, Anne Martel, Carlos Martín-Isla, Bjoern Menze, Chinedu Innocent Nwoye, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Carole Sudre, Kimberlin van Wijnen, Armine Vardazaryan, Tom Vercauteren, Martin Wagner, Chuanbo Wang, Moi Hoon Yap, Zeyun Yu, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Rina Bao, Chanyeol Choi, Andrew Cohen, Oleh Dzyubachyk, Adrian Galdran, Tianyuan Gan, Tianqi Guo, Pradyumna Gupta, Mahmood Haithami, Edward Ho, Ikbeom Jang, Zhili Li, Zhengbo Luo, Filip Lux, Sokratis Makrogiannis, Dominik Müller, Young-tack Oh, Subeen Pang, Constantin Pape, Gorkem Polat, Charlotte Rosalie Reed, Kanghyun Ryu, Tim Scherr, Vajira Thambawita, Haoyu Wang, Xinliang Wang, Kele Xu, Hung Yeh, Doyeob Yeo, Yixuan Yuan, Yan Zeng, Xin Zhao, Julian Abbing, Jannes Adam, Nagesh Adluru, Niklas Agethen, Salman Ahmed, Yasmina Al Khalil, Mireia Alenyà, Esa Alhoniemi, Chengyang An, Talha Anwar, Tewodros Weldebirhan Arega, Netanell Avisdris, Dogu Baran Aydogan, Yingbin Bai, Maria Baldeon Calisto, Berke Doga Basaran, Marcel Beetz, Cheng Bian, Hao Bian, Kevin Blansit, Louise Bloch, Robert Bohnsack, Sara Bosticardo, Jack Breen, Mikael Brudfors, Raphael Brüngel, Mariano Cabezas, Alberto Cacciola, Zhiwei Chen, Yucong Chen, Daniel Tianming Chen, Minjeong Cho, Min-Kook Choi, Chuantao Xie Chuantao Xie, Dana Cobzas, Julien Cohen-Adad, Jorge Corral Acero, Sujit Kumar Das, Marcela de Oliveira, Hanqiu Deng, Guiming Dong, Lars Doorenbos, Cory Efird, Di Fan, Mehdi Fatan Serj, Alexandre Fenneteau, Lucas Fidon, Patryk Filipiak, René Finzel, Nuno R. Freitas, Christoph M. Friedrich, Mitchell Fulton, Finn Gaida, Francesco Galati, Christoforos Galazis, Chang Hee Gan, Zheyao Gao, Shengbo Gao, Matej Gazda, Beerend Gerats, Neil Getty, Adam Gibicar, Ryan Gifford, Sajan Gohil, Maria Grammatikopoulou, Daniel Grzech, Orhun Güley, Timo Günnemann, Chunxu Guo, Sylvain Guy, Heonjin Ha, Luyi Han, Il Song Han, Ali Hatamizadeh, Tian He, Jimin Heo, Sebastian Hitziger, SeulGi Hong, SeungBum Hong, Rian Huang, Ziyan Huang, Markus Huellebrand, Stephan Huschauer, Mustaffa Hussain, Tomoo Inubushi, Ece Isik Polat, Mojtaba Jafaritadi, SeongHun Jeong, Bailiang Jian, Yuanhong Jiang, Zhifan Jiang, Yueming Jin, Smriti Joshi, Abdolrahim Kadkhodamohammadi, Reda Abdellah Kamraoui, Inha Kang, Junghwa Kang, Davood Karimi, April Khademi, Muhammad Irfan Khan, Suleiman A. Khan, Rishab Khantwal, Kwang-Ju Kim, Timothy Kline, Satoshi Kondo, Elina Kontio, Adrian Krenzer, Artem Kroviakov, Hugo Kuijf, Satyadwyoom Kumar, Francesco La Rosa, Abhi Lad, Doohee Lee, Minho Lee, Chiara Lena, Hao Li, Ling Li, Xingyu Li, Fuyuan Liao, KuanLun Liao, Arlindo Limede Oliveira, Chaonan Lin, Shan Lin, Akis Linardos, Marius George Linguraru, Han Liu, Tao Liu, Di Liu, Yanling Liu, João Lourenço-Silva, Jingpei Lu, Jiangshan Lu, Imanol Luengo, Christina B. Lund, Huan Minh Luu, Yi Lv, Uzay Macar, Leon Maechler, Sina Mansour L., Kenji Marshall, Moona Mazher, Richard McKinley, Alfonso Medela, Felix Meissen, Mingyuan Meng, Dylan Miller, Seyed Hossein Mirjahanmardi, Arnab Mishra, Samir Mitha, Hassan Mohy-ud-Din, Tony Chi Wing Mok, Gowtham Krishnan Murugesan, Enamundram Naga Karthik, Sahil Nalawade, Jakub Nalepa, Mohamed Naser, Ramin Nateghi, Hammad Naveed, Quang-Minh Nguyen, Cuong Nguyen Quoc, Brennan Nichyporuk, Bruno Oliveira, David Owen, Jimut Bahan Pal, Junwen Pan, Wentao Pan, Winnie Pang, Bogyu Park, Vivek Pawar, Kamlesh Pawar, Michael Peven, Lena Philipp, Tomasz Pieciak, Szymon Plotka, Marcel Plutat, Fattaneh Pourakpour, Domen Preložnik, Kumaradevan Punithakumar, Abdul Qayyum, Sandro Queirós, Arman Rahmim, Salar Razavi, Jintao Ren, Mina Rezaei, Jonathan Adam Rico, ZunHyan Rieu, Markus Rink, Johannes Roth, Yusely Ruiz-Gonzalez, Numan Saeed, Anindo Saha, Mostafa Salem, Ricardo Sanchez-Matilla, Kurt Schilling, Wei Shao, Zhiqiang Shen, Ruize Shi, Pengcheng Shi, Daniel Sobotka, Théodore Soulier, Bella Specktor Fadida, Danail Stoyanov, Timothy Sum Hon Mun, Xiaowu Sun, Rong Tao, Franz Thaler, Antoine Théberge, Felix Thielke, Helena Torres, Kareem A. Wahid, Jiacheng Wang, YiFei Wang, Wei Wang, Xiong Wang, Jianhui Wen, Ning Wen, Marek Wodzinski, Ye Wu, Fangfang Xia, Tianqi Xiang, Chen Xiaofei, Lizhan Xu, Tingting Xue, Yuxuan Yang, Lin Yang, Kai Yao, Huifeng Yao, Amirsaeed Yazdani, Michael Yip, Hwanseung Yoo, Fereshteh Yousefirizi, Shunkai Yu, Lei Yu, Jonathan Zamora, Ramy Ashraf Zeineldin, Dewen Zeng, Jianpeng Zhang, Bokai Zhang, Jiapeng Zhang, Fan Zhang, Huahong Zhang, Zhongchen Zhao, Zixuan Zhao, Jiachen Zhao, Can Zhao, Qingshuo Zheng, Yuheng Zhi, Ziqi Zhou, Baosheng Zou, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge