Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Matthias Eisenmann, Annika Reinke, Vivienn Weru, Minu Dietlinde Tizabi, Fabian Isensee, Tim J. Adler, Patrick Godau, Veronika Cheplygina, Michal Kozubek, Sharib Ali, Anubha Gupta, Jan Kybic, Alison Noble, Carlos Ortiz de Solórzano, Samiksha Pachade, Caroline Petitjean, Daniel Sage, Donglai Wei, Elizabeth Wilden, Deepak Alapatt, Vincent Andrearczyk, Ujjwal Baid, Spyridon Bakas, Niranjan Balu, Sophia Bano, Vivek Singh Bawa, Jorge Bernal, Sebastian Bodenstedt, Alessandro Casella, Jinwook Choi, Olivier Commowick, Marie Daum, Adrien Depeursinge, Reuben Dorent, Jan Egger, Hannah Eichhorn, Sandy Engelhardt, Melanie Ganz, Gabriel Girard, Lasse Hansen, Mattias Heinrich, Nicholas Heller, Alessa Hering, Arnaud Huaulmé, Hyunjeong Kim, Bennett Landman, Hongwei Bran Li, Jianning Li, Jun Ma, Anne Martel, Carlos Martín-Isla, Bjoern Menze, Chinedu Innocent Nwoye, Valentin Oreiller, Nicolas Padoy, Sarthak Pati, Kelly Payette, Carole Sudre, Kimberlin van Wijnen, Armine Vardazaryan, Tom Vercauteren, Martin Wagner, Chuanbo Wang, Moi Hoon Yap, Zeyun Yu, Chun Yuan, Maximilian Zenk, Aneeq Zia, David Zimmerer, Rina Bao, Chanyeol Choi, Andrew Cohen, Oleh Dzyubachyk, Adrian Galdran, Tianyuan Gan, Tianqi Guo, Pradyumna Gupta, Mahmood Haithami, Edward Ho, Ikbeom Jang, Zhili Li, Zhengbo Luo, Filip Lux, Sokratis Makrogiannis, Dominik Müller, Young-tack Oh, Subeen Pang, Constantin Pape, Gorkem Polat, Charlotte Rosalie Reed, Kanghyun Ryu, Tim Scherr, Vajira Thambawita, Haoyu Wang, Xinliang Wang, Kele Xu, Hung Yeh, Doyeob Yeo, Yixuan Yuan, Yan Zeng, Xin Zhao, Julian Abbing, Jannes Adam, Nagesh Adluru, Niklas Agethen, Salman Ahmed, Yasmina Al Khalil, Mireia Alenyà, Esa Alhoniemi, Chengyang An, Talha Anwar, Tewodros Weldebirhan Arega, Netanell Avisdris, Dogu Baran Aydogan, Yingbin Bai, Maria Baldeon Calisto, Berke Doga Basaran, Marcel Beetz, Cheng Bian, Hao Bian, Kevin Blansit, Louise Bloch, Robert Bohnsack, Sara Bosticardo, Jack Breen, Mikael Brudfors, Raphael Brüngel, Mariano Cabezas, Alberto Cacciola, Zhiwei Chen, Yucong Chen, Daniel Tianming Chen, Minjeong Cho, Min-Kook Choi, Chuantao Xie Chuantao Xie, Dana Cobzas, Julien Cohen-Adad, Jorge Corral Acero, Sujit Kumar Das, Marcela de Oliveira, Hanqiu Deng, Guiming Dong, Lars Doorenbos, Cory Efird, Di Fan, Mehdi Fatan Serj, Alexandre Fenneteau, Lucas Fidon, Patryk Filipiak, René Finzel, Nuno R. Freitas, Christoph M. Friedrich, Mitchell Fulton, Finn Gaida, Francesco Galati, Christoforos Galazis, Chang Hee Gan, Zheyao Gao, Shengbo Gao, Matej Gazda, Beerend Gerats, Neil Getty, Adam Gibicar, Ryan Gifford, Sajan Gohil, Maria Grammatikopoulou, Daniel Grzech, Orhun Güley, Timo Günnemann, Chunxu Guo, Sylvain Guy, Heonjin Ha, Luyi Han, Il Song Han, Ali Hatamizadeh, Tian He, Jimin Heo, Sebastian Hitziger, SeulGi Hong, SeungBum Hong, Rian Huang, Ziyan Huang, Markus Huellebrand, Stephan Huschauer, Mustaffa Hussain, Tomoo Inubushi, Ece Isik Polat, Mojtaba Jafaritadi, SeongHun Jeong, Bailiang Jian, Yuanhong Jiang, Zhifan Jiang, Yueming Jin, Smriti Joshi, Abdolrahim Kadkhodamohammadi, Reda Abdellah Kamraoui, Inha Kang, Junghwa Kang, Davood Karimi, April Khademi, Muhammad Irfan Khan, Suleiman A. Khan, Rishab Khantwal, Kwang-Ju Kim, Timothy Kline, Satoshi Kondo, Elina Kontio, Adrian Krenzer, Artem Kroviakov, Hugo Kuijf, Satyadwyoom Kumar, Francesco La Rosa, Abhi Lad, Doohee Lee, Minho Lee, Chiara Lena, Hao Li, Ling Li, Xingyu Li, Fuyuan Liao, KuanLun Liao, Arlindo Limede Oliveira, Chaonan Lin, Shan Lin, Akis Linardos, Marius George Linguraru, Han Liu, Tao Liu, Di Liu, Yanling Liu, João Lourenço-Silva, Jingpei Lu, Jiangshan Lu, Imanol Luengo, Christina B. Lund, Huan Minh Luu, Yi Lv, Uzay Macar, Leon Maechler, Sina Mansour L., Kenji Marshall, Moona Mazher, Richard McKinley, Alfonso Medela, Felix Meissen, Mingyuan Meng, Dylan Miller, Seyed Hossein Mirjahanmardi, Arnab Mishra, Samir Mitha, Hassan Mohy-ud-Din, Tony Chi Wing Mok, Gowtham Krishnan Murugesan, Enamundram Naga Karthik, Sahil Nalawade, Jakub Nalepa, Mohamed Naser, Ramin Nateghi, Hammad Naveed, Quang-Minh Nguyen, Cuong Nguyen Quoc, Brennan Nichyporuk, Bruno Oliveira, David Owen, Jimut Bahan Pal, Junwen Pan, Wentao Pan, Winnie Pang, Bogyu Park, Vivek Pawar, Kamlesh Pawar, Michael Peven, Lena Philipp, Tomasz Pieciak, Szymon Plotka, Marcel Plutat, Fattaneh Pourakpour, Domen Preložnik, Kumaradevan Punithakumar, Abdul Qayyum, Sandro Queirós, Arman Rahmim, Salar Razavi, Jintao Ren, Mina Rezaei, Jonathan Adam Rico, ZunHyan Rieu, Markus Rink, Johannes Roth, Yusely Ruiz-Gonzalez, Numan Saeed, Anindo Saha, Mostafa Salem, Ricardo Sanchez-Matilla, Kurt Schilling, Wei Shao, Zhiqiang Shen, Ruize Shi, Pengcheng Shi, Daniel Sobotka, Théodore Soulier, Bella Specktor Fadida, Danail Stoyanov, Timothy Sum Hon Mun, Xiaowu Sun, Rong Tao, Franz Thaler, Antoine Théberge, Felix Thielke, Helena Torres, Kareem A. Wahid, Jiacheng Wang, YiFei Wang, Wei Wang, Xiong Wang, Jianhui Wen, Ning Wen, Marek Wodzinski, Ye Wu, Fangfang Xia, Tianqi Xiang, Chen Xiaofei, Lizhan Xu, Tingting Xue, Yuxuan Yang, Lin Yang, Kai Yao, Huifeng Yao, Amirsaeed Yazdani, Michael Yip, Hwanseung Yoo, Fereshteh Yousefirizi, Shunkai Yu, Lei Yu, Jonathan Zamora, Ramy Ashraf Zeineldin, Dewen Zeng, Jianpeng Zhang, Bokai Zhang, Jiapeng Zhang, Fan Zhang, Huahong Zhang, Zhongchen Zhao, Zixuan Zhao, Jiachen Zhao, Can Zhao, Qingshuo Zheng, Yuheng Zhi, Ziqi Zhou, Baosheng Zou, Klaus Maier-Hein, Paul F. Jäger, Annette Kopp-Schneider, Lena Maier-Hein
The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Deep Learning Methods for Lung Cancer Segmentation in Whole-slide Histopathology Images -- the ACDC@LungHP Challenge 2019
Aug 21, 2020Zhang Li, Jiehua Zhang, Tao Tan, Xichao Teng, Xiaoliang Sun, Yang Li, Lihong Liu, Yang Xiao, Byungjae Lee, Yilong Li, Qianni Zhang, Shujiao Sun, Yushan Zheng, Junyu Yan, Ni Li, Yiyu Hong, Junsu Ko, Hyun Jung, Yanling Liu, Yu-cheng Chen, Ching-wei Wang, Vladimir Yurovskiy, Pavel Maevskikh, Vahid Khanagha, Yi Jiang, Xiangjun Feng, Zhihong Liu, Daiqiang Li, Peter J. Schüffler, Qifeng Yu, Hui Chen, Yuling Tang, Geert Litjens
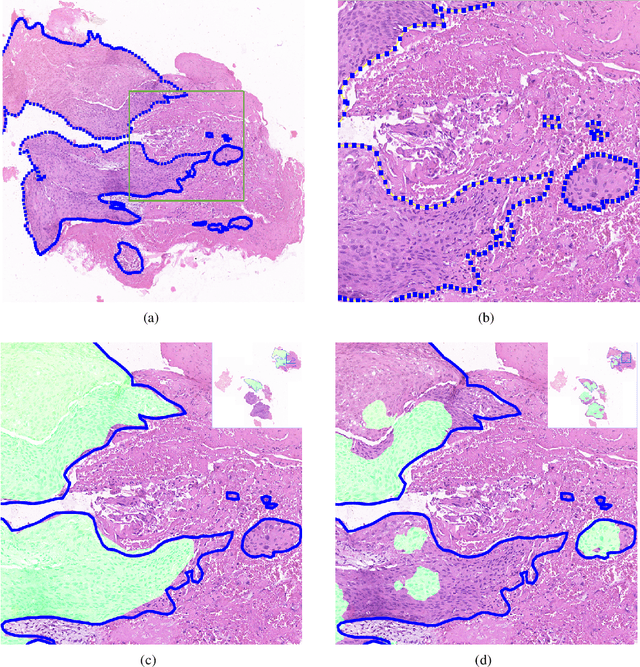
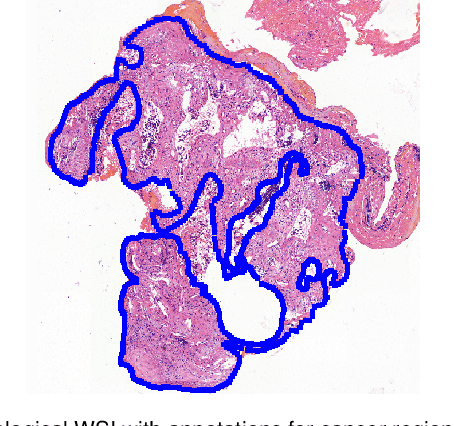
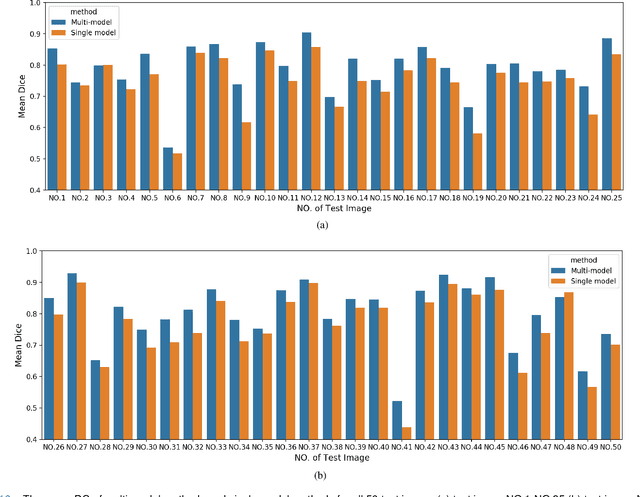
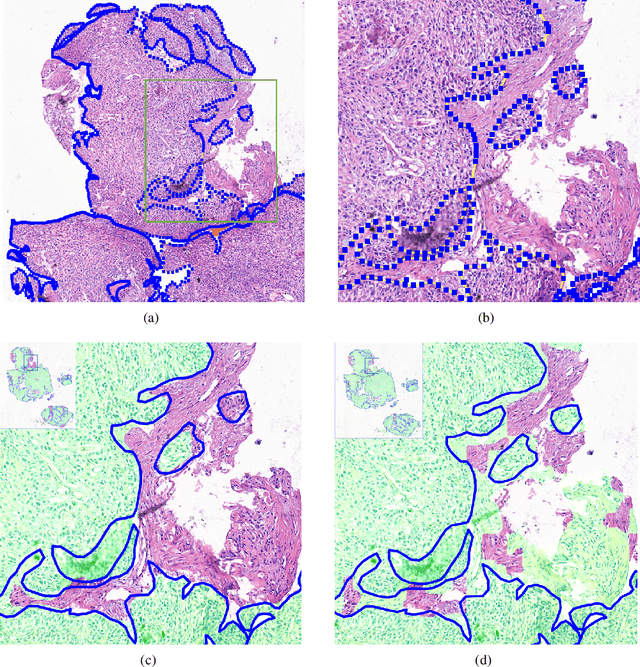

Accurate segmentation of lung cancer in pathology slides is a critical step in improving patient care. We proposed the ACDC@LungHP (Automatic Cancer Detection and Classification in Whole-slide Lung Histopathology) challenge for evaluating different computer-aided diagnosis (CADs) methods on the automatic diagnosis of lung cancer. The ACDC@LungHP 2019 focused on segmentation (pixel-wise detection) of cancer tissue in whole slide imaging (WSI), using an annotated dataset of 150 training images and 50 test images from 200 patients. This paper reviews this challenge and summarizes the top 10 submitted methods for lung cancer segmentation. All methods were evaluated using the false positive rate, false negative rate, and DICE coefficient (DC). The DC ranged from 0.7354$\pm$0.1149 to 0.8372$\pm$0.0858. The DC of the best method was close to the inter-observer agreement (0.8398$\pm$0.0890). All methods were based on deep learning and categorized into two groups: multi-model method and single model method. In general, multi-model methods were significantly better ($\textit{p}$<$0.01$) than single model methods, with mean DC of 0.7966 and 0.7544, respectively. Deep learning based methods could potentially help pathologists find suspicious regions for further analysis of lung cancer in WSI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge