How Important Is Tokenization in French Medical Masked Language Models?
Feb 22, 2024Yanis Labrak, Adrien Bazoge, Beatrice Daille, Mickael Rouvier, Richard Dufour
Subword tokenization has become the prevailing standard in the field of natural language processing (NLP) over recent years, primarily due to the widespread utilization of pre-trained language models. This shift began with Byte-Pair Encoding (BPE) and was later followed by the adoption of SentencePiece and WordPiece. While subword tokenization consistently outperforms character and word-level tokenization, the precise factors contributing to its success remain unclear. Key aspects such as the optimal segmentation granularity for diverse tasks and languages, the influence of data sources on tokenizers, and the role of morphological information in Indo-European languages remain insufficiently explored. This is particularly pertinent for biomedical terminology, characterized by specific rules governing morpheme combinations. Despite the agglutinative nature of biomedical terminology, existing language models do not explicitly incorporate this knowledge, leading to inconsistent tokenization strategies for common terms. In this paper, we seek to delve into the complexities of subword tokenization in French biomedical domain across a variety of NLP tasks and pinpoint areas where further enhancements can be made. We analyze classical tokenization algorithms, including BPE and SentencePiece, and introduce an original tokenization strategy that integrates morpheme-enriched word segmentation into existing tokenization methods.
DrBenchmark: A Large Language Understanding Evaluation Benchmark for French Biomedical Domain
Feb 20, 2024Yanis Labrak, Adrien Bazoge, Oumaima El Khettari, Mickael Rouvier, Pacome Constant dit Beaufils, Natalia Grabar, Beatrice Daille, Solen Quiniou, Emmanuel Morin, Pierre-Antoine Gourraud, Richard Dufour
The biomedical domain has sparked a significant interest in the field of Natural Language Processing (NLP), which has seen substantial advancements with pre-trained language models (PLMs). However, comparing these models has proven challenging due to variations in evaluation protocols across different models. A fair solution is to aggregate diverse downstream tasks into a benchmark, allowing for the assessment of intrinsic PLMs qualities from various perspectives. Although still limited to few languages, this initiative has been undertaken in the biomedical field, notably English and Chinese. This limitation hampers the evaluation of the latest French biomedical models, as they are either assessed on a minimal number of tasks with non-standardized protocols or evaluated using general downstream tasks. To bridge this research gap and account for the unique sensitivities of French, we present the first-ever publicly available French biomedical language understanding benchmark called DrBenchmark. It encompasses 20 diversified tasks, including named-entity recognition, part-of-speech tagging, question-answering, semantic textual similarity, and classification. We evaluate 8 state-of-the-art pre-trained masked language models (MLMs) on general and biomedical-specific data, as well as English specific MLMs to assess their cross-lingual capabilities. Our experiments reveal that no single model excels across all tasks, while generalist models are sometimes still competitive.
BioMistral: A Collection of Open-Source Pretrained Large Language Models for Medical Domains
Feb 15, 2024Yanis Labrak, Adrien Bazoge, Emmanuel Morin, Pierre-Antoine Gourraud, Mickael Rouvier, Richard Dufour
Large Language Models (LLMs) have demonstrated remarkable versatility in recent years, offering potential applications across specialized domains such as healthcare and medicine. Despite the availability of various open-source LLMs tailored for health contexts, adapting general-purpose LLMs to the medical domain presents significant challenges. In this paper, we introduce BioMistral, an open-source LLM tailored for the biomedical domain, utilizing Mistral as its foundation model and further pre-trained on PubMed Central. We conduct a comprehensive evaluation of BioMistral on a benchmark comprising 10 established medical question-answering (QA) tasks in English. We also explore lightweight models obtained through quantization and model merging approaches. Our results demonstrate BioMistral's superior performance compared to existing open-source medical models and its competitive edge against proprietary counterparts. Finally, to address the limited availability of data beyond English and to assess the multilingual generalization of medical LLMs, we automatically translated and evaluated this benchmark into 7 other languages. This marks the first large-scale multilingual evaluation of LLMs in the medical domain. Datasets, multilingual evaluation benchmarks, scripts, and all the models obtained during our experiments are freely released.
A Zero-shot and Few-shot Study of Instruction-Finetuned Large Language Models Applied to Clinical and Biomedical Tasks
Jul 22, 2023Yanis Labrak, Mickael Rouvier, Richard Dufour
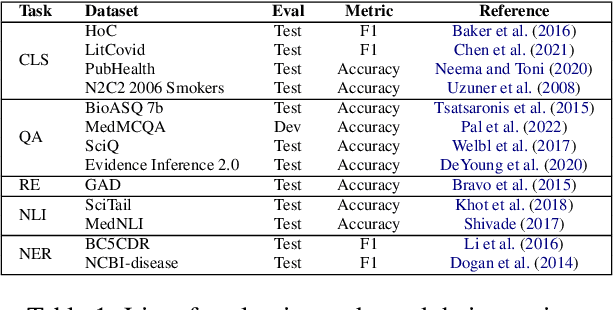
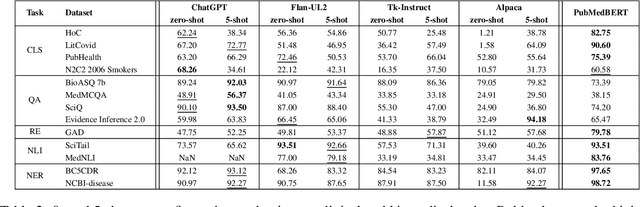
We evaluate four state-of-the-art instruction-tuned large language models (LLMs) -- ChatGPT, Flan-T5 UL2, Tk-Instruct, and Alpaca -- on a set of 13 real-world clinical and biomedical natural language processing (NLP) tasks in English, such as named-entity recognition (NER), question-answering (QA), relation extraction (RE), etc. Our overall results demonstrate that the evaluated LLMs begin to approach performance of state-of-the-art models in zero- and few-shot scenarios for most tasks, and particularly well for the QA task, even though they have never seen examples from these tasks before. However, we observed that the classification and RE tasks perform below what can be achieved with a specifically trained model for the medical field, such as PubMedBERT. Finally, we noted that no LLM outperforms all the others on all the studied tasks, with some models being better suited for certain tasks than others.
FrenchMedMCQA: A French Multiple-Choice Question Answering Dataset for Medical domain
Apr 09, 2023Yanis Labrak, Adrien Bazoge, Richard Dufour, Mickael Rouvier, Emmanuel Morin, Béatrice Daille, Pierre-Antoine Gourraud


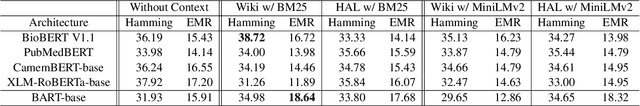
This paper introduces FrenchMedMCQA, the first publicly available Multiple-Choice Question Answering (MCQA) dataset in French for medical domain. It is composed of 3,105 questions taken from real exams of the French medical specialization diploma in pharmacy, mixing single and multiple answers. Each instance of the dataset contains an identifier, a question, five possible answers and their manual correction(s). We also propose first baseline models to automatically process this MCQA task in order to report on the current performances and to highlight the difficulty of the task. A detailed analysis of the results showed that it is necessary to have representations adapted to the medical domain or to the MCQA task: in our case, English specialized models yielded better results than generic French ones, even though FrenchMedMCQA is in French. Corpus, models and tools are available online.
DrBERT: A Robust Pre-trained Model in French for Biomedical and Clinical domains
Apr 03, 2023Yanis Labrak, Adrien Bazoge, Richard Dufour, Mickael Rouvier, Emmanuel Morin, Béatrice Daille, Pierre-Antoine Gourraud




In recent years, pre-trained language models (PLMs) achieve the best performance on a wide range of natural language processing (NLP) tasks. While the first models were trained on general domain data, specialized ones have emerged to more effectively treat specific domains. In this paper, we propose an original study of PLMs in the medical domain on French language. We compare, for the first time, the performance of PLMs trained on both public data from the web and private data from healthcare establishments. We also evaluate different learning strategies on a set of biomedical tasks. In particular, we show that we can take advantage of already existing biomedical PLMs in a foreign language by further pre-train it on our targeted data. Finally, we release the first specialized PLMs for the biomedical field in French, called DrBERT, as well as the largest corpus of medical data under free license on which these models are trained.
BLOOM: A 176B-Parameter Open-Access Multilingual Language Model
Nov 09, 2022Teven Le Scao, Angela Fan, Christopher Akiki, Ellie Pavlick, Suzana Ilić, Daniel Hesslow, Roman Castagné, Alexandra Sasha Luccioni, François Yvon, Matthias Gallé, Jonathan Tow, Alexander M. Rush, Stella Biderman, Albert Webson, Pawan Sasanka Ammanamanchi, Thomas Wang, Benoît Sagot, Niklas Muennighoff, Albert Villanova del Moral, Olatunji Ruwase, Rachel Bawden, Stas Bekman, Angelina McMillan-Major, Iz Beltagy, Huu Nguyen, Lucile Saulnier, Samson Tan, Pedro Ortiz Suarez, Victor Sanh, Hugo Laurençon, Yacine Jernite, Julien Launay, Margaret Mitchell, Colin Raffel, Aaron Gokaslan, Adi Simhi, Aitor Soroa, Alham Fikri Aji, Amit Alfassy, Anna Rogers, Ariel Kreisberg Nitzav, Canwen Xu, Chenghao Mou, Chris Emezue, Christopher Klamm, Colin Leong, Daniel van Strien, David Ifeoluwa Adelani, Dragomir Radev, Eduardo González Ponferrada, Efrat Levkovizh, Ethan Kim, Eyal Bar Natan, Francesco De Toni, Gérard Dupont, Germán Kruszewski, Giada Pistilli, Hady Elsahar, Hamza Benyamina, Hieu Tran, Ian Yu, Idris Abdulmumin, Isaac Johnson, Itziar Gonzalez-Dios, Javier de la Rosa, Jenny Chim, Jesse Dodge, Jian Zhu, Jonathan Chang, Jörg Frohberg, Joseph Tobing, Joydeep Bhattacharjee, Khalid Almubarak, Kimbo Chen, Kyle Lo, Leandro Von Werra, Leon Weber, Long Phan, Loubna Ben allal, Ludovic Tanguy, Manan Dey, Manuel Romero Muñoz, Maraim Masoud, María Grandury, Mario Šaško, Max Huang, Maximin Coavoux, Mayank Singh, Mike Tian-Jian Jiang, Minh Chien Vu, Mohammad A. Jauhar, Mustafa Ghaleb, Nishant Subramani, Nora Kassner, Nurulaqilla Khamis, Olivier Nguyen, Omar Espejel, Ona de Gibert, Paulo Villegas, Peter Henderson, Pierre Colombo, Priscilla Amuok, Quentin Lhoest, Rheza Harliman, Rishi Bommasani, Roberto Luis López, Rui Ribeiro, Salomey Osei, Sampo Pyysalo, Sebastian Nagel, Shamik Bose, Shamsuddeen Hassan Muhammad, Shanya Sharma, Shayne Longpre, Somaieh Nikpoor, Stanislav Silberberg, Suhas Pai, Sydney Zink, Tiago Timponi Torrent, Timo Schick, Tristan Thrush, Valentin Danchev, Vassilina Nikoulina, Veronika Laippala, Violette Lepercq, Vrinda Prabhu, Zaid Alyafeai, Zeerak Talat, Arun Raja, Benjamin Heinzerling, Chenglei Si, Elizabeth Salesky, Sabrina J. Mielke, Wilson Y. Lee, Abheesht Sharma, Andrea Santilli, Antoine Chaffin, Arnaud Stiegler, Debajyoti Datta, Eliza Szczechla, Gunjan Chhablani, Han Wang, Harshit Pandey, Hendrik Strobelt, Jason Alan Fries, Jos Rozen, Leo Gao, Lintang Sutawika, M Saiful Bari, Maged S. Al-shaibani, Matteo Manica, Nihal Nayak, Ryan Teehan, Samuel Albanie, Sheng Shen, Srulik Ben-David, Stephen H. Bach, Taewoon Kim, Tali Bers, Thibault Fevry, Trishala Neeraj, Urmish Thakker, Vikas Raunak, Xiangru Tang, Zheng-Xin Yong, Zhiqing Sun, Shaked Brody, Yallow Uri, Hadar Tojarieh, Adam Roberts, Hyung Won Chung, Jaesung Tae, Jason Phang, Ofir Press, Conglong Li, Deepak Narayanan, Hatim Bourfoune, Jared Casper, Jeff Rasley, Max Ryabinin, Mayank Mishra, Minjia Zhang, Mohammad Shoeybi, Myriam Peyrounette, Nicolas Patry, Nouamane Tazi, Omar Sanseviero, Patrick von Platen, Pierre Cornette, Pierre François Lavallée, Rémi Lacroix, Samyam Rajbhandari, Sanchit Gandhi, Shaden Smith, Stéphane Requena, Suraj Patil, Tim Dettmers, Ahmed Baruwa, Amanpreet Singh, Anastasia Cheveleva, Anne-Laure Ligozat, Arjun Subramonian, Aurélie Névéol, Charles Lovering, Dan Garrette, Deepak Tunuguntla, Ehud Reiter, Ekaterina Taktasheva, Ekaterina Voloshina, Eli Bogdanov, Genta Indra Winata, Hailey Schoelkopf, Jan-Christoph Kalo, Jekaterina Novikova, Jessica Zosa Forde, Jordan Clive, Jungo Kasai, Ken Kawamura, Liam Hazan, Marine Carpuat, Miruna Clinciu, Najoung Kim, Newton Cheng, Oleg Serikov, Omer Antverg, Oskar van der Wal, Rui Zhang, Ruochen Zhang, Sebastian Gehrmann, Shani Pais, Tatiana Shavrina, Thomas Scialom, Tian Yun, Tomasz Limisiewicz, Verena Rieser, Vitaly Protasov, Vladislav Mikhailov, Yada Pruksachatkun, Yonatan Belinkov, Zachary Bamberger, Zdeněk Kasner, Alice Rueda, Amanda Pestana, Amir Feizpour, Ammar Khan, Amy Faranak, Ana Santos, Anthony Hevia, Antigona Unldreaj, Arash Aghagol, Arezoo Abdollahi, Aycha Tammour, Azadeh HajiHosseini, Bahareh Behroozi, Benjamin Ajibade, Bharat Saxena, Carlos Muñoz Ferrandis, Danish Contractor, David Lansky, Davis David, Douwe Kiela, Duong A. Nguyen, Edward Tan, Emi Baylor, Ezinwanne Ozoani, Fatima Mirza, Frankline Ononiwu, Habib Rezanejad, Hessie Jones, Indrani Bhattacharya, Irene Solaiman, Irina Sedenko, Isar Nejadgholi, Jesse Passmore, Josh Seltzer, Julio Bonis Sanz, Karen Fort, Livia Dutra, Mairon Samagaio, Maraim Elbadri, Margot Mieskes, Marissa Gerchick, Martha Akinlolu, Michael McKenna, Mike Qiu, Muhammed Ghauri, Mykola Burynok, Nafis Abrar, Nazneen Rajani, Nour Elkott, Nour Fahmy, Olanrewaju Samuel, Ran An, Rasmus Kromann, Ryan Hao, Samira Alizadeh, Sarmad Shubber, Silas Wang, Sourav Roy, Sylvain Viguier, Thanh Le, Tobi Oyebade, Trieu Le, Yoyo Yang, Zach Nguyen, Abhinav Ramesh Kashyap, Alfredo Palasciano, Alison Callahan, Anima Shukla, Antonio Miranda-Escalada, Ayush Singh, Benjamin Beilharz, Bo Wang, Caio Brito, Chenxi Zhou, Chirag Jain, Chuxin Xu, Clémentine Fourrier, Daniel León Periñán, Daniel Molano, Dian Yu, Enrique Manjavacas, Fabio Barth, Florian Fuhrimann, Gabriel Altay, Giyaseddin Bayrak, Gully Burns, Helena U. Vrabec, Imane Bello, Ishani Dash, Jihyun Kang, John Giorgi, Jonas Golde, Jose David Posada, Karthik Rangasai Sivaraman, Lokesh Bulchandani, Lu Liu, Luisa Shinzato, Madeleine Hahn de Bykhovetz, Maiko Takeuchi, Marc Pàmies, Maria A Castillo, Marianna Nezhurina, Mario Sänger, Matthias Samwald, Michael Cullan, Michael Weinberg, Michiel De Wolf, Mina Mihaljcic, Minna Liu, Moritz Freidank, Myungsun Kang, Natasha Seelam, Nathan Dahlberg, Nicholas Michio Broad, Nikolaus Muellner, Pascale Fung, Patrick Haller, Ramya Chandrasekhar, Renata Eisenberg, Robert Martin, Rodrigo Canalli, Rosaline Su, Ruisi Su, Samuel Cahyawijaya, Samuele Garda, Shlok S Deshmukh, Shubhanshu Mishra, Sid Kiblawi, Simon Ott, Sinee Sang-aroonsiri, Srishti Kumar, Stefan Schweter, Sushil Bharati, Tanmay Laud, Théo Gigant, Tomoya Kainuma, Wojciech Kusa, Yanis Labrak, Yash Shailesh Bajaj, Yash Venkatraman, Yifan Xu, Yingxin Xu, Yu Xu, Zhe Tan, Zhongli Xie, Zifan Ye, Mathilde Bras, Younes Belkada, Thomas Wolf
Large language models (LLMs) have been shown to be able to perform new tasks based on a few demonstrations or natural language instructions. While these capabilities have led to widespread adoption, most LLMs are developed by resource-rich organizations and are frequently kept from the public. As a step towards democratizing this powerful technology, we present BLOOM, a 176B-parameter open-access language model designed and built thanks to a collaboration of hundreds of researchers. BLOOM is a decoder-only Transformer language model that was trained on the ROOTS corpus, a dataset comprising hundreds of sources in 46 natural and 13 programming languages (59 in total). We find that BLOOM achieves competitive performance on a wide variety of benchmarks, with stronger results after undergoing multitask prompted finetuning. To facilitate future research and applications using LLMs, we publicly release our models and code under the Responsible AI License.
BigBIO: A Framework for Data-Centric Biomedical Natural Language Processing
Jun 30, 2022Jason Alan Fries, Leon Weber, Natasha Seelam, Gabriel Altay, Debajyoti Datta, Samuele Garda, Myungsun Kang, Ruisi Su, Wojciech Kusa, Samuel Cahyawijaya, Fabio Barth, Simon Ott, Matthias Samwald, Stephen Bach, Stella Biderman, Mario Sänger, Bo Wang, Alison Callahan, Daniel León Periñán, Théo Gigant, Patrick Haller, Jenny Chim, Jose David Posada, John Michael Giorgi, Karthik Rangasai Sivaraman, Marc Pàmies, Marianna Nezhurina, Robert Martin, Michael Cullan, Moritz Freidank, Nathan Dahlberg, Shubhanshu Mishra, Shamik Bose, Nicholas Michio Broad, Yanis Labrak, Shlok S Deshmukh, Sid Kiblawi, Ayush Singh, Minh Chien Vu, Trishala Neeraj, Jonas Golde, Albert Villanova del Moral, Benjamin Beilharz
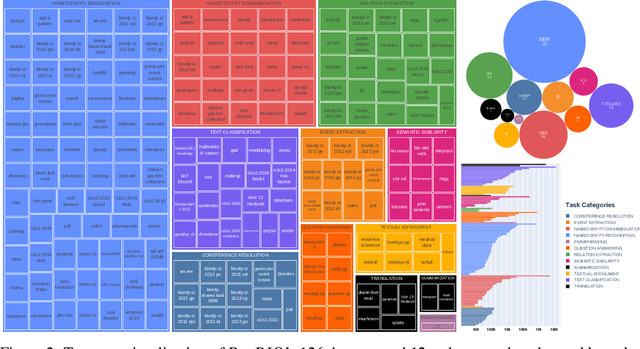
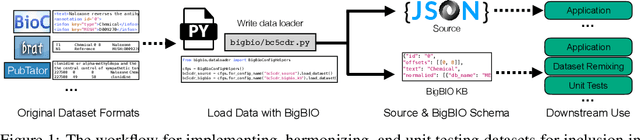
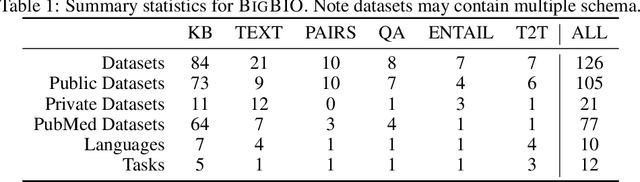

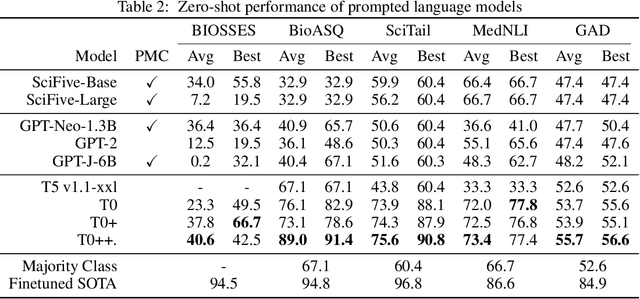
Training and evaluating language models increasingly requires the construction of meta-datasets --diverse collections of curated data with clear provenance. Natural language prompting has recently lead to improved zero-shot generalization by transforming existing, supervised datasets into a diversity of novel pretraining tasks, highlighting the benefits of meta-dataset curation. While successful in general-domain text, translating these data-centric approaches to biomedical language modeling remains challenging, as labeled biomedical datasets are significantly underrepresented in popular data hubs. To address this challenge, we introduce BigBIO a community library of 126+ biomedical NLP datasets, currently covering 12 task categories and 10+ languages. BigBIO facilitates reproducible meta-dataset curation via programmatic access to datasets and their metadata, and is compatible with current platforms for prompt engineering and end-to-end few/zero shot language model evaluation. We discuss our process for task schema harmonization, data auditing, contribution guidelines, and outline two illustrative use cases: zero-shot evaluation of biomedical prompts and large-scale, multi-task learning. BigBIO is an ongoing community effort and is available at https://github.com/bigscience-workshop/biomedical
Multi-label classification for biomedical literature: an overview of the BioCreative VII LitCovid Track for COVID-19 literature topic annotations
Apr 20, 2022Qingyu Chen, Alexis Allot, Robert Leaman, Rezarta Islamaj Doğan, Jingcheng Du, Li Fang, Wang Kai, Shuo Xu, Yuefu Zhang, Parsa Bagherzadeh, Sabine Bergler, Aakash Bhatnagar, Nidhir Bhavsar, Yung-Chun Chang, Sheng-Jie Lin, Wentai Tang, Hongtong Zhang, Ilija Tavchioski, Shubo Tian, Jinfeng Zhang, Yulia Otmakhova, Antonio Jimeno Yepes, Hang Dong, Honghan Wu, Richard Dufour, Yanis Labrak, Niladri Chatterjee, Kushagri Tandon, Fréjus Laleye, Loïc Rakotoson, Emmanuele Chersoni, Jinghang Gu, Annemarie Friedrich, Subhash Chandra Pujari, Mariia Chizhikova, Naveen Sivadasan, Zhiyong Lu
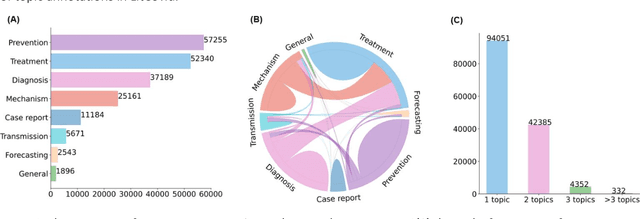
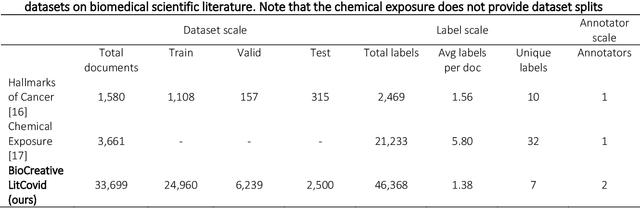
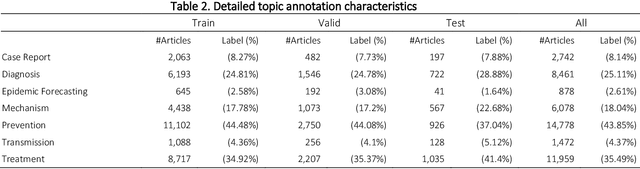
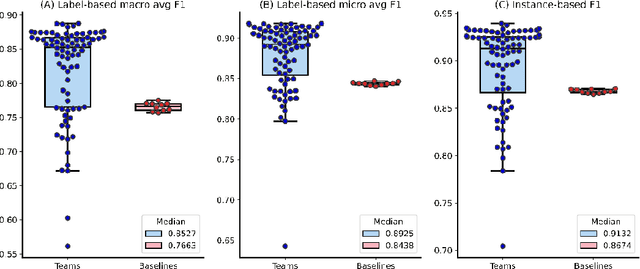
The COVID-19 pandemic has been severely impacting global society since December 2019. Massive research has been undertaken to understand the characteristics of the virus and design vaccines and drugs. The related findings have been reported in biomedical literature at a rate of about 10,000 articles on COVID-19 per month. Such rapid growth significantly challenges manual curation and interpretation. For instance, LitCovid is a literature database of COVID-19-related articles in PubMed, which has accumulated more than 200,000 articles with millions of accesses each month by users worldwide. One primary curation task is to assign up to eight topics (e.g., Diagnosis and Treatment) to the articles in LitCovid. Despite the continuing advances in biomedical text mining methods, few have been dedicated to topic annotations in COVID-19 literature. To close the gap, we organized the BioCreative LitCovid track to call for a community effort to tackle automated topic annotation for COVID-19 literature. The BioCreative LitCovid dataset, consisting of over 30,000 articles with manually reviewed topics, was created for training and testing. It is one of the largest multilabel classification datasets in biomedical scientific literature. 19 teams worldwide participated and made 80 submissions in total. Most teams used hybrid systems based on transformers. The highest performing submissions achieved 0.8875, 0.9181, and 0.9394 for macro F1-score, micro F1-score, and instance-based F1-score, respectively. The level of participation and results demonstrate a successful track and help close the gap between dataset curation and method development. The dataset is publicly available via https://ftp.ncbi.nlm.nih.gov/pub/lu/LitCovid/biocreative/ for benchmarking and further development.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge