Real-Time 4K Super-Resolution of Compressed AVIF Images. AIS 2024 Challenge Survey
Apr 25, 2024Marcos V. Conde, Zhijun Lei, Wen Li, Cosmin Stejerean, Ioannis Katsavounidis, Radu Timofte, Kihwan Yoon, Ganzorig Gankhuyag, Jiangtao Lv, Long Sun, Jinshan Pan, Jiangxin Dong, Jinhui Tang, Zhiyuan Li, Hao Wei, Chenyang Ge, Dongyang Zhang, Tianle Liu, Huaian Chen, Yi Jin, Menghan Zhou, Yiqiang Yan, Si Gao, Biao Wu, Shaoli Liu, Chengjian Zheng, Diankai Zhang, Ning Wang, Xintao Qiu, Yuanbo Zhou, Kongxian Wu, Xinwei Dai, Hui Tang, Wei Deng, Qingquan Gao, Tong Tong, Jae-Hyeon Lee, Ui-Jin Choi, Min Yan, Xin Liu, Qian Wang, Xiaoqian Ye, Zhan Du, Tiansen Zhang, Long Peng, Jiaming Guo, Xin Di, Bohao Liao, Zhibo Du, Peize Xia, Renjing Pei, Yang Wang, Yang Cao, Zhengjun Zha, Bingnan Han, Hongyuan Yu, Zhuoyuan Wu, Cheng Wan, Yuqing Liu, Haodong Yu, Jizhe Li, Zhijuan Huang, Yuan Huang, Yajun Zou, Xianyu Guan, Qi Jia, Heng Zhang, Xuanwu Yin, Kunlong Zuo, Hyeon-Cheol Moon, Tae-hyun Jeong, Yoonmo Yang, Jae-Gon Kim, Jinwoo Jeong, Sunjei Kim
This paper introduces a novel benchmark as part of the AIS 2024 Real-Time Image Super-Resolution (RTSR) Challenge, which aims to upscale compressed images from 540p to 4K resolution (4x factor) in real-time on commercial GPUs. For this, we use a diverse test set containing a variety of 4K images ranging from digital art to gaming and photography. The images are compressed using the modern AVIF codec, instead of JPEG. All the proposed methods improve PSNR fidelity over Lanczos interpolation, and process images under 10ms. Out of the 160 participants, 25 teams submitted their code and models. The solutions present novel designs tailored for memory-efficiency and runtime on edge devices. This survey describes the best solutions for real-time SR of compressed high-resolution images.
NTIRE 2024 Challenge on Low Light Image Enhancement: Methods and Results
Apr 22, 2024Xiaoning Liu, Zongwei Wu, Ao Li, Florin-Alexandru Vasluianu, Yulun Zhang, Shuhang Gu, Le Zhang, Ce Zhu, Radu Timofte, Zhi Jin, Hongjun Wu, Chenxi Wang, Haitao Ling, Yuanhao Cai, Hao Bian, Yuxin Zheng, Jing Lin, Alan Yuille, Ben Shao, Jin Guo, Tianli Liu, Mohao Wu, Yixu Feng, Shuo Hou, Haotian Lin, Yu Zhu, Peng Wu, Wei Dong, Jinqiu Sun, Yanning Zhang, Qingsen Yan, Wenbin Zou, Weipeng Yang, Yunxiang Li, Qiaomu Wei, Tian Ye, Sixiang Chen, Zhao Zhang, Suiyi Zhao, Bo Wang, Yan Luo, Zhichao Zuo, Mingshen Wang, Junhu Wang, Yanyan Wei, Xiaopeng Sun, Yu Gao, Jiancheng Huang, Hongming Chen, Xiang Chen, Hui Tang, Yuanbin Chen, Yuanbo Zhou, Xinwei Dai, Xintao Qiu, Wei Deng, Qinquan Gao, Tong Tong, Mingjia Li, Jin Hu, Xinyu He, Xiaojie Guo, Sabarinathan, K Uma, A Sasithradevi, B Sathya Bama, S. Mohamed Mansoor Roomi, V. Srivatsav, Jinjuan Wang, Long Sun, Qiuying Chen, Jiahong Shao, Yizhi Zhang, Marcos V. Conde, Daniel Feijoo, Juan C. Benito, Alvaro García, Jaeho Lee, Seongwan Kim, Sharif S M A, Nodirkhuja Khujaev, Roman Tsoy, Ali Murtaza, Uswah Khairuddin, Ahmad 'Athif Mohd Faudzi, Sampada Malagi, Amogh Joshi, Nikhil Akalwadi, Chaitra Desai, Ramesh Ashok Tabib, Uma Mudenagudi, Wenyi Lian, Wenjing Lian, Jagadeesh Kalyanshetti, Vijayalaxmi Ashok Aralikatti, Palani Yashaswini, Nitish Upasi, Dikshit Hegde, Ujwala Patil, Sujata C, Xingzhuo Yan, Wei Hao, Minghan Fu, Pooja choksy, Anjali Sarvaiya, Kishor Upla, Kiran Raja, Hailong Yan, Yunkai Zhang, Baiang Li, Jingyi Zhang, Huan Zheng
This paper reviews the NTIRE 2024 low light image enhancement challenge, highlighting the proposed solutions and results. The aim of this challenge is to discover an effective network design or solution capable of generating brighter, clearer, and visually appealing results when dealing with a variety of conditions, including ultra-high resolution (4K and beyond), non-uniform illumination, backlighting, extreme darkness, and night scenes. A notable total of 428 participants registered for the challenge, with 22 teams ultimately making valid submissions. This paper meticulously evaluates the state-of-the-art advancements in enhancing low-light images, reflecting the significant progress and creativity in this field.
Innovative Quantitative Analysis for Disease Progression Assessment in Familial Cerebral Cavernous Malformations
Mar 23, 2024Ruige Zong, Tao Wang, Chunwang Li, Xinlin Zhang, Yuanbin Chen, Longxuan Zhao, Qixuan Li, Qinquan Gao, Dezhi Kang, Fuxin Lin, Tong Tong
Familial cerebral cavernous malformation (FCCM) is a hereditary disorder characterized by abnormal vascular structures within the central nervous system. The FCCM lesions are often numerous and intricate, making quantitative analysis of the lesions a labor-intensive task. Consequently, clinicians face challenges in quantitatively assessing the severity of lesions and determining whether lesions have progressed. To alleviate this problem, we propose a quantitative statistical framework for FCCM, comprising an efficient annotation module, an FCCM lesion segmentation module, and an FCCM lesion quantitative statistics module. Our framework demonstrates precise segmentation of the FCCM lesion based on efficient data annotation, achieving a Dice coefficient of 93.22\%. More importantly, we focus on quantitative statistics of lesions, which is combined with image registration to realize the quantitative comparison of lesions between different examinations of patients, and a visualization framework has been established for doctors to comprehensively compare and analyze lesions. The experimental results have demonstrated that our proposed framework not only obtains objective, accurate, and comprehensive quantitative statistical information, which provides a quantitative assessment method for disease progression and drug efficacy study, but also considerably reduces the manual measurement and statistical workload of lesions, assisting clinical decision-making for FCCM and accelerating progress in FCCM clinical research. This highlights the potential of practical application of the framework in FCCM clinical research and clinical decision-making. The codes are available at https://github.com/6zrg/Quantitative-Statistics-of-FCCM.
Distance Guided Generative Adversarial Network for Explainable Binary Classifications
Dec 29, 2023Xiangyu Xiong, Yue Sun, Xiaohong Liu, Wei Ke, Chan-Tong Lam, Jiangang Chen, Mingfeng Jiang, Mingwei Wang, Hui Xie, Tong Tong, Qinquan Gao, Hao Chen, Tao Tan
Despite the potential benefits of data augmentation for mitigating the data insufficiency, traditional augmentation methods primarily rely on the prior intra-domain knowledge. On the other hand, advanced generative adversarial networks (GANs) generate inter-domain samples with limited variety. These previous methods make limited contributions to describing the decision boundaries for binary classification. In this paper, we propose a distance guided GAN (DisGAN) which controls the variation degrees of generated samples in the hyperplane space. Specifically, we instantiate the idea of DisGAN by combining two ways. The first way is vertical distance GAN (VerDisGAN) where the inter-domain generation is conditioned on the vertical distances. The second way is horizontal distance GAN (HorDisGAN) where the intra-domain generation is conditioned on the horizontal distances. Furthermore, VerDisGAN can produce the class-specific regions by mapping the source images to the hyperplane. Experimental results show that DisGAN consistently outperforms the GAN-based augmentation methods with explainable binary classification. The proposed method can apply to different classification architectures and has potential to extend to multi-class classification.
Toward Real World Stereo Image Super-Resolution via Hybrid Degradation Model and Discriminator for Implied Stereo Image Information
Dec 13, 2023Yuanbo Zhou, Yuyang Xue, Jiang Bi, Wenlin He, Xinlin Zhang, Jiajun Zhang, Wei Deng, Ruofeng Nie, Junlin Lan, Qinquan Gao, Tong Tong
Real-world stereo image super-resolution has a significant influence on enhancing the performance of computer vision systems. Although existing methods for single-image super-resolution can be applied to improve stereo images, these methods often introduce notable modifications to the inherent disparity, resulting in a loss in the consistency of disparity between the original and the enhanced stereo images. To overcome this limitation, this paper proposes a novel approach that integrates a implicit stereo information discriminator and a hybrid degradation model. This combination ensures effective enhancement while preserving disparity consistency. The proposed method bridges the gap between the complex degradations in real-world stereo domain and the simpler degradations in real-world single-image super-resolution domain. Our results demonstrate impressive performance on synthetic and real datasets, enhancing visual perception while maintaining disparity consistency. The complete code is available at the following \href{https://github.com/fzuzyb/SCGLANet}{link}.
A Parameterized Generative Adversarial Network Using Cyclic Projection for Explainable Medical Image Classification
Dec 07, 2023Xiangyu Xiong, Yue Sun, Xiaohong Liu, Chan-Tong Lam, Tong Tong, Hao Chen, Qinquan Gao, Wei Ke, Tao Tan
Although current data augmentation methods are successful to alleviate the data insufficiency, conventional augmentation are primarily intra-domain while advanced generative adversarial networks (GANs) generate images remaining uncertain, particularly in small-scale datasets. In this paper, we propose a parameterized GAN (ParaGAN) that effectively controls the changes of synthetic samples among domains and highlights the attention regions for downstream classification. Specifically, ParaGAN incorporates projection distance parameters in cyclic projection and projects the source images to the decision boundary to obtain the class-difference maps. Our experiments show that ParaGAN can consistently outperform the existing augmentation methods with explainable classification on two small-scale medical datasets.
Pseudo Label-Guided Data Fusion and Output Consistency for Semi-Supervised Medical Image Segmentation
Nov 17, 2023Tao Wang, Yuanbin Chen, Xinlin Zhang, Yuanbo Zhou, Junlin Lan, Bizhe Bai, Tao Tan, Min Du, Qinquan Gao, Tong Tong
Supervised learning algorithms based on Convolutional Neural Networks have become the benchmark for medical image segmentation tasks, but their effectiveness heavily relies on a large amount of labeled data. However, annotating medical image datasets is a laborious and time-consuming process. Inspired by semi-supervised algorithms that use both labeled and unlabeled data for training, we propose the PLGDF framework, which builds upon the mean teacher network for segmenting medical images with less annotation. We propose a novel pseudo-label utilization scheme, which combines labeled and unlabeled data to augment the dataset effectively. Additionally, we enforce the consistency between different scales in the decoder module of the segmentation network and propose a loss function suitable for evaluating the consistency. Moreover, we incorporate a sharpening operation on the predicted results, further enhancing the accuracy of the segmentation. Extensive experiments on three publicly available datasets demonstrate that the PLGDF framework can largely improve performance by incorporating the unlabeled data. Meanwhile, our framework yields superior performance compared to six state-of-the-art semi-supervised learning methods. The codes of this study are available at https://github.com/ortonwang/PLGDF.
Dual-Decoder Consistency via Pseudo-Labels Guided Data Augmentation for Semi-Supervised Medical Image Segmentation
Aug 31, 2023Yuanbin Chen, Tao Wang, Hui Tang, Longxuan Zhao, Ruige Zong, Tao Tan, Xinlin Zhang, Tong Tong
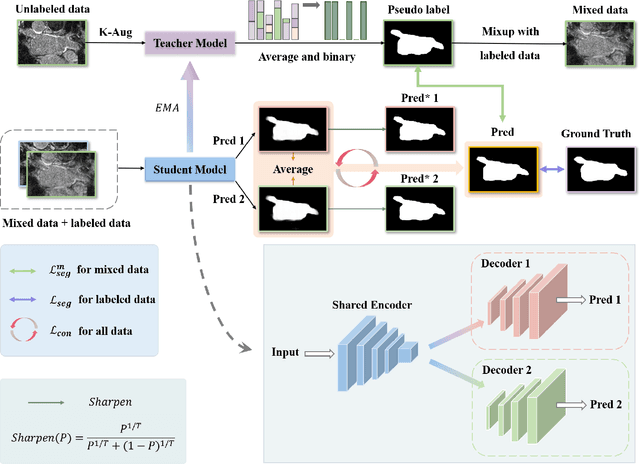
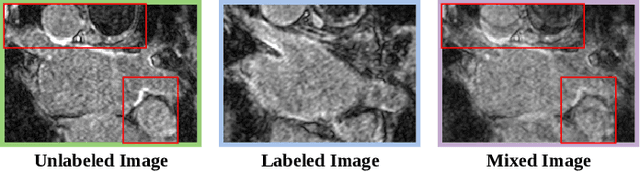

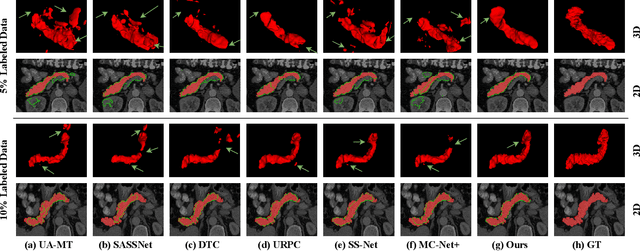
Medical image segmentation methods often rely on fully supervised approaches to achieve excellent performance, which is contingent upon having an extensive set of labeled images for training. However, annotating medical images is both expensive and time-consuming. Semi-supervised learning offers a solution by leveraging numerous unlabeled images alongside a limited set of annotated ones. In this paper, we introduce a semi-supervised medical image segmentation method based on the mean-teacher model, referred to as Dual-Decoder Consistency via Pseudo-Labels Guided Data Augmentation (DCPA). This method combines consistency regularization, pseudo-labels, and data augmentation to enhance the efficacy of semi-supervised segmentation. Firstly, the proposed model comprises both student and teacher models with a shared encoder and two distinct decoders employing different up-sampling strategies. Minimizing the output discrepancy between decoders enforces the generation of consistent representations, serving as regularization during student model training. Secondly, we introduce mixup operations to blend unlabeled data with labeled data, creating mixed data and thereby achieving data augmentation. Lastly, pseudo-labels are generated by the teacher model and utilized as labels for mixed data to compute unsupervised loss. We compare the segmentation results of the DCPA model with six state-of-the-art semi-supervised methods on three publicly available medical datasets. Beyond classical 10\% and 20\% semi-supervised settings, we investigate performance with less supervision (5\% labeled data). Experimental outcomes demonstrate that our approach consistently outperforms existing semi-supervised medical image segmentation methods across the three semi-supervised settings.
PCDAL: A Perturbation Consistency-Driven Active Learning Approach for Medical Image Segmentation and Classification
Jun 29, 2023Tao Wang, Xinlin Zhang, Yuanbo Zhou, Junlin Lan, Tao Tan, Min Du, Qinquan Gao, Tong Tong
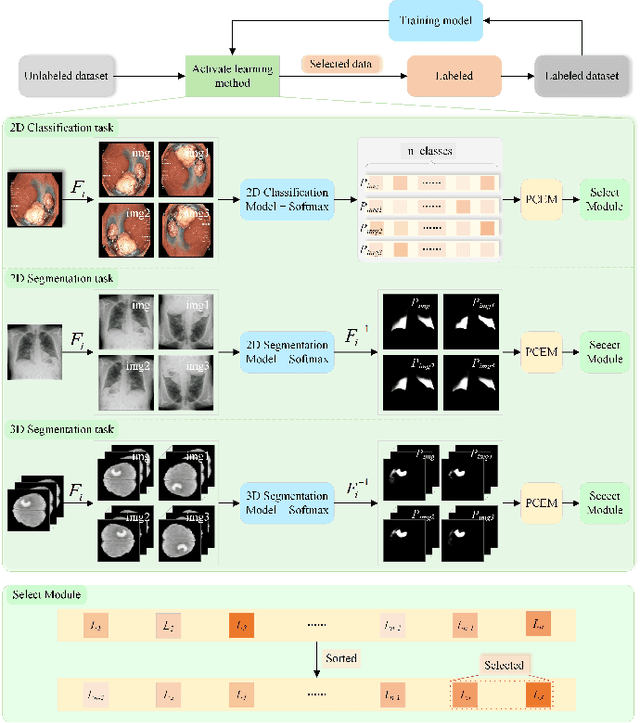
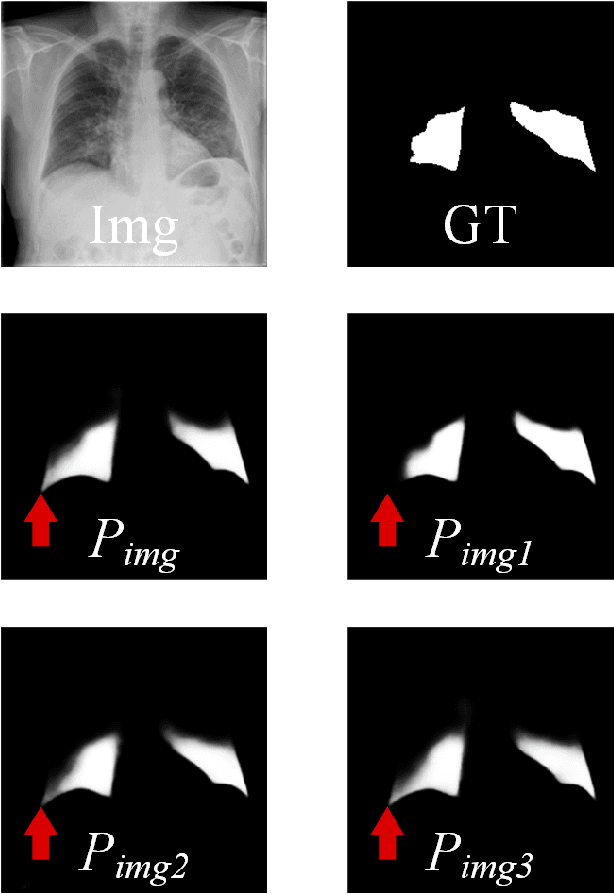
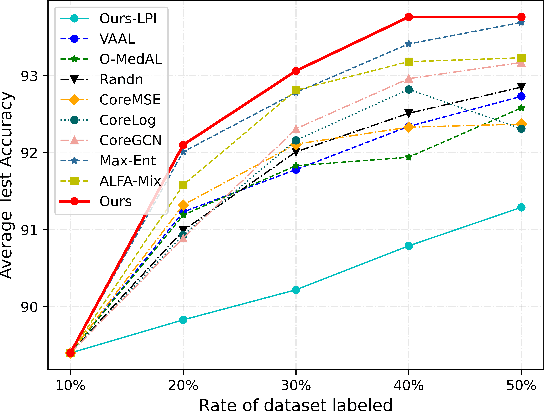
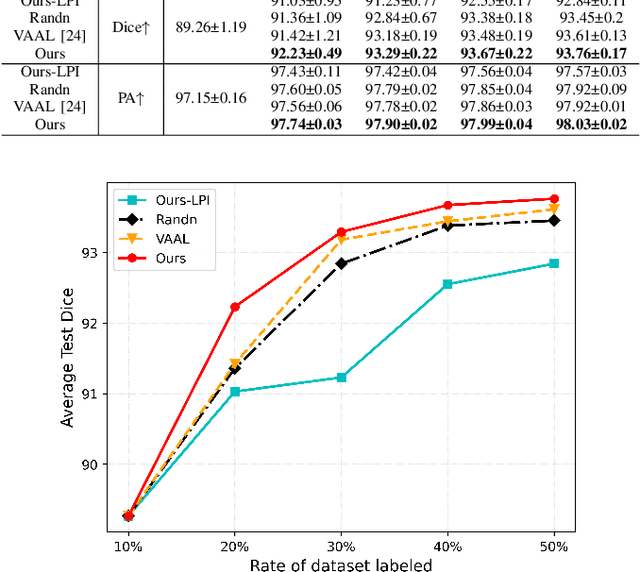
In recent years, deep learning has become a breakthrough technique in assisting medical image diagnosis. Supervised learning using convolutional neural networks (CNN) provides state-of-the-art performance and has served as a benchmark for various medical image segmentation and classification. However, supervised learning deeply relies on large-scale annotated data, which is expensive, time-consuming, and even impractical to acquire in medical imaging applications. Active Learning (AL) methods have been widely applied in natural image classification tasks to reduce annotation costs by selecting more valuable examples from the unlabeled data pool. However, their application in medical image segmentation tasks is limited, and there is currently no effective and universal AL-based method specifically designed for 3D medical image segmentation. To address this limitation, we propose an AL-based method that can be simultaneously applied to 2D medical image classification, segmentation, and 3D medical image segmentation tasks. We extensively validated our proposed active learning method on three publicly available and challenging medical image datasets, Kvasir Dataset, COVID-19 Infection Segmentation Dataset, and BraTS2019 Dataset. The experimental results demonstrate that our PCDAL can achieve significantly improved performance with fewer annotations in 2D classification and segmentation and 3D segmentation tasks. The codes of this study are available at https://github.com/ortonwang/PCDAL.
Architecture Disentanglement for Deep Neural Networks
Mar 30, 2020Jie Hu, Rongrong Ji, Qixiang Ye, Tong Tong, ShengChuan Zhang, Ke Li, Feiyue Huang, Ling Shao

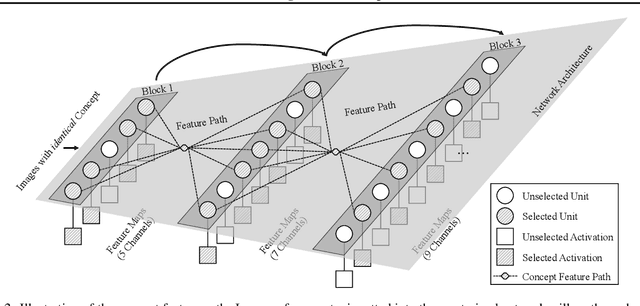
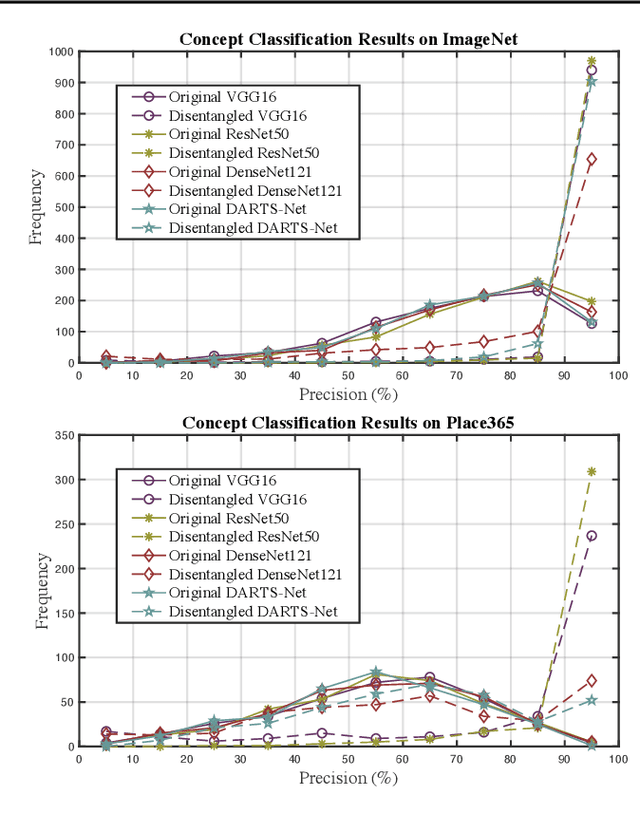
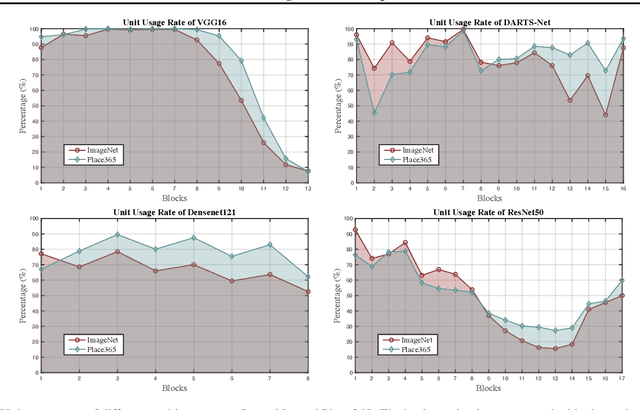
Deep Neural Networks (DNNs) are central to deep learning, and understanding their internal working mechanism is crucial if they are to be used for emerging applications in medical and industrial AI. To this end, the current line of research typically involves linking semantic concepts to a DNN's units or layers. However, this fails to capture the hierarchical inference procedure throughout the network. To address this issue, we introduce the novel concept of Neural Architecture Disentanglement (NAD) in this paper. Specifically, we disentangle a pre-trained network into hierarchical paths corresponding to specific concepts, forming the concept feature paths, i.e., the concept flows from the bottom to top layers of a DNN. Such paths further enable us to quantify the interpretability of DNNs according to the learned diversity of human concepts. We select four types of representative architectures ranging from handcrafted to autoML-based, and conduct extensive experiments on object-based and scene-based datasets. Our NAD sheds important light on the information flow of semantic concepts in DNNs, and provides a fundamental metric that will facilitate the design of interpretable network architectures. Code will be available at: https://github.com/hujiecpp/NAD.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge